Week 2 HW: DNA Read, Write, & Edit


Part 1 – Benchling & In-silico Gel Art
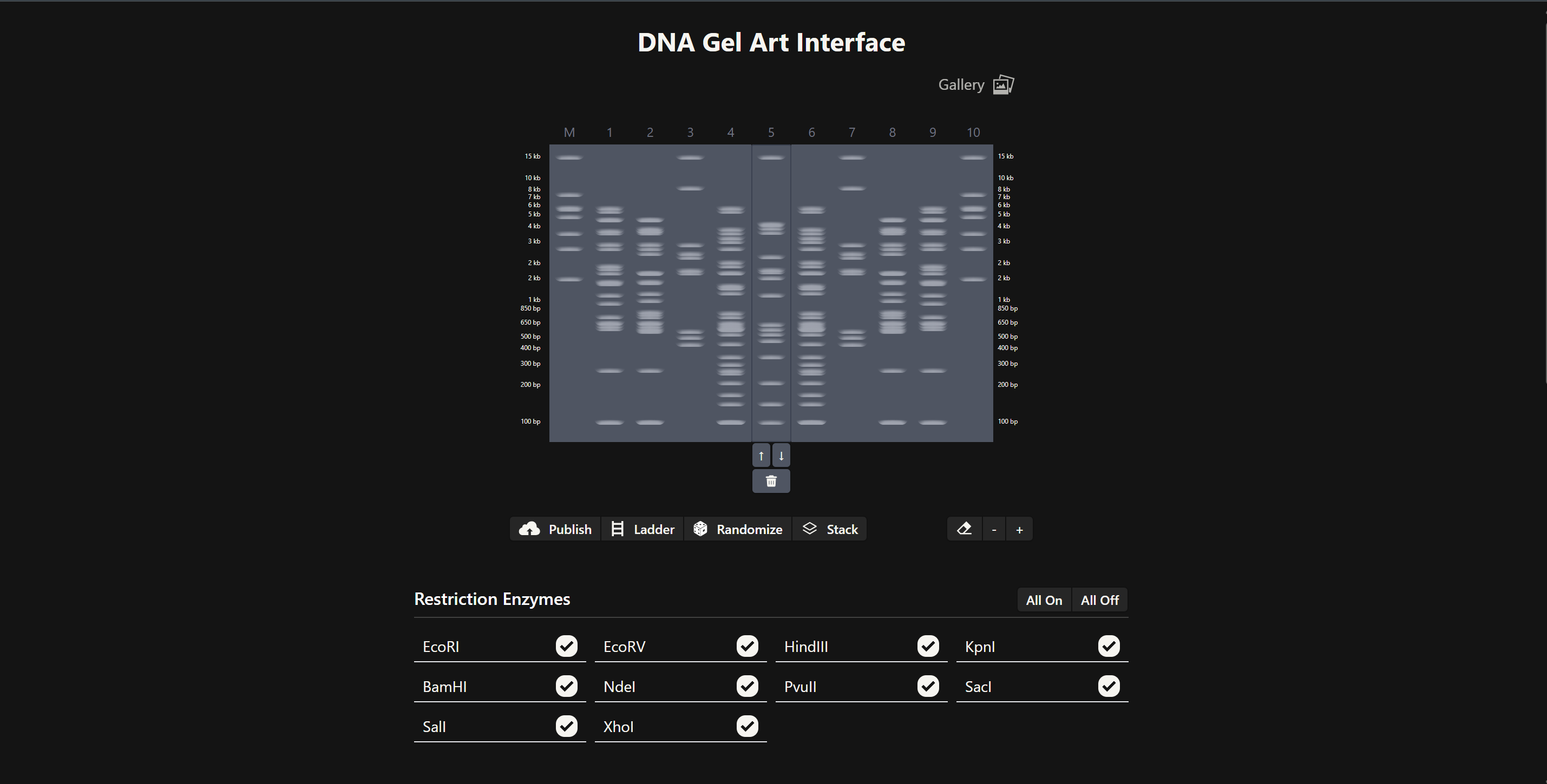
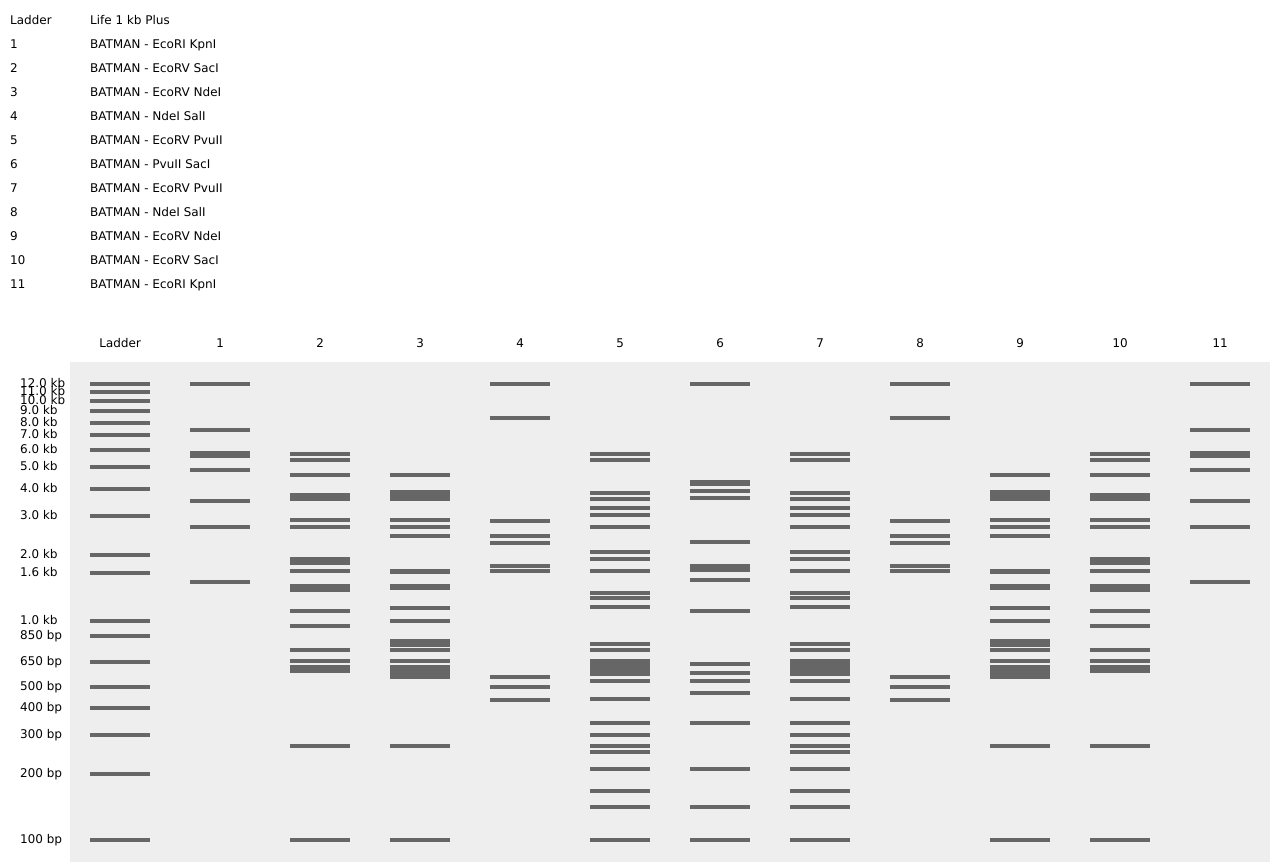
I used Benchling to design an in‑silico restriction digest of Lambda DNA. In Benchling, I created a customized