Week 2 HW: DNA Read, Write, & Edit
Part 0: Basics of Gel Electrophoresis
Completed
Part 1: Benchling & In-silico Gel Art
After opening my Benchling account and joining the HTGAA group in Benchling, I imported the Lambda DNA.
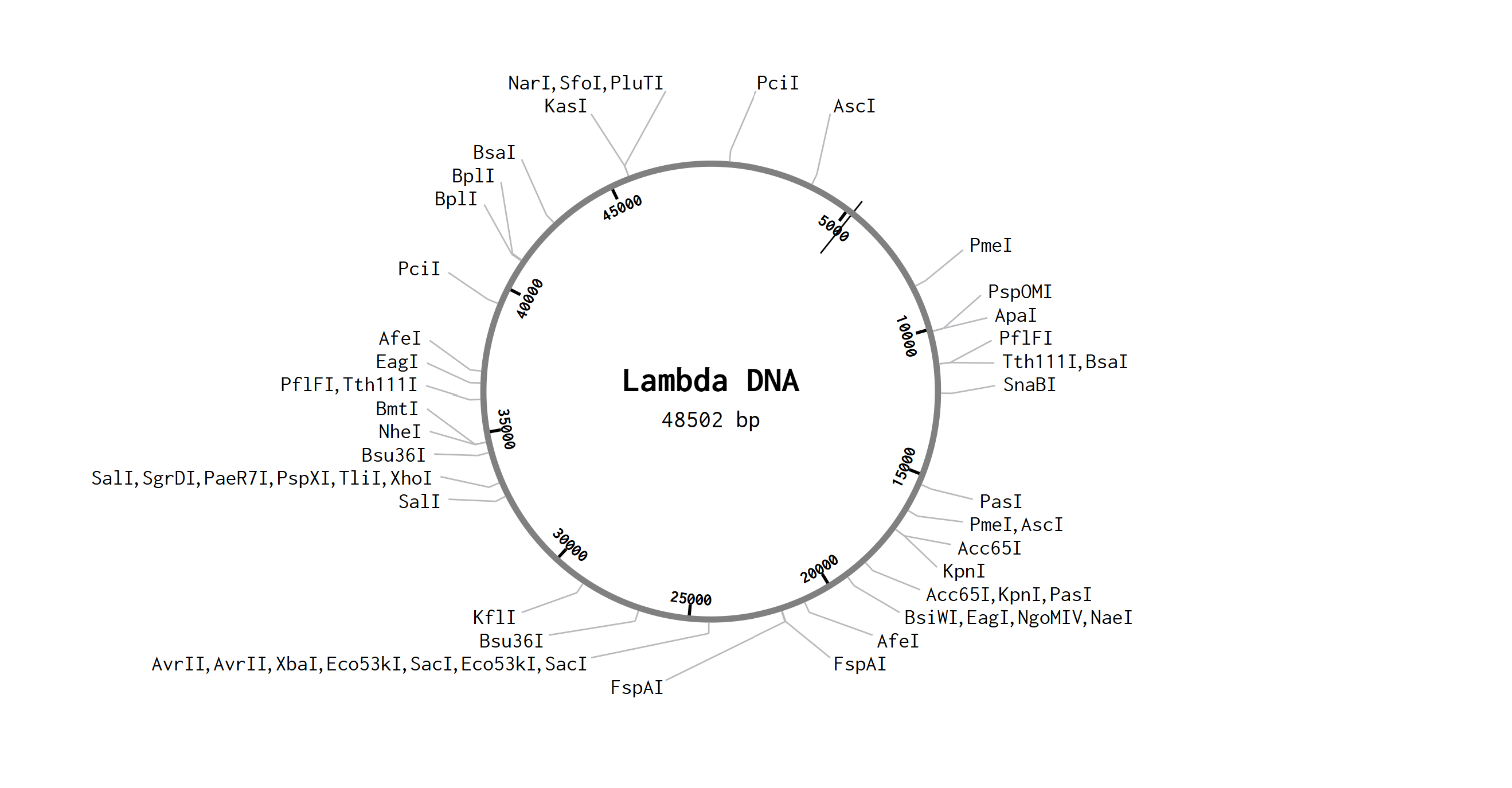

A restriction enzyme digestion was simulated in Benchling with the following enzymes:
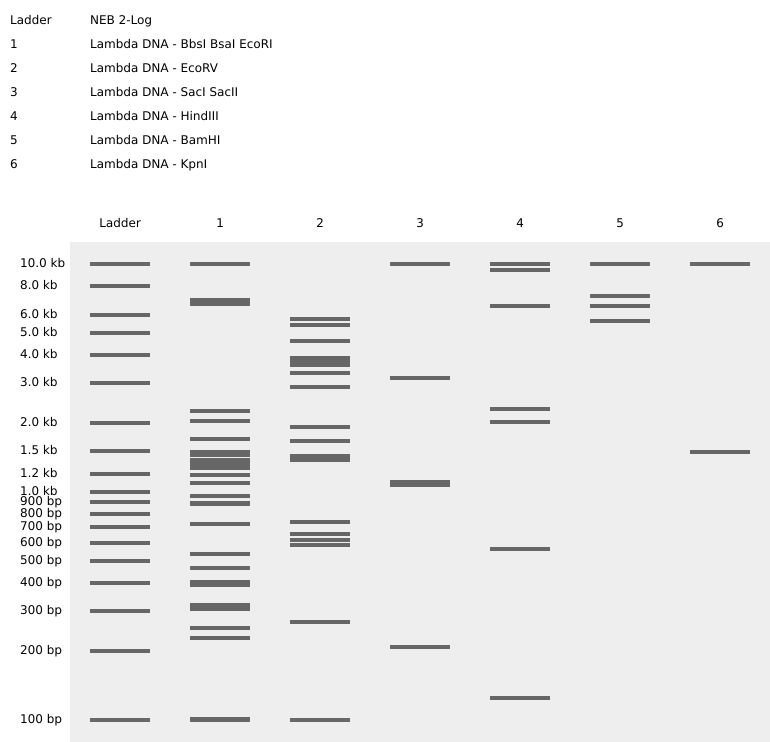


The image I tried to recreate in the style of Paul Vanouse’s latent Figure Protocol was a bird flying as seen below.




Part 2: Gel Art - Restriction Digests and Gel Electrophoresis
I do not have lab access.
Part 3: DNA Design Challenge
3.1. Choose your protein
I chose bacteriorhodopsin because it is a light-activated membrane protein that can convert light into electrochemical signals. This makes it interesting for synthetic biology applications in bio-electronics and biological computing which is what I would like to do my final project on. In particular, bacteriorhodopsin has been used in optical memory systems and protein-based neural networks, where its light-induced conformational changes allow it to function similarly to a switch or synaptic element. I am