Week 4 HW: Protein Design Part 1
Part A. Conceptual Questions
1. How many molecules of amino acids do you take with a piece of 500 grams of meat? (on average an amino acid is ~100 Daltons)
Amino acids have an average mass of about 100 Daltons, which is approximately 100 g per mole. If we assume that most of the mass of meat comes from proteins, we can estimate the number of amino acids in 500 g of meat to be 3*10^24 amino acid molecules.
2. Why do humans eat beef but do not become a cow, eat fish but do not become fish?
When we eat food, our digestive system breaks proteins down into individual amino acids using enzymes. These amino acids are absorbed into the bloodstream and used by our cells to build human proteins, not cow or fish proteins.
The instructions for building proteins come from our DNA, which determines the structure of the proteins our cells produce. Therefore, even though the amino acids come from other organisms, they are reassembled according to human genetic instructions, so we remain human.
3. Why are there only 20 natural amino acids?
There are 20 standard amino acids used in proteins because evolution selected a set that provides enough chemical diversity to build stable and functional proteins. These amino acids include different properties such as hydrophobic, polar, charged, aromatic.
Together, they allow proteins to fold into many complex structures and perform many functions. While other amino acids exist, the 20 canonical ones became part of the genetic code early in evolution, and this system remained conserved because it works efficiently for life.
5. Where did amino acids come from before enzymes that make them, and before life started?
Before life existed, amino acids were likely formed through prebiotic chemical reactions on early Earth. Experiments like the Miller-Urey experiment showed that amino acids can form when simple molecules such as methane, ammonia, water, and hydrogen are exposed to energy sources like lightning.
Amino acids may also have come from meteorites, which have been found to contain organic molecules. These sources suggest that amino acids could form naturally in the environment before enzymes or living organisms existed.
6. If you make an α-helix using D-amino acids, what handedness (right or left) would you expect?
Proteins made from L-amino acids, which are the natural form in biology, usually form right-handed α-helices.
If the helix were made from D-amino acids, the geometry would be mirrored, so the helix would form a left-handed α-helix.
7. Can you discover additional helices in proteins?
Yes, it is possible to discover additional helices. The α-helix is the most common helix found in proteins, but other helical structures exist, such as 3₁₀ helices and π-helices.
Using computational protein design, structural biology, and protein engineering, scientists can also design new helices with different properties. Advances in synthetic biology and protein modeling tools like AlphaFold or Rosetta make it easier to identify or design new helical structures.
8. Why are most molecular helices right-handed?
Most helices in biological proteins are right-handed because proteins are built from L-amino acids. The stereochemistry of L-amino acids favors a right-handed helix because it allows better bond angles and fewer steric clashes between atoms.
This configuration is therefore energetically more stable, which is why evolution selected it.
9. Why do β-sheets tend to aggregate? - What is the driving force for β-sheet aggregation?##
β-sheets tend to aggregate because their structure allows strong hydrogen bonding between neighbouring strands. When β-sheets from different proteins align, they can form extended networks of hydrogen bonds.
The main driving forces are hydrogen bonding between peptide backbones, hydrophobic interactions, and stacking of β-strands. These interactions stabilize large sheet-like structures, which can lead to aggregation.
9. Why do many amyloid diseases form β-sheets? - Can you use amyloid β-sheets as materials?
Many amyloid diseases form β-sheets because misfolded proteins can rearrange into stable β-sheet-rich structures. These sheets stack together into amyloid fibrils, which are very stable and difficult for cells to break down. This aggregation can disrupt normal cellular function and lead to diseases such as Alzheimer’s or Parkinson’s disease.
However, amyloid β-sheets also have useful properties. Because they are extremely strong, stable, and self-assembling, scientists are studying them as materials for nanofibers, biomaterials, scaffolds for tissue engineering, and nanotechnology applications. So although amyloids can cause disease, their structural properties could also be used in synthetic biology and materials science.
Part B: Protein Analysis and Visualization
Briefly describe the protein you selected and why you selected it.
The protein I selected is NaChBac (Bacterial Voltage-Gated Sodium Channel). NaChBac is a voltage-gated sodium ion channel originally discovered in bacteria. It allows sodium ions to pass through the cell membrane when the membrane voltage changes. This flow of ions can create electrical signals similar to action potentials in neurons. I selected NaChBac because sodium channels are essential for neuronal signaling and spike generation, which are key mechanisms in biological neural networks. NaChBac is simpler than human sodium channels and easier to study or engineer, which makes it useful for synthetic biology and bio-inspired computing systems. Using proteins like NaChBac could help design biological circuits that behave similarly to neurons, which could eventually contribute to new approaches to neuromorphic or bio-based AI systems which is what I would like to work in for my final project.
After searching for its sequence in PDB, there were multiple structures for this protein. I selected PDB structure 6VWX, which represents the NaChBac sodium channel in a lipid nanodisc at 3.1 Å resolution determined by cryo-electron microscopy. This structure provides the highest resolution available among the listed options and represents the channel in a membrane-like environment, which is important because NaChBac is a transmembrane ion channel. Using a lipid nanodisc preserves the native conformation of membrane proteins better than detergent conditions. Therefore, 6VWX provides the most biologically relevant structure for analyzing the architecture and function of the NaChBac channel.
Identify the amino acid sequence of your protein.
Amino acid sequence:
MKMEARQKQNSFTSKMQKIVNHRAFTFTVIALILFNALIVGIETYPRIYADHKWLFYRIDLVILWIFTIEIAMRFLASNPKSAFFRSSWNWDFLIVAAGHIFAGAQFVTIVLRILRVLRVLRAISVVPSLRRLVDALVMTIPALGNILILMSIFFYIFAVIGTMLFQHVSPEYFGNLQLSLLTLFQVVTLESWASGVMRPIFAEVPWSWLYFVSFVLIGTFITFNLFIGVIVNNVEKAELTDNEEDGEADGLKQEISALRKDVAELKSLLKQSK
- How long is it?
274 aminoacids.
- What is the most frequent amino acid?
Leucine (L), which appears 34 times.
- How many protein sequence homologs are there for your protein?
2: A0A4Q0VN24 and A0ABS6JN79
- Does your protein belong to any protein family?
Yes. NaChBac belongs to the voltage-gated sodium channel (Nav) protein family.
Identify the structure page of your protein in RCSB
- When was the structure solved?
The structure was solved in 2020, deposited on February 20, 2020 and released to PDB on June 24, 2020.
- Is it a good quality structure? Good quality structure is the one with good resolution. Smaller the better.
The resolution is 3.10 Å, determined using cryo-electron microscopy. This number indicates that the overall protein structure and most side chains can be reliably modeled.
- Are there any other molecules in the solved structure apart from protein?
Yes, besides the protein, the structure also contains sodium ions (Na⁺) and lipid molecules (POV). These molecules help stabilize the sodium channel and mimic the membrane environment in which the protein normally functions.
- Does your protein belong to any structure classification family?
Yes. NaChBac belongs to the voltage-gated sodium channel structural family, which is part of the larger voltage-gated ion channel superfamily.
Structure of your protein in the 3D molecule visualization software PyMol
- Cartoon visualization

- Ribbon visualization


- Ball and stick visualization


- Color the protein by secondary structure. Does it have more helices or sheets?


Color the protein by residue type. What can you tell about the distribution of hydrophobic vs hydrophilic residues?
Visualize the surface of the protein. Does it have any “holes” (aka binding pockets)?
Part C. Using ML-Based Protein Design Tools
C1. Protein Language Modeling
C2. Protein Folding
C3. Protein Generation
Part D. Group Brainstorm on Bacteriophage Engineering
Computational Engineering of the MS2 Lysis Protein to Improve Stability, Titers, and Toxicity
Group members: Emmanuel Pereyra, Sergio Cuiza, Domenica Vizcaino, Diana Grimaldos
Selected Goals
After reviewing the provided literature on the MS2 lysis protein (L) and discussing the project aims, our group has decided to focus on three interconnected goals:
Goal 1: Increase the stability of the L protein.
Rationale: As the “easiest” goal, it is the most computationally tractable. A stabilized protein is less prone to degradation and misfolding, which could directly lead to higher functional titers and serve as a robust starting point for any subsequent engineering.
Goal 2: Increase bacteriophage titers through improved lysis efficiency.
Phage therapy relies on high phage titers for effective bacterial killing and scalable manufacturing, but phage production can be limited by inefficient lysis or poor coordination between phage replication and host destruction. Improving the efficiency and timing of host cell lysis can therefore directly increase the number of phage particles released per infected cell.
The MS2 L protein is a small 75–amino acid membrane protein that triggers bacterial lysis and is essential for the release of new phage particles. In the paper Mutational analysis of the MS2 lysis protein L, it is described how MS2 L functions as a single-gene lysis protein that disrupts bacterial cell envelope integrity without classical enzymatic activity. Additionally, L interacts with the host chaperone DnaJ, which modulates its activity and timing of lysis. In MS2 Lysis of Escherichia coli Depends on Host Chaperone DnaJ it is shown that lysis timing strongly affects the number of virions produced before the host cell bursts, meaning that engineering improved L variants may increase overall phage titers.
Goal 3: Increase the toxicity of the lysis protein.
This proposal addresses the subproblem of increasing the toxicity of the L lysis protein from Bacteriophage MS2. Instead of random mutagenesis, toxicity will be approached as a multi-factor optimization problem involving structural stability, membrane insertion, oligomerization efficiency, and expression kinetics in Escherichia coli. The objective is to design L variants that enhance membrane disruption while maintaining proper folding and stability.
Additionally, we will explore disrupting the interaction between the L protein and the E. coli chaperone DnaJ.
Rationale: The reading “Identification MS2 lysis protein dependency on DnaJ” establishes this interaction as critical for function. By computationally predicting and then disrupting this interface, we can test its necessity and potentially create a DnaJ-independent lysis mechanism, offering a new avenue for controlling lysis timing.
Together, these three goals form a coherent strategy: stabilizing the L protein may improve its folding and expression, which can increase functional titers, while further engineering of membrane disruption and host interactions may increase toxicity and lysis efficiency.
Proposed Computational Tools and Approaches
Proposed Tools and Approaches We will build a computational pipeline using the tools introduced in recitation and the provided resources. The key steps and tools are:
Step 1: Structural Modeling of the L Protein
Tool: AlphaFold2 (via ColabFold for ease of use).
Why: No high-resolution experimental structure of the full-length MS2 L protein exists. A reliable 3D model is the absolute foundation for all downstream analysis, allowing us to visualize which parts are structured vs. disordered.
Step 2: Modeling the L-DnaJ Complex
Tool: AlphaFold-Multimer.
Why: To disrupt the interaction, we first need to know where it occurs. AlphaFold-Multimer is the current state-of-the-art for predicting protein-protein complexes and will generate a testable model of the L protein bound to E. coli DnaJ.
Step 3: In Silico Mutagenesis for Stability
Tool: Rosetta (or FoldX). Specifically, the ddg_monomer application for predicting changes in folding free energy (ΔΔG).
Why: These tools are parameterized using vast amounts of experimental data on protein stability. They can systematically mutate each residue in our L protein model and predict whether the change (e.g., A->V) makes the protein more stable (negative ΔΔG) or less stable (positive ΔΔG).
Step 4: Visualizing and Selecting Interface Mutations
Tool: PyMOL and the HTGAA Protein Engineering Tools spreadsheet.
Why: We will use PyMOL to visually inspect the predicted L-DnaJ complex from Step 2 and select residues at the interface. We will then use the spreadsheet to check the conservation of those residues and manually design mutations (e.g., swapping a large hydrophobic residue for a charged one) predicted to break the interaction.
Protein Language Models (PLMs)
Protein language models such as ESM or ProtBERT will be used to perform in silico mutagenesis on the MS2 L protein sequence. These models can suggest mutations that preserve structural and functional constraints learned from large protein datasets.
This approach allows us to generate multiple candidate mutations across the L protein, avoid mutations likely to disrupt folding, and explore sequence space beyond naturally occurring variants.
AlphaFold Structure Prediction
Each candidate L variant will be analyzed using AlphaFold to predict protein structure and membrane topology. Since the C-terminal transmembrane region is essential for lytic activity, structural prediction will help identify mutations that preserve this functional domain.
Structural predictions will also help identify:
- misfolded variants
- mutations that destabilize the transmembrane region
- variants that may alter oligomerization or membrane insertion
Interaction Modeling with Host Proteins
Because MS2 L interacts with the DnaJ chaperone, which affects lysis timing, candidate variants can be evaluated using AlphaFold-Multimer to predict changes in the L–DnaJ interaction.
This could help identify variants that:
- maintain necessary folding assistance
- reduce excessive dependency on host chaperones
- improve robustness of lysis across physiological conditions
Proposed Computational Strategy
First, protein language models (e.g., ESM-2, ProtT5) will be used to perform directed in silico mutagenesis. These models capture evolutionary constraints and residue interactions, enabling the generation of structurally plausible variants while identifying mutation-tolerant and functionally critical positions. This step efficiently reduces the combinatorial search space.
Second, predicted variants will be structurally evaluated using AlphaFold2 for monomer folding and AlphaFold - Multimer to assess oligomerization and interaction with host factors such as DnaJ.
Third, membrane compatibility will be analyzed using membrane-aware modeling (RosettaMP) and selected molecular dynamics simulations.
Fourth, ΔΔG prediction tools (e.g., FoldX, Rosetta energy functions) will filter out destabilizing mutations.
In parallel, codon optimization algorithms will redesign selected variants for improved expression in E. coli, as toxicity depends on both structure and intracellular concentration.
Why These Tools Will Help
Why These Tools Will Help This pipeline is powerful because it moves from the general to the specific.
AlphaFold2/3 provides the necessary atomic-resolution context, transforming a sequence into a tangible structure we can analyze.
Rosetta leverages that structural context to make quantitative, physics-based predictions about stability.
AlphaFold-Multimer extends this to the biological mechanism, allowing us to generate a hypothesis about the DnaJ interaction that is currently unknown.
PyMOL enables the crucial final step of human intuition, allowing us to filter computational predictions through biological reasoning.
Rationale
Toxicity emerges from the combination of folding stability, cooperative oligomerization, membrane insertion, and sufficient expression levels.
These computational tools allow us to screen large numbers of protein variants without performing wet-lab experiments first.
Previous studies show that mutations in specific regions of L can abolish lysis function, indicating that the protein’s structure and interactions are highly sensitive to sequence changes.
Additionally, new AI-based methods are increasingly being used to design bacteriophages and improve phage performance.
Potential Pitfalls
Pitfall 1: Dynamic Regions and Model Quality
The L protein is small and likely has flexible/disordered regions, especially in its N-terminal domain.
Pitfall 2: Stability vs. Function Trade-off
A mutation that makes the protein more stable in its monomeric state might prevent it from undergoing the necessary conformational changes to oligomerize and form a pore in the membrane.
Pitfall 3: Lack of Membrane Context
Our stability predictions (Rosetta) are performed in a virtual “aqueous” environment and do not account for the energetic complexity of the lipid bilayer.
Limited biological data: There is still limited structural and mechanistic knowledge about MS2 L.
Cellular context not captured computationally Protein modeling tools may not fully capture membrane environment.
One limitation is the scarcity of quantitative datasets linking specific mutations to measured lysis kinetics.
Pipeline
We have developed three different pipelines to address each goal more specifically. The images were generated with AI:
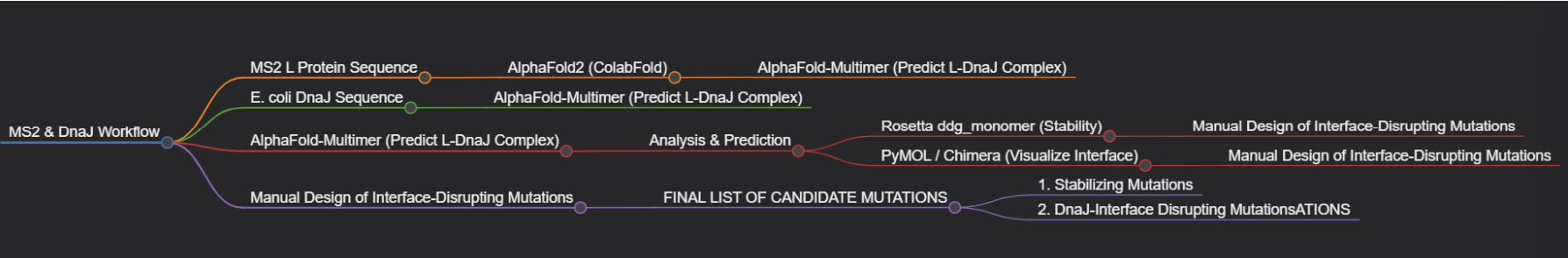
Goal 1.