restriction enzyme list for smoother later operations that included all the enzymes provided in the Week 2 HTGAA homework
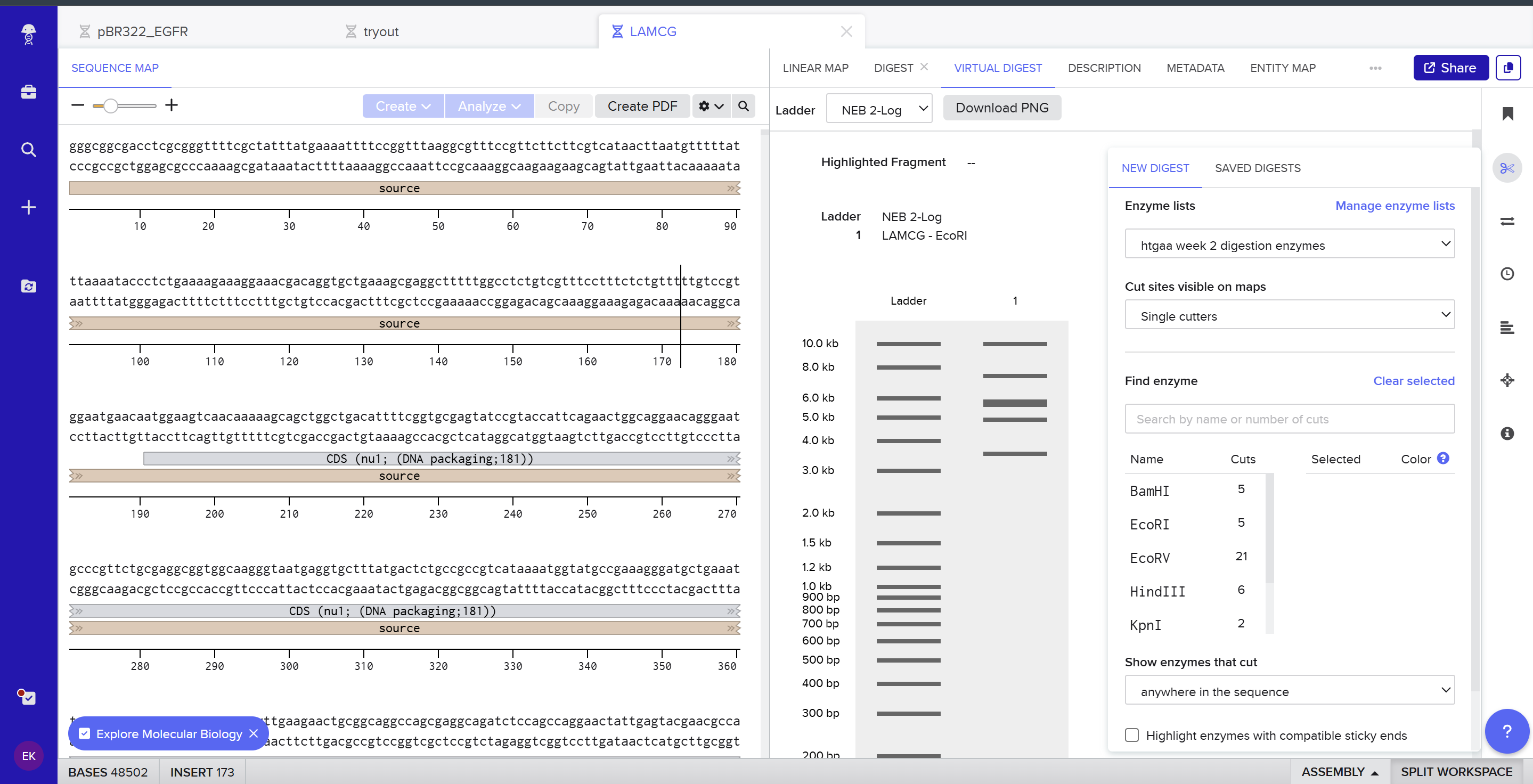

Using Ronan’s website, I tried to create a “Bat signal” 🦇 pattern on the gel (hopefully you can see my vision too!)
![]()

This was my first attempt, where the lanes did not appear in the order I expected, so the pattern looked wrong…
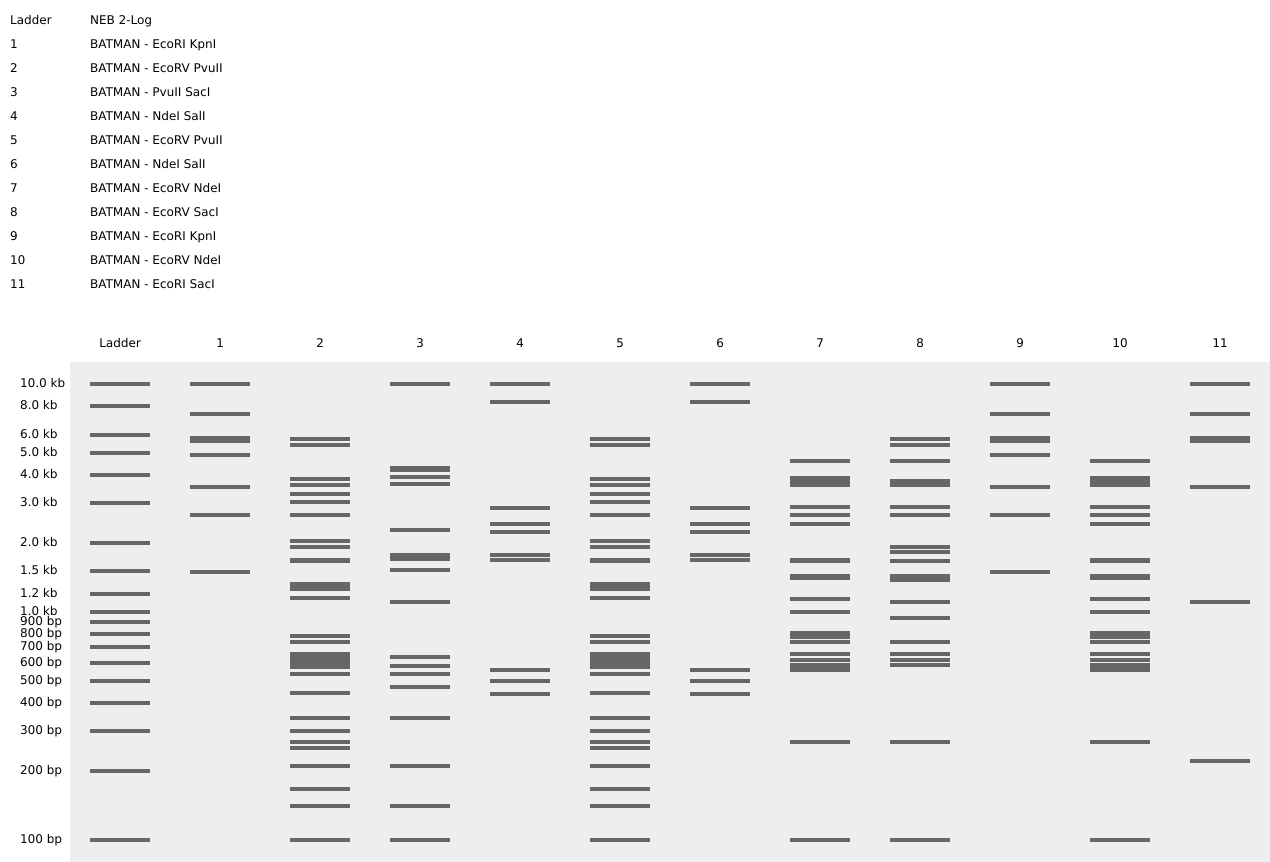

To fix this, I renamed each “Digest” tab with numbers, because every new digest was appearing in a random order.
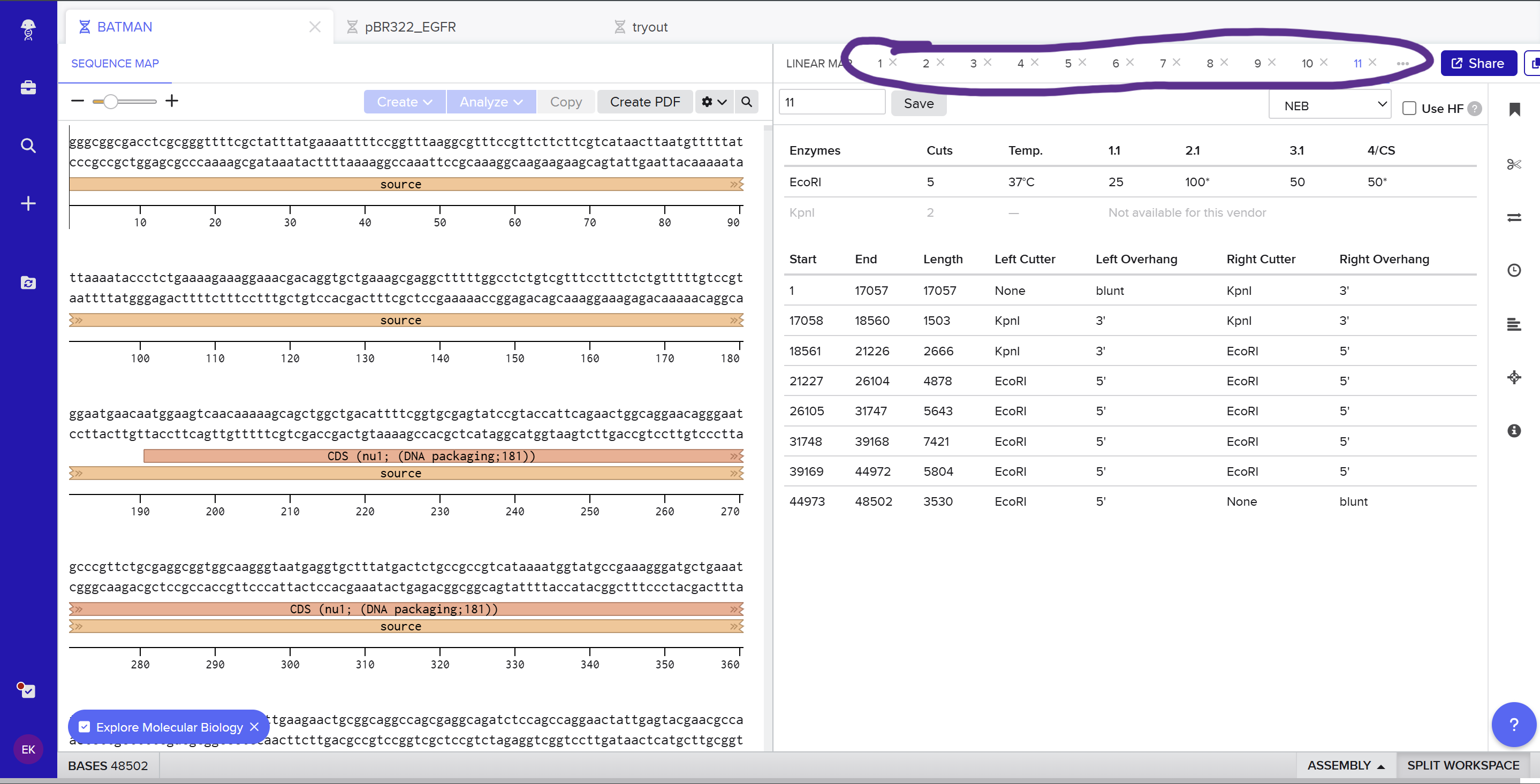

After running all the digests and then ordering the numbered lanes correctly, I finally obtained my intended DNA gel “Batman” pattern.

Part 3 - DNA Design Challenge
Protein – TRPV1 (heat and “spicy” pain sensation)
- cation channel expressed in nociceptive sensory neurons, where it detects noxious heat, low pH, and capsaicin (main compound in chili peppers) 🌶️. I chose TRPV1 because it directly links physical stimuli at the skin (heat or spicy chemicals) to electrical activity in pain pathways, making it a clear molecular mediator of sensory perception. Engineering the DNA sequence that encodes TRPV1 could tune its expression or gating properties, which is relevant for altering thermal pain sensitivity or designing cells that report damaging levels of heat.
Sequence from UniProt
sp|Q8NER1|TRPV1_HUMAN Transient receptor potential cation channel subfamily V member 1 OS=Homo sapiens OX=9606 GN=TRPV1 PE=1 SV=2 MKKWSSTDLGAAADPLQKDTCPDPLDGDPNSRPPPAKPQLSTAKSRTRLFGKGDSEEAFP VDCPHEEGELDSCPTITVSPVITIQRPGDGPTGARLLSQDSVAASTEKTLRLYDRRSIFE AVAQNNCQDLESLLLFLQKSKKHLTDNEFKDPETGKTCLLKAMLNLHDGQNTTIPLLLEI ARQTDSLKELVNASYTDSYYKGQTALHIAIERRNMALVTLLVENGADVQAAAHGDFFKKT KGRPGFYFGELPLSLAACTNQLGIVKFLLQNSWQTADISARDSVGNTVLHALVEVADNTA DNTKFVTSMYNEILMLGAKLHPTLKLEELTNKKGMTPLALAAGTGKIGVLAYILQREIQE PECRHLSRKFTEWAYGPVHSSLYDLSCIDTCEKNSVLEVIAYSSSETPNRHDMLLVEPLN RLLQDKWDRFVKRIFYFNFLVYCLYMIIFTMAAYYRPVDGLPPFKMEKTGDYFRVTGEIL SVLGGVYFFFRGIQYFLQRRPSMKTLFVDSYSEMLFFLQSLFMLATVVLYFSHLKEYVAS MVFSLALGWTNMLYYTRGFQQMGIYAVMIEKMILRDLCRFMFVYIVFLFGFSTAVVTLIE DGKNDSLPSESTSHRWRGPACRPPDSSYNSLYSTCLELFKFTIGMGDLEFTENYDFKAVF IILLLAYVILTYILLLNMLIALMGETVNKIAQESKNIWKLQRAITILDTEKSFLKCMRKA FRSGKLLQVGYTPDGKDDYRWCFRVDEVNWTTWNTNVGIINEDPGNCEGVKRTLSFSLRS SRVSGRHWKNFALVPLLREASARDRQSAQPEEVYLRQFSGSLKPEDAEVFKSPAASGEK
Reverse translated DNA sequence
atgaaraartggwsnwsnacngayytnggngcngcngcngayccnytncaraargayacn tgyccngayccnytngayggngayccnaaywsnmgnccnccnccngcnaarccncarytn wsnacngcnaarwsnmgnacnmgnytnttyggnaarggngaywsngargargcnttyccn gtngaytgyccncaygargarggngarytngaywsntgyccnacnathacngtnwsnccn gtnathacnathcarmgnccnggngayggnccnacnggngcnmgnytnytnwsncargay wsngtngcngcnwsnacngaraaracnytnmgnytntaygaymgnmgnwsnathttygar gcngtngcncaraayaaytgycargayytngarwsnytnytnytnttyytncaraarwsn aaraarcayytnacngayaaygarttyaargayccngaracnggnaaracntgyytnytn aargcnatgytnaayytncaygayggncaraayacnacnathccnytnytnytngarath gcnmgncaracngaywsnytnaargarytngtnaaygcnwsntayacngaywsntaytay aarggncaracngcnytncayathgcnathgarmgnmgnaayatggcnytngtnacnytn ytngtngaraayggngcngaygtncargcngcngcncayggngayttyttyaaraaracn aarggnmgnccnggnttytayttyggngarytnccnytnwsnytngcngcntgyacnaay carytnggnathgtnaarttyytnytncaraaywsntggcaracngcngayathwsngcn mgngaywsngtnggnaayacngtnytncaygcnytngtngargtngcngayaayacngcn gayaayacnaarttygtnacnwsnatgtayaaygarathytnatgytnggngcnaarytn cayccnacnytnaarytngargarytnacnaayaaraarggnatgacnccnytngcnytn gcngcnggnacnggnaarathggngtnytngcntayathytncarmgngarathcargar ccngartgymgncayytnwsnmgnaarttyacngartgggcntayggnccngtncaywsn wsnytntaygayytnwsntgyathgayacntgygaraaraaywsngtnytngargtnath gcntaywsnwsnwsngaracnccnaaymgncaygayatgytnytngtngarccnytnaay mgnytnytncargayaartgggaymgnttygtnaarmgnathttytayttyaayttyytn gtntaytgyytntayatgathathttyacnatggcngcntaytaymgnccngtngayggn ytnccnccnttyaaratggaraaracnggngaytayttymgngtnacnggngarathytn wsngtnytnggnggngtntayttyttyttymgnggnathcartayttyytncarmgnmgn ccnwsnatgaaracnytnttygtngaywsntaywsngaratgytnttyttyytncarwsn ytnttyatgytngcnacngtngtnytntayttywsncayytnaargartaygtngcnwsn atggtnttywsnytngcnytnggntggacnaayatgytntaytayacnmgnggnttycar caratgggnathtaygcngtnatgathgaraaratgathytnmgngayytntgymgntty atgttygtntayathgtnttyytnttyggnttywsnacngcngtngtnacnytnathgar gayggnaaraaygaywsnytnccnwsngarwsnacnwsncaymgntggmgnggnccngcn tgymgnccnccngaywsnwsntayaaywsnytntaywsnacntgyytngarytnttyaar ttyacnathggnatgggngayytngarttyacngaraaytaygayttyaargcngtntty athathytnytnytngcntaygtnathytnacntayathytnytnytnaayatgytnath gcnytnatgggngaracngtnaayaarathgcncargarwsnaaraayathtggaarytn carmgngcnathacnathytngayacngaraarwsnttyytnaartgyatgmgnaargcn ttymgnwsnggnaarytnytncargtnggntayacnccngayggnaargaygaytaymgn tggtgyttymgngtngaygargtnaaytggacnacntggaayacnaaygtnggnathath aaygargayccnggnaaytgygarggngtnaarmgnacnytnwsnttywsnytnmgnwsn wsnmgngtnwsnggnmgncaytggaaraayttygcnytngtnccnytnytnmgngargcn wsngcnmgngaymgncarwsngcncarccngargargtntayytnmgncarttywsnggn wsnytnaarccngargaygcngargtnttyaarwsnccngcngcnwsnggngaraar
Codon Optimization For codon optimization, I planned to take my reverse‑translated TRPV1 coding sequence and run it through an online codon optimization tool to adapt codon usage to E. coli, replacing rare codons, adjusting GC content, and removing unwanted motifs while keeping the amino‑acid sequence unchanged. However, the TwistBioscience optimization tool was unavailable and other available web tools repeatedly failed on my long TRPV1 sequence, so for this homework I kept the reverse‑translated sequence from Part 3.2 as my working TRPV1 coding sequence and discussed codon optimization conceptually instead of providing a fully optimized sequence.