interested in future applications where biological molecules are integrated with electronic or neuromorphic systems for sensing, computing, or personalized medical technologies, so bacteriorhodopsin is a useful model protein to study.
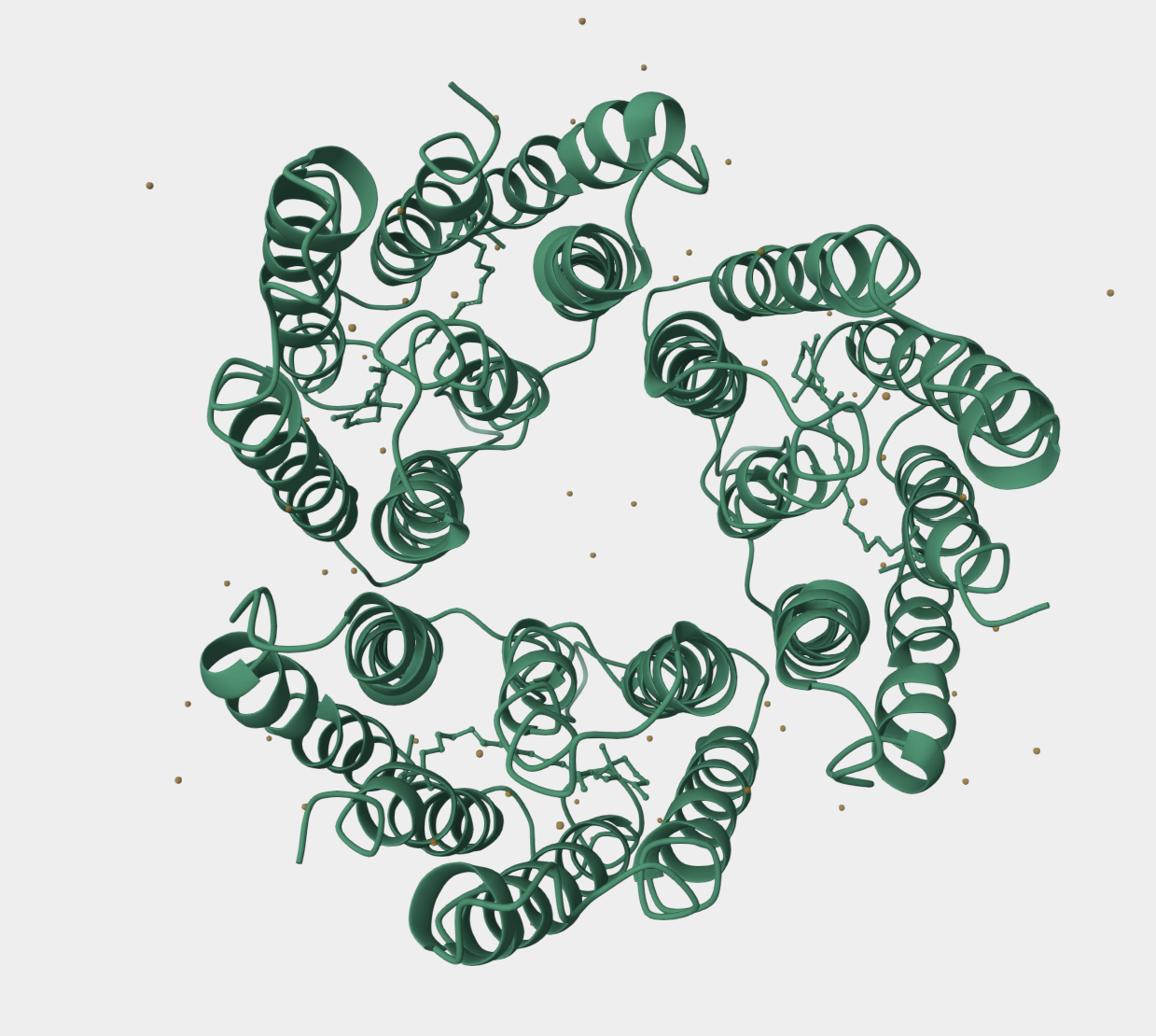

The following information was taken from UniProt:
UniProt entry: P02945 Organism: Halobacterium salinarum Protein: bacteriorhodopsin Organism taxonomy ID: 64091 Protein existence evidence: 1 Sequence version: 2
UniProt Format:
sp|P02945|BACR_HALSA Bacteriorhodopsin OS=Halobacterium salinarum OX=64091 PE=1 SV=2 MLELLPTAVEGVSQAQITGRPEWIWLALGTALMGLGTLYFLVKGMGVSDPDAKKFYAITTLVPAIAFTMYLSMLLGYGLTMVPFGGEQNPIYWARYADWLFTTPLLLLDLALLVDADQGTILALVGADGIMIGTGLVGALTKVYSYRFVWWAISTAAMLYILYVLFFGFTSKAESMRPEVASTFKVLRNVTVVLWSAYPVVWLIGSEGAGIVPLNIETLLFMVLDVSAKVGFGLILLRSRAIFGEAEAPEPSAGDGAAATSD
This protein has 262 amino acids.
3.2. Reverse Translate: Protein (amino acid) sequence to DNA (nucleotide) sequence
This sequence includes a stop codon at the end TAA
ATGTTAGAATTATTACCTACTGCTGTTGAAGGTGTTTCTCAAGCTCAAATTACTGGTCGTCCTGAATGGATTTGGTTAGCTTTAGGTACTGCTTTAATGGGTTTAGGTACTTTATATTTTTTAGTTAAAGGTATGGGTGTTTCTGATCCTGATGCTAAAAAATTTTATGCTATTACTACTTTAGTTCCTGCTATTGCTTTTACTATGTATTTATCTATGTTATTAGGTTATGGTTTAACTATGGTTCCTTTTGGTGGTGAACAAAATCCTATTTATTGGGCTCGTTATGCTGATTGGTTATTTACTACTCCTTTATTATTATTAGATTTAGCTTTATTAGTTGATGCTGATCAAGGTACTATTTTAGCTTTAGTTGGTGCTGATGGTATTATGATTGGTACTGGTTTAGTTGGTGCTTTAACTAAAGTTTATTCTTATCGTTTTGTTTGGTGGGCTATTTCTACTGCTGCTATGTTATATATTTTATATGTTTTATTTTTTGGTTTTACTTCTAAAGCTGAATCTATGCGTCCTGAAGTTGCTTCTACTTTTAAAGTTTTACGTAATGTTACTGTTGTTTTATGGTCTGCTTATCCTGTTGTTTGGTTAATTGGTTCTGAAGGTGCTGGTATTGTTCCTTTAAATATTGAAACTTTATTATTTATGGTTTTAGATGTTTCTGCTAAAGTTGGTTTTGGTTTAATTTTATTACGTTCTCGTGCTATTTTTGGTGAAGCTGAAGCTCCTGAACCTTCTGCTGGTGATGGTGCTGCTGCTACTTCTGATTAA
3.3. Codon optimization
Although many different DNA sequences can encode the same protein, organisms prefer certain codons over others. This is called codon bias. If a gene uses codons that are rare in the host organism, the protein may be expressed poorly because the necessary tRNAs are less abundant. Codon optimization adjusts the DNA sequence so that it uses codons preferred by the host organism while still encoding the same amino-acid sequence. This improves translation efficiency, protein yield, and overall stability of expression.
