Week 3 HW: Lab automation
Python Script for Opentrons Artwork
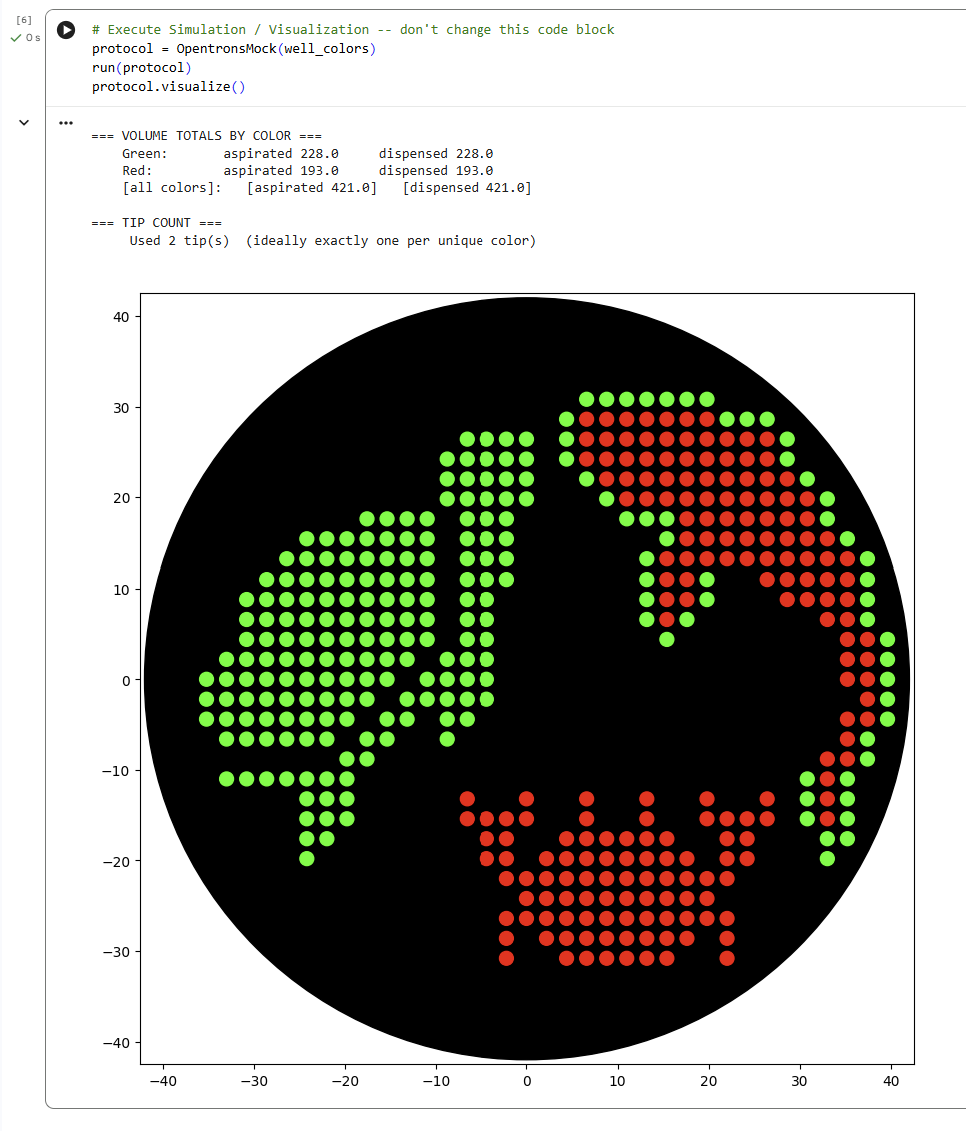
For this lab, I generated an artistic design using the GUI at opentrons-art.rcdonovan.com. I decided to design some undersea animals because I really like being underwater and observing their nature. Given that in my node’s lab there are only fluorescent green and red, I decided to draw a red crab, a green turtle, and a red and green fish.


The coordinates given are the following: mrfp1_points = [(6.6,28.6), (8.8,28.6), (11,28.6), (13.2,28.6), (15.4,28.6), (17.6,28.6), (19.8,28.6), (6.6,26.4), (8.8,26.4), (11,26.4), (13.2,26.4), (15.4,26.4), (17.6,26.4), (19.8,26.4), (22,26.4), (24.2,26.4), (26.4,26.4), (6.6,24.2), (8.8,24.2), (11,24.2), (13.2,24.2), (15.4,24.2), (17.6,24.2), (19.8,24.2), (22,24.2), (24.2,24.2), (26.4,24.2), (8.8,22), (11,22), (13.2,22), (15.4,22), (17.6,22), (19.8,22), (22,22), (24.2,22), (26.4,22), (28.6,22), (11,19.8), (13.2,19.8), (15.4,19.8), (17.6,19.8), (19.8,19.8), (22,19.8), (24.2,19.8), (26.4,19.8), (28.6,19.8), (30.8,19.8), (17.6,17.6), (19.8,17.6), (22,17.6), (24.2,17.6), (26.4,17.6), (28.6,17.6), (30.8,17.6), (17.6,15.4), (19.8,15.4), (22,15.4), (24.2,15.4), (26.4,15.4), (28.6,15.4), (30.8,15.4), (33,15.4), (15.4,13.2), (17.6,13.2), (19.8,13.2), (22,13.2), (24.2,13.2), (26.4,13.2), (28.6,13.2), (30.8,13.2), (33,13.2), (35.2,13.2), (15.4,11), (17.6,11), (26.4,11), (28.6,11), (30.8,11), (33,11), (35.2,11), (15.4,8.8), (17.6,8.8), (28.6,8.8), (30.8,8.8), (33,8.8), (35.2,8.8), (15.4,6.6), (33,6.6), (35.2,6.6), (35.2,4.4), (37.4,4.4), (35.2,2.2), (37.4,2.2), (35.2,0), (37.4,0), (37.4,-2.2), (35.2,-4.4), (37.4,-4.4), (35.2,-6.6), (33,-8.8), (35.2,-8.8), (33,-11), (-6.6,-13.2), (0,-13.2), (6.6,-13.2), (13.2,-13.2), (19.8,-13.2), (26.4,-13.2), (33,-13.2), (-6.6,-15.4), (-4.4,-15.4), (-2.2,-15.4), (0,-15.4), (6.6,-15.4), (13.2,-15.4), (19.8,-15.4), (22,-15.4), (24.2,-15.4), (26.4,-15.4), (33,-15.4), (-4.4,-17.6), (-2.2,-17.6), (4.4,-17.6), (6.6,-17.6), (8.8,-17.6), (11,-17.6), (13.2,-17.6), (15.4,-17.6), (22,-17.6), (24.2,-17.6), (-4.4,-19.8), (-2.2,-19.8), (2.2,-19.8), (4.4,-19.8), (6.6,-19.8), (8.8,-19.8), (11,-19.8), (13.2,-19.8), (15.4,-19.8), (17.6,-19.8), (22,-19.8), (24.2,-19.8), (-2.2,-22), (0,-22), (2.2,-22), (4.4,-22), (6.6,-22), (8.8,-22), (11,-22), (13.2,-22), (15.4,-22), (17.6,-22), (19.8,-22), (22,-22), (0,-24.2), (2.2,-24.2), (4.4,-24.2), (6.6,-24.2), (8.8,-24.2), (11,-24.2), (13.2,-24.2), (15.4,-24.2), (17.6,-24.2), (19.8,-24.2), (-2.2,-26.4), (0,-26.4), (2.2,-26.4), (4.4,-26.4), (6.6,-26.4), (8.8,-26.4), (11,-26.4), (13.2,-26.4), (15.4,-26.4), (17.6,-26.4), (19.8,-26.4), (22,-26.4), (-2.2,-28.6), (2.2,-28.6), (4.4,-28.6), (6.6,-28.6), (8.8,-28.6), (11,-28.6), (13.2,-28.6), (15.4,-28.6), (17.6,-28.6), (22,-28.6), (-2.2,-30.8), (4.4,-30.8), (6.6,-30.8), (8.8,-30.8), (11,-30.8), (13.2,-30.8), (15.4,-30.8), (22,-30.8)] sfgfp_points = [(6.6,30.8), (8.8,30.8), (11,30.8), (13.2,30.8), (15.4,30.8), (17.6,30.8), (19.8,30.8), (4.4,28.6), (22,28.6), (24.2,28.6), (26.4,28.6), (-6.6,26.4), (-4.4,26.4), (-2.2,26.4), (0,26.4), (4.4,26.4), (28.6,26.4), (-8.8,24.2), (-6.6,24.2), (-4.4,24.2), (-2.2,24.2), (0,24.2), (4.4,24.2), (28.6,24.2), (-8.8,22), (-6.6,22), (-4.4,22), (-2.2,22), (0,22), (6.6,22), (30.8,22), (-8.8,19.8), (-6.6,19.8), (-4.4,19.8), (-2.2,19.8), (0,19.8), (8.8,19.8), (33,19.8), (-17.6,17.6), (-15.4,17.6), (-13.2,17.6), (-11,17.6), (-6.6,17.6), (-4.4,17.6), (-2.2,17.6), (11,17.6), (13.2,17.6), (15.4,17.6), (33,17.6), (-24.2,15.4), (-22,15.4), (-19.8,15.4), (-17.6,15.4), (-15.4,15.4), (-13.2,15.4), (-11,15.4), (-6.6,15.4), (-4.4,15.4), (-2.2,15.4), (15.4,15.4), (35.2,15.4), (-26.4,13.2), (-24.2,13.2), (-22,13.2), (-19.8,13.2), (-17.6,13.2), (-15.4,13.2), (-13.2,13.2), (-11,13.2), (-6.6,13.2), (-4.4,13.2), (-2.2,13.2), (13.2,13.2), (37.4,13.2), (-28.6,11), (-26.4,11), (-24.2,11), (-22,11), (-19.8,11), (-17.6,11), (-15.4,11), (-13.2,11), (-11,11), (-6.6,11), (-4.4,11), (-2.2,11), (13.2,11), (19.8,11), (37.4,11), (-30.8,8.8), (-28.6,8.8), (-26.4,8.8), (-24.2,8.8), (-22,8.8), (-19.8,8.8), (-17.6,8.8), (-15.4,8.8), (-13.2,8.8), (-11,8.8), (-6.6,8.8), (-4.4,8.8), (13.2,8.8), (19.8,8.8), (37.4,8.8), (-30.8,6.6), (-28.6,6.6), (-26.4,6.6), (-24.2,6.6), (-22,6.6), (-19.8,6.6), (-17.6,6.6), (-15.4,6.6), (-13.2,6.6), (-11,6.6), (-6.6,6.6), (-4.4,6.6), (13.2,6.6), (17.6,6.6), (37.4,6.6), (-30.8,4.4), (-28.6,4.4), (-26.4,4.4), (-24.2,4.4), (-22,4.4), (-19.8,4.4), (-17.6,4.4), (-15.4,4.4), (-13.2,4.4), (-11,4.4), (-6.6,4.4), (-4.4,4.4), (15.4,4.4), (39.6,4.4), (-33,2.2), (-30.8,2.2), (-28.6,2.2), (-26.4,2.2), (-24.2,2.2), (-22,2.2), (-19.8,2.2), (-17.6,2.2), (-15.4,2.2), (-13.2,2.2), (-8.8,2.2), (-6.6,2.2), (-4.4,2.2), (39.6,2.2), (-35.2,0), (-33,0), (-30.8,0), (-28.6,0), (-26.4,0), (-24.2,0), (-22,0), (-19.8,0), (-17.6,0), (-15.4,0), (-11,0), (-8.8,0), (-6.6,0), (-4.4,0), (39.6,0), (-35.2,-2.2), (-33,-2.2), (-30.8,-2.2), (-28.6,-2.2), (-26.4,-2.2), (-24.2,-2.2), (-22,-2.2), (-19.8,-2.2), (-17.6,-2.2), (-13.2,-2.2), (-11,-2.2), (-8.8,-2.2), (-6.6,-2.2), (-4.4,-2.2), (39.6,-2.2), (-35.2,-4.4), (-33,-4.4), (-30.8,-4.4), (-28.6,-4.4), (-26.4,-4.4), (-24.2,-4.4), (-22,-4.4), (-19.8,-4.4), (-15.4,-4.4), (-13.2,-4.4), (-8.8,-4.4), (-6.6,-4.4), (39.6,-4.4), (-33,-6.6), (-30.8,-6.6), (-28.6,-6.6), (-26.4,-6.6), (-24.2,-6.6), (-22,-6.6), (-17.6,-6.6), (-15.4,-6.6), (-8.8,-6.6), (37.4,-6.6), (-19.8,-8.8), (-17.6,-8.8), (37.4,-8.8), (-33,-11), (-30.8,-11), (-28.6,-11), (-26.4,-11), (-24.2,-11), (-22,-11), (-19.8,-11), (30.8,-11), (35.2,-11), (-24.2,-13.2), (-22,-13.2), (-19.8,-13.2), (30.8,-13.2), (35.2,-13.2), (-24.2,-15.4), (-22,-15.4), (-19.8,-15.4), (30.8,-15.4), (35.2,-15.4), (-24.2,-17.6), (-22,-17.6), (33,-17.6), (35.2,-17.6), (-24.2,-19.8), (33,-19.8)]