Group’s Short Plan for Engineering a Bacteriophage: Our group will computationally engineer the MS2 lysis protein to enhance its utility. First, we will use AlphaFold to model the protein and its complex with the host factor DnaJ. We will then employ Rosetta to perform in silico saturation mutagenesis, identifying point mutations that increase the protein’s predicted stability. Concurrently, using the AlphaFold-Multimer model, we will design mutations at the L-DnaJ interface intended to disrupt this key interaction.

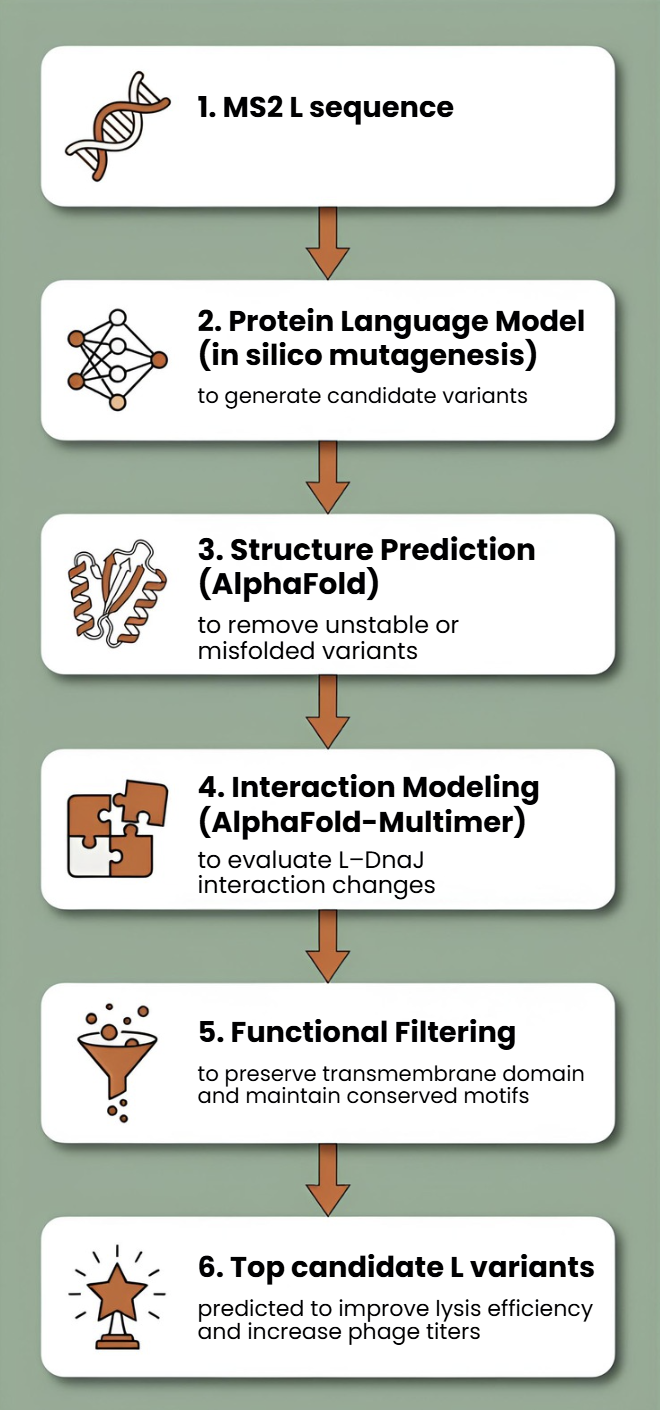
Goal 2.


Goal 3.
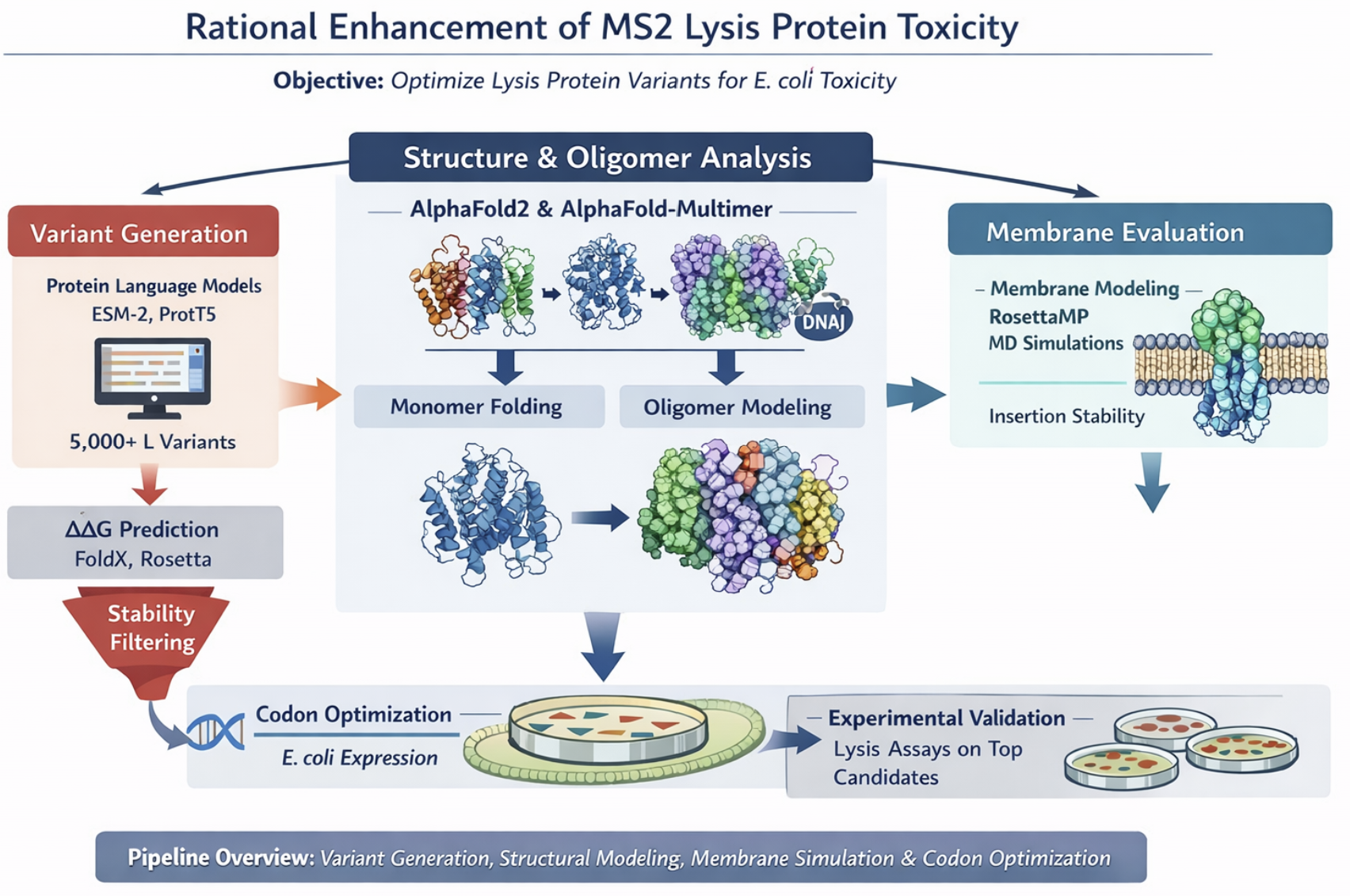
Generate ~5,000 variants with protein LLMs, filter by ΔΔG stability, predict structure and oligomers with AlphaFold, evaluate membrane behavior, optimize codons, select top candidates for experimental lysis assays.