Codon optimization can also remove problematic sequences such as restriction enzyme sites, repeats, or regions that form strong secondary structures, which can interfere with cloning or gene synthesis.
I optimized the bacteriorhodopsin coding sequence for Escherichia coli expression. E. coli is one of the most commonly used organisms in synthetic biology because it grows quickly, is easy to genetically engineer, and has well-characterized expression systems. Optimizing the gene for E. coli would make it easier to clone and express bacteriorhodopsin in a laboratory setting for research or synthetic biology applications.
The sequence was codon optimized using the IDT codon optimization tool, avoiding Type IIs enzyme recognition sites BsaI, BsmBI, and BbsI. These were avoided because restriction enzymes like BsaI, BsmBI, and BbsI are commonly used in synthetic biology cloning methods (especially Golden Gate assembly). These enzymes recognize very specific DNA sequences and cut at those sites. If the gene contains one of those recognition sequences internally, the enzyme will cut the gene in the middle when trying to clone it. That would break the construct and prevent proper assembly.
ATG TTG GAA CTG TTA CCG ACC GCG GTT GAA GGA GTC TCT CAG GCG CAG ATT ACG GGG CGT CCA GAA TGG ATC TGG TTG GCA CTG GGA ACA GCT TTA ATG GGC TTA GGA ACT CTG TAT TTT TTA GTC AAG GGA ATG GGC GTT TCC GAC CCT GAT GCT AAA AAA TTT TAT GCA ATT ACC ACA CTG GTG CCA GCT ATT GCC TTC ACC ATG TAC TTG TCT ATG CTT CTG GGG TAT GGA CTT ACA ATG GTT CCG TTT GGC GGT GAG CAA AAT CCG ATT TAT TGG GCA CGT TAC GCG GAC TGG CTT TTT ACG ACG CCG TTA CTG TTA TTG GAT CTT GCA CTT CTG GTG GAC GCG GAT CAA GGT ACC ATT CTT GCA TTG GTG GGG GCT GAT GGG ATA ATG ATA GGC ACC GGT TTA GTG GGT GCA CTG ACA AAA GTA TAT TCA TAT CGC TTC GTG TGG TGG GCC ATC TCA ACA GCC GCC ATG CTT TAC ATA TTG TAT GTA TTG TTT TTT GGC TTT ACC TCG AAA GCA GAG AGT ATG CGT CCG GAA GTA GCG TCT ACA TTC AAA GTG TTG CGC AAT GTT ACG GTG GTG TTA TGG TCA GCC TAC CCC GTA GTA TGG CTG ATT GGT AGC GAG GGA GCG GGT ATT GTA CCG CTT AAT ATC GAA ACC CTG CTG TTC ATG GTA CTG GAC GTG TCG GCC AAG GTG GGC TTC GGC CTG ATA CTG TTA CGT AGC AGA GCA ATT TTC GGA GAA GCT GAA GCT CCA GAG CCT AGT GCA GGT GAT GGT GCA GCC GCA ACG TCA GAT TAA
3.4. You have a sequence! Now what?
The bacteriorhodopsin protein can be produced by first introducing the codon-optimized DNA sequence into an expression system. This can be done using either cell-dependent (living cells) or cell-free protein expression technologies.
Cell-dependent
One common method is to insert the codon-optimized gene into a plasmid vector and transform it into a host organism such as Escherichia coli. Inside the cell, the DNA is first transcribed into messenger RNA (mRNA) by RNA polymerase. The mRNA is then translated by ribosomes, which read the codons in the mRNA and assemble the corresponding amino acids into the bacteriorhodopsin protein. Transfer RNAs (tRNAs) bring amino acids to the ribosome based on codon-anticodon pairing, allowing the polypeptide chain to form. After translation, the protein folds into its functional structure and can be integrated into membranes or purified for further use.
Cell-free
The gene can be expressed using a cell-free protein synthesis system, which contains ribosomes, enzymes, nucleotides, and tRNAs without living cells. In this system, the DNA template is added directly to the reaction mixture, where it is transcribed into mRNA and translated into protein in vitro. Cell-free systems allow rapid protein production and easier control of conditions, which can be useful for synthetic biology and protein engineering experiments.
Part 4: Prepare a Twist DNA Synthesis Order
After opening my Twist and Benchling accounts, I followed the steps mentioned and obtained the following plasmid.
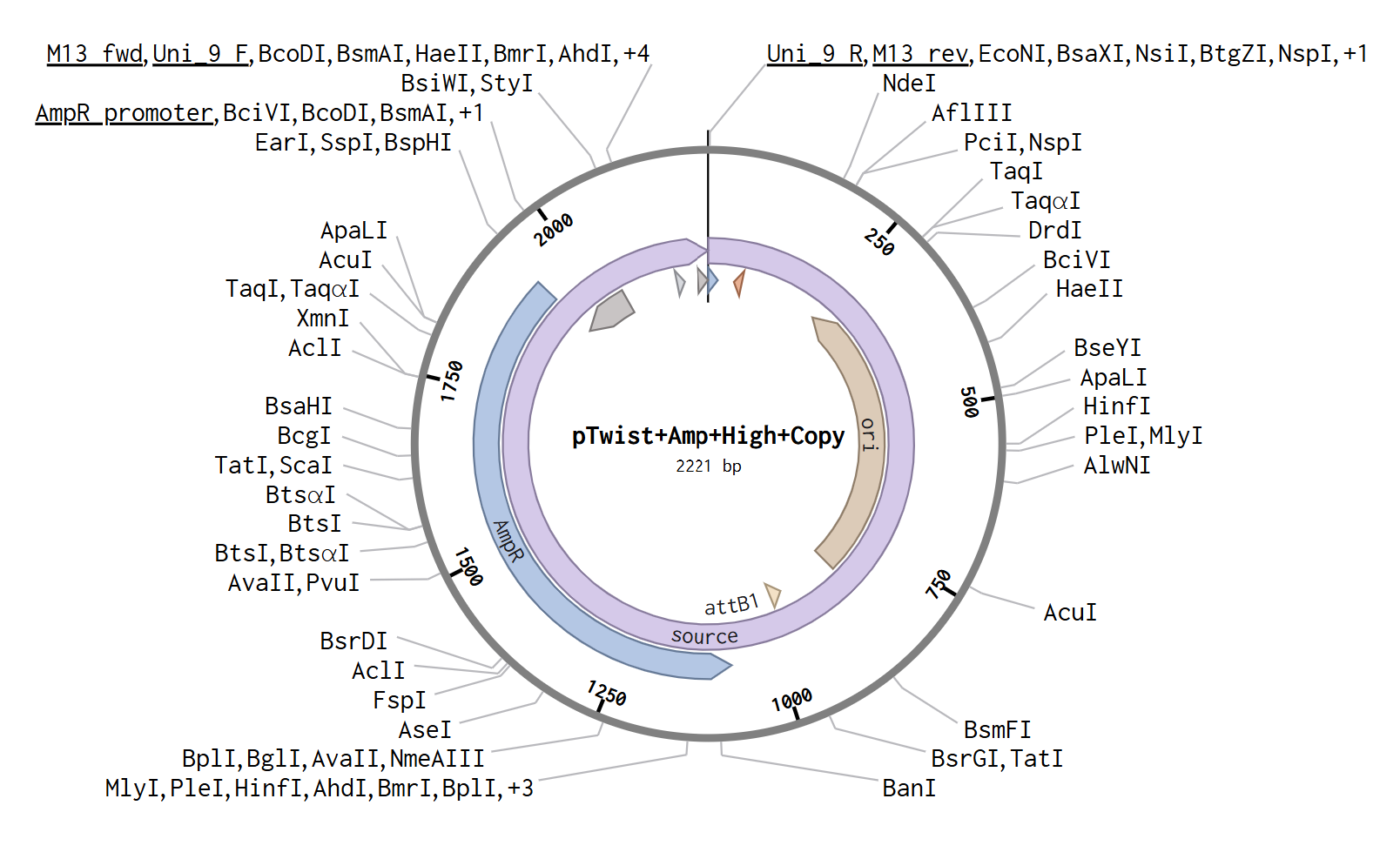

Part 5: DNA Read/Write/Edit
5.1 DNA Read
(i) What DNA would you want to sequence (e.g., read) and why?