Later, I simulated the design in the Opentrons Colab. This protocol programs the Opentrons robot to create an artistic pattern on an agar plate using two fluorescent proteins (mRFP1 in red and sfGFP in green).
First, the robot loads the necessary labware: a 20 µL tip rack, a temperature module holding the color plate (agar plate with dyes), and the agar plate where the design will be drawn. The pipette is initialized and set to start from a specific tip.
The script defines helper functions:
location_of_color() finds the well containing a specific color.
dispense_and_detach() carefully deposits a droplet onto the agar surface by approaching from above, dispensing, and lifting back up to avoid smearing.
Two lists of (x, y) coordinates define the artwork. These coordinates represent positions relative to the center of the agar plate.
The code: Download my Opentron Python script
After doing the simulation, I filled out the Google form and signed up for a timeslot to run the code on the robot:

Post-Lab Questions
Paper: BOTany Methods: Accessible Automation for Plant Synthetic Biology
The paper I found describes how researchers used automated liquid handling systems to process biological samples in a standardized way. Instead of manually pipetting every reaction, they used robotic tools to precisely mix reagents, prepare samples, and run assays with much less human intervention. This allowed them to reduce variability, increase reproducibility, and process many samples at the same time. Automation was especially important because small pipetting differences can affect biological results. By using robotics, the researchers were able to make their workflow more scalable and reliable, which is critical in biomedical research where consistency matters.
For my final project, I would like to use an Opentrons robot to automate the testing of DNA-based logic circuits using cell-free protein synthesis. The robot would transfer DNA constructs into a well plate, add the expression mix, dispense cofactors, and incubate the reactions before measuring fluorescence output. This would allow me to test many different circuit designs at once and compare their outputs automatically. I could also design a small 3D-printed holder to support custom plates or patterned substrates. Overall, automation would help me prototype biological computation systems faster and more accurately, while reducing human error and making the experiments more scalable.
Final Project Ideas
In slides for committed listeners.