Week 6 HW: Genetic Circuits Part I: Assembly Technologies
DNA Assembly
What are some components in the Phusion High-Fidelity PCR Master Mix and what is their purpose?
Phusion DNA Polymerase*
- Chimeric enzyme that catalyzes the synthesis of ew DNA strand in the 5 -> 3 direction with high-fidelity
dNTPs
- four chemical building blocks ($dATP, dTTP, dCTP, dGTP$) used to construct the DNA. They provide both the physical material and the energy required for the polymerase to grow the new strand
Reaction Buffer
- Maintains the optimal pH and ionic environment for the reaction. It ensures the enzyme remains stable and functional throughout the high-temperature cycles of PCR.
Magnesium Chloride
- Co-factor for the polymerase enzyme. Without magnesium ions, the enzyme cannot catalyze the chemical reaction needed to link the DNA building blocks together
Additives & Stabilizers
- Chemicals like glycerol or detergents protect the enzyme from degradation. Their purpose is to keep the master mix stable during storage and prevent the proteins from sticking to the plastic tube walls.
What are some factors that determine primer annealing temperature during PCR?
The primer annealing temperature ensures primers stick specifically to the target DNA. Main factors are:
- Primer Length and Composition: ratio of G-C to A-T bases (G-C pairs have three hydrogen bonds)
- Primer Concentración: higher concentration can increase binding
- Salt concentration: cations like K+ and Mg2+ stabilize the DNA backbone, which increases the melting temperature
- Base Mismatches
There are two methods from this class that create linear fragments of DNA: PCR, and restriction enzyme digests. Compare and contrast these two methods, both in terms of protocol as well as when one may be preferable to use over the other.
Both PCR and Restriction Enzyme Digests are fundamental techniques for generating DNA fragments; they differ in the process of generating these fragments PCR
- requires a template of DNA
- uses heat and a polymerase to synthesize new copies of a specific region
- main components are primers, dNTPs, DNA polymerase, and thermal cycler
- depend on the designed primer
Use when we have a low DNA sample volume/ we want to create a fragment of a very specific, non-common length
Restriction Enzyme Digest
- requires a high concentration of purified DNA
- uses molecular scissors to physically cut existing DNA
- main components are restriction enzymes and a stable heat incubator
- depend on the presence of specific recognition sequences (sites)
Use when we want to cut out a gene or insert from a circular plasmid to move to another vector/we want to check a piece of DNA (Diagnostic Digest)/when we know the restriction sites already exist
How can you ensure that the DNA sequences that you have digested and PCR-ed will be appropriate for Gibson cloning?
To ensure DNA fragments are ready for Gibson Assembly, we have to focus on the end of the sequences since Gibson uses overlapping DNA sequences, not the ‘sticky ends’from restriction enzymes.
- Check for overlapping ends - each fragment must share an identical sequence with the fragment next to it.
- Verify Clean Ends - since Gibson relies on Exonuclease, we must ensure there are no extra A overhangs or that enzymes have reached complete digestion.
- Check the chemical environment - must remove polymerase, dNTPs, and salts from the PCR reaction
- Sequence accuracy - verify the final assembled plasmid with the Sanger Sequencing kit at the junction points
How does the plasmid DNA enter the E. coli cells during transformation?
The process is called Transformation. Steps:
- Preparation - before DNA enters, the cells are soaked in a solution of calcium chloride, so that Ca+ would neutralize the negative charge of the DNA and the cell membrane, allowing them to get close to each other.
- Entry point = SHOCK - once mixed, the cells with DNA are moved to a 42 C water bath for 30-60 seconds to create a temporary “pressure difference” and physical holes in the cell membrane. Plasmid DNA is sweeped into these pores
- Recovery - put the cells back on ice to seal holes.