I would want to sequence DNA used in synthetic biological computing systems, particularly DNA strands that encode digital information or act as molecular logic circuits. DNA can be used to store data, perform computations, and respond to biological signals. Sequencing these DNA molecules would allow us to read stored information, verify computational outputs, and detect mutations or errors that occur over time.
In the long term, sequencing DNA-based computing systems could enable personalized medicine applications. For example, DNA circuits inside cells could record molecular events such as exposure to drugs, stress signals, or disease biomarkers. Sequencing these DNA records would allow clinicians to understand how a patient’s cells responded to treatment and adjust therapies accordingly. Accurate DNA reading is therefore essential for both bio-computing and medical diagnostics.
(ii) In lecture, a variety of sequencing technologies were mentioned. What technology or technologies would you use to perform sequencing on your DNA and why?
I would use a combination of Illumina sequencing (second-generation) and Oxford Nanopore sequencing (third-generation).
Illumina sequencing provides high accuracy, which is useful for verifying computational DNA outputs and detecting small errors. Nanopore sequencing allows long reads and real-time sequencing, which is useful for quickly decoding DNA circuits or recording information from living cells. Input would be synthetic DNA strands or DNA extracted from engineered cells.
Preparation steps:
- DNA extraction
- Fragmentation (if needed)
- Adapter ligation
- PCR amplification (for Illumina)
- Loading onto sequencing platform
Illumina sequencing uses fluorescently labeled nucleotides. As each base is incorporated during synthesis, a fluorescent signal is detected and recorded to determine the sequence. Nanopore sequencing detects changes in electrical current as DNA passes through a nanopore, and software converts those signals into base calls.
The output is digital sequence data (FASTQ files) that can be decoded into stored information or analyzed to determine how the DNA circuit functioned.
5.2 DNA Write
(i) What DNA would you want to synthesize (e.g., write) and why?
I would synthesize DNA sequences that function as biological computing circuits. These sequences could encode logic operations, data storage elements, or sensing modules that respond to cellular signals. For example, a DNA circuit could detect inflammation markers in a patient and record that information into DNA for later sequencing.
Such systems could eventually be used in personalized medicine, where engineered cells monitor patient health and record molecular events. The synthesized DNA could include logic gates, memory elements, and protein-coding sequences that allow cells to process information and respond to disease states.
(ii) What DNA synthesis technology would you use and why?
I would use phosphoramidite-based oligonucleotide synthesis followed by DNA assembly methods such as Gibson Assembly or Golden Gate assembly.
Essential steps
- Chemical synthesis of short DNA oligos
- Purification
- Assembly into longer constructs
- Cloning into plasmids
- Sequence verification
Limitations
- Errors increase with longer DNA
- Cost of long constructs
- Some sequences are difficult to synthesize
- Assembly required for long circuits
Despite these limitations, modern DNA synthesis allows rapid construction of custom genetic circuits for research and therapeutic applications.
DNA Edit
(i) What DNA would you want to edit and why?
I would want to edit DNA in engineered cells used for biological computing and personalized medicine. For example, cells could be edited to contain DNA circuits that sense disease markers, process information, and respond by producing therapeutic molecules. Editing could also be used to improve stability and accuracy of DNA-based memory systems. In medicine, editing patient-derived cells could allow personalized treatments tailored to an individual’s genetic and molecular profile.
(ii) What editing technology would you use and why?
I would use CRISPR-Cas9 and base editing technologies to make precise modifications to DNA. A guide RNA directs the Cas9 enzyme to a specific DNA sequence. Cas9 creates a cut at that location, and the cell’s repair machinery introduces the desired edit using a provided template or repair process.