3.4: What technologies could be used to produce this protein from your DNA? Describe in your words the DNA sequence can be transcribed and translated into a protein. You may describe either cell-dependent or cell-free methods, or both. Once I have a coding DNA sequence for TRPV1, I can synthesize it and clone it into an expression plasmid with a suitable promoter, ribosome binding site, and terminator. After transforming this plasmid into host cells such as E. coli or mammalian cells, RNA polymerase transcribes the TRPV1 gene into mRNA, and ribosomes translate the mRNA into the TRPV1 channel, which is inserted into the plasma membrane and opens in response to heat or capsaicin to generate pain signals. The same DNA sequence could also be used in a cell‑free transcription–translation mix to produce TRPV1 in vitro, still following the central dogma from DNA to RNA to protein
Part 4
I created a new linear DNA sequence in Benchling named sfGFP, set the nucleotide type to DNA, and topology to Linear. In the sequence editor I pasted, in order, the example promoter BBa_J23106, RBS BBa_B0034 with spacer, start codon (ATG), the provided codon‑optimized sfGFP coding sequence, a 7×His tag at the C‑terminus, a stop codon (TAA), and the BBa_B0015 terminator, and added annotations for each feature (Promoter, RBS, sfGFP CDS, 7×His tag, Stop, Terminator). Here you can see the screenshot from Benchling showing the sequence map: (https://benchling.com/s/seq-KNkSG9FjYrEgCrgZE0Id?m=slm-aiflv0AFXb7Fro539sLk)
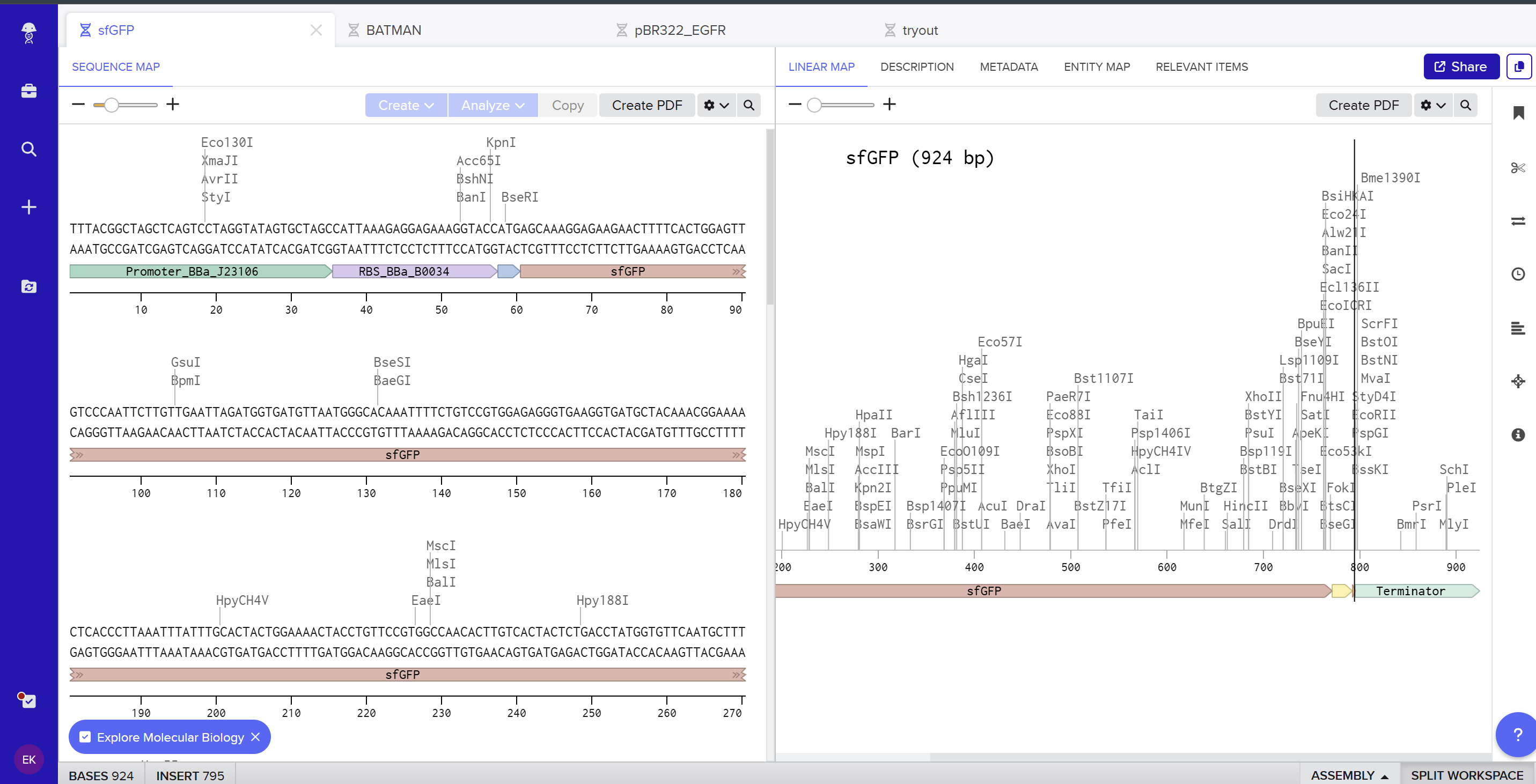

On the Twist portal I selected the “Genes” product and chose the “Clonal Genes” option, since this provides my insert in a circular plasmid that can be transformed directly into E. coli. I imported the FASTA file of my sfGFP expression cassette as a nucleotide sequence, then chose a Twist cloning vector (pTwist Amp High Copy) as the backbone so that the final construct includes an origin of replication and ampicillin resistance. After Twist generated the plasmid design, I downloaded the GenBank file and re‑imported it into Benchling to view the full plasmid map with my annotated sfGFP expression cassette inserted:
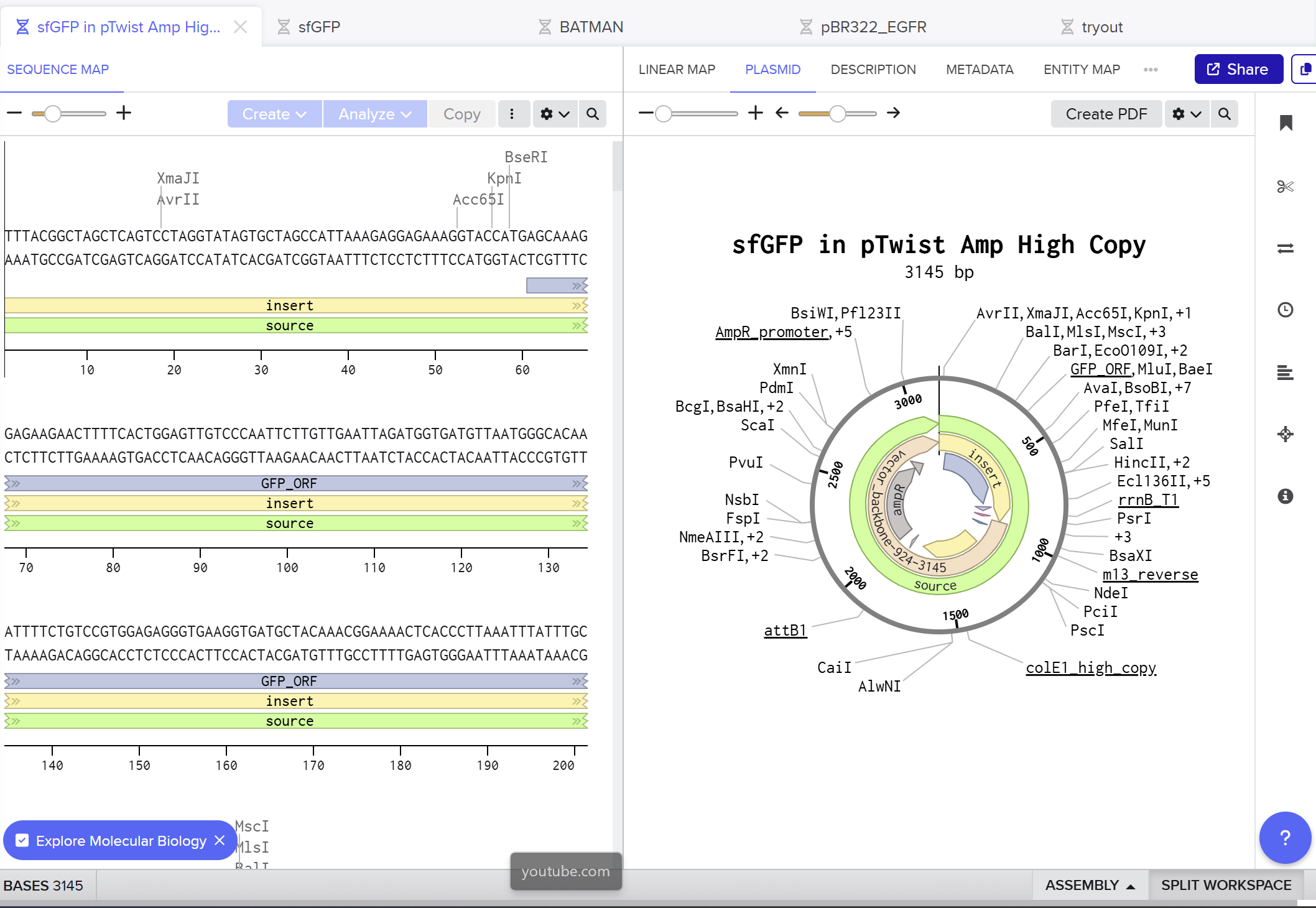

Part 5
DNA Read 📖
What DNA would you want to sequence and why?


I would like to sequence DNA from banana (Musa species) to explore how similar or different it is from the human genome, especially because of the known fun fact stating that humans “share around half their genes” with banana. By sequencing banana DNA, I would wanna compare it to human gene sets and get the idea where these similarities come from and what they lead to. 🍌
What technology would you use and why?
I would use Illumina sequencing‑by‑synthesis (second‑generation NGS), possibly complemented by nanopore (third‑generation) for long reads.
Input and prep: extract banana genomic DNA, fragment it, repair ends, ligate Illumina adapters, PCR‑amplify, then load on a flow cell
How it reads bases: clusters are formed on the flow cell. In each cycle, fluorescently labeled nucleotides are added, one base at a time, and the machine takes a picture. The color of each spot in each cycle tells you which base (A, T, C, or G) was added there.
Output: millions of short reads in FASTQ format, which can be assembled and compared to human genes
DNA Write ✍🏽
What DNA would you want to synthesize (e.g., write) and why? I would like to synthesize a genetic circuit for a “self‑adjusting” biomaterial, where cells inside a hydrogel can sense mechanical stress and then change the stiffness of the material. The idea is to have a material that becomes stiffer when it needs more support and softer when stress is too high, using gene expression instead of external tools. This could be useful for tissue engineering and mechanobiology, because many studies show that cell fate and behavior depend not only on stiffness, but also on how stiffness changes over time
What technology would you use to perform this DNA synthesis and why? To build this circuit, I would use chip‑based DNA oligo synthesis plus clonal gene synthesis, and then assemble the parts into an expression cassette. Chip‑based synthesis is good for designing and producing many regulatory variants (different mechanosensitive promoters, crosslinker genes, degradation domains) in parallel, which is important when tuning a dynamic material
Essential steps
- Design the circuit in silico: pick mechanosensitive promoter elements, choose coding sequences for matrix‑building proteins and matrix‑remodeling enzymes, then add RBSs and terminators
- Order synthetic DNA fragments or full clonal genes from a synthesis provider, using chip‑based oligo synthesis to keep costs down for complex designs.
- Assemble the fragments into plasmids, transform them into the chosen cell chassis, and verify by sequencing
Limitations
- Complex construction can have a high error rate
- Synthesis and clonign might take several days to weeks
- Mechanosensitive elements characterized in 2D cultures may behave differently in 3D hydrogels
DNA Edit 🖆

What DNA would you want to edit and why?
I would like to edit DNA in cartilage‑related cells for athletes. The example would be figure skaters who often perform repeated high jumps and landings that produce a very high impact on the knee and ankle. Most figure skaters frequently develop overuse injuries and early degenerative changes in the ankle/knee joints. This leads to the early retirement of athletes in their early teens and extensive health problems.
Editing joint cartilage cells to be more regenerative, so that damaged cartilage can be repaired more effectively over time.
The target gene would be SOX9 and TGF-Beta pathway genes, since they are known to be the main pro-generative genes in cartilage.
The reason why I wouldn’t want to explicitly target genes related to the defensive functions of cartilage to prevent injuries is that it would raise some ethical concerns.
What technology or technologies would you use to perform these DNA edits and why? I would use CRISPR-based gene activation in joint-derived stem cells to upregulate SOX9 and TGF-Beta pathways genes. This technology would guide RNAs targeting promoters to boost cells’ own existing genes without cutting DNA. This would explicitly focus on existing injuries.
Essential steps
- Confirm that SOX9 and key TGF genes are pro-generative in articular cartilage and design guide RNAs that bind promoter regions of SOX9 adn TGFB-pathways genes in human joint cells
- Build dCas9-activator plasmids for designed gRNAs
- Deliver dCas9-activator and gRNA to the cell
- Culture and differentiate edited cells towards cartilage
Preparation and inputs
- Extensive research and selection of targeted genes and regulatory regions in human joint cartilage
- design of guide RNA
- selection of dCas9-activator
- Inputs: DNA templates, plasmids, viral vectors encoding dCas9-activator, plasmids for gRNAs, patient derived MSCs cells
Limitations
- Since dCas9 does not cut DNA, there is a possibility of upregulation of unintended genes, because of the off-target binding
- There should be controlled upregulation, since over-activation of these genes can lead to fibrosis or abnormal tissue growth