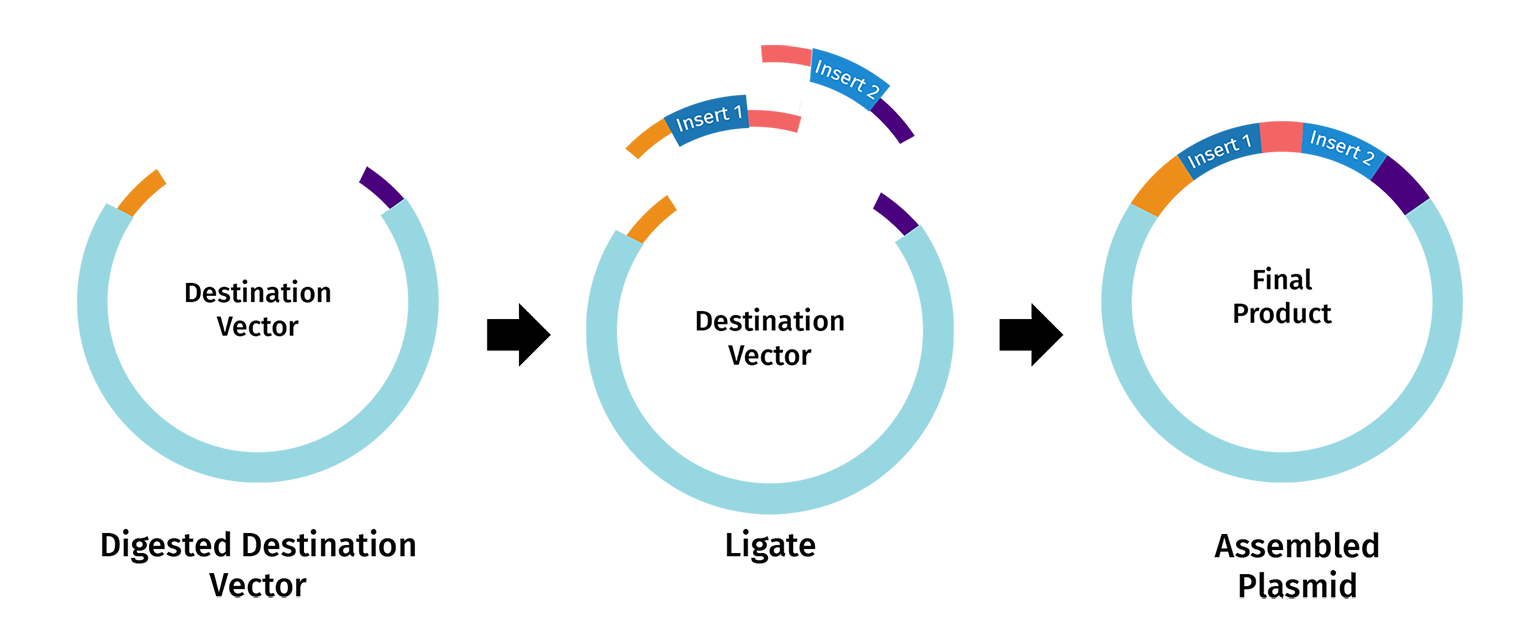
Describe another assembly method in detail (such as Golden Gate Assembly)
Golden Gate Assembly is a molecular cloning method that allows for the simultaneous, “one-pot” assembly of multiple DNA fragments using Type IIS restriction enzymes and T4 DNA ligase. Unlike standard enzymes, Type IIS enzymes (like BsaI) cut outside of their recognition sites, creating unique 4-base overhangs that can be customized to dictate the assembly order. Because the recognition sites are placed at the very ends of the fragments and are “cut off” during the reaction, the final product is seamless and lacks the original restriction sites, preventing the enzyme from re-cutting the finished plasmid. This “scarless” assembly is highly efficient, often reaching nearly 100% accuracy even when joining ten or more fragments at once. The entire process occurs in a single tube through a series of temperature cycles that alternate between the optimal conditions for digestion and ligation.