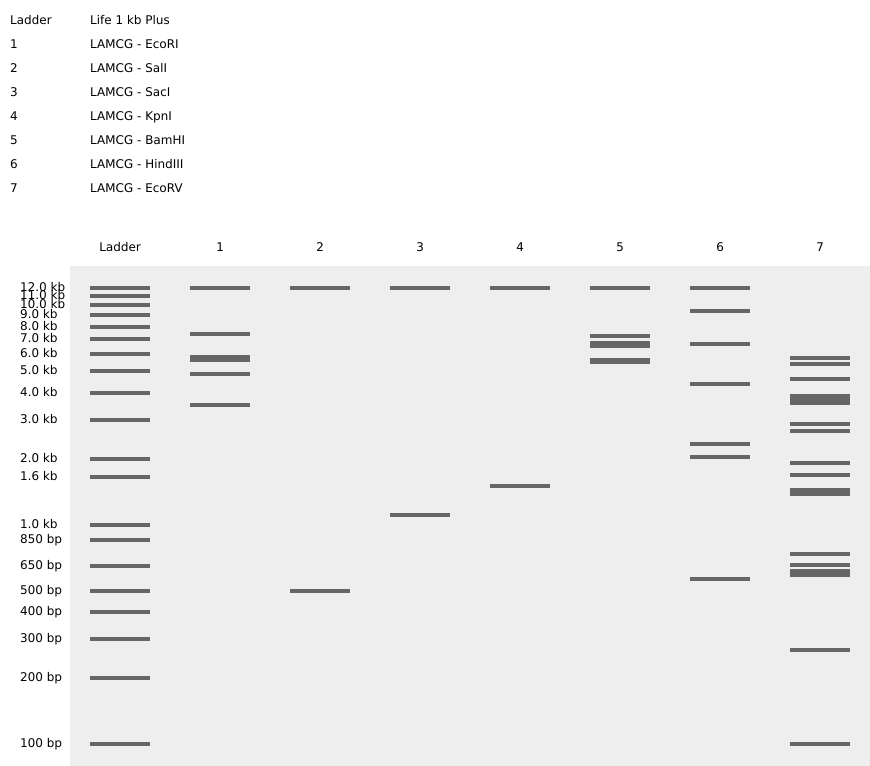
First, describe a biological engineering application or tool you want to develop and why.
Currently one area of interest is the development of a biotechnological tool capable of inhibiting and remediating toxic mould species commonly found in damp and poorly ventilated residential environments. In England alone, it is estimated that between 962,000 and 6.5 million households experience damp and mould, with up to 58% of tenants in the private rented sector affected (BRE, 2021). The prevalence is particularly acute in London, where housing commodification has intensified overcrowding and substandard living conditions. The populations most impacted include low-income households, residents of social housing, migrants, and racialized communities. Exposure to indoor mould is strongly associated with respiratory conditions, asthma exacerbation, allergic reactions, and increased vulnerability among infants, children, the elderly, and individuals with pre-existing respiratory diseases. The National Health Service (NHS) in England is estimated to spend approximately £1.4 billion annually on treating illnesses linked to cold and damp housing conditions (BRE, 2021).
Current market remediation strategies are limited in effectiveness and safety. Chemical agents such as sodium hypochlorite (bleach), ammonium compounds, and hydrogen peroxide are widely used but pose risks to human health, indoor air quality, and building materials. Conversely, domestic remedies such as vinegar or sodium bicarbonate often lack sufficient antifungal efficacy, particularly against toxigenic species such as Stachybotrys chartarum and Aspergillus spp.
This project proposes the development of a bioengineered antifungal building material or coating capable of both remediating existing mould infestations and preventing future colonization, while simultaneously restoring structural integrity through biologically driven repair mechanisms. It could look at combining various enzymes and organisms such as chitinases derived from fungal antagonists and urease producing bacteria used in self-healing concrete or fibre forming bacterial cellulose for self-repair after mould damage.
Next, describe one or more governance/policy goals related to ensuring that this application or tool contributes to an “ethical” future, like ensuring non-malfeasance (preventing harm). Break big goals down into two or more specific sub-goals.
Goal 1: Ensuring non-malfeasance
- Use products that have a low risk of virulence and infection
- toxicological safety assurance - require comprehensive testing
Goal 2: Ensure the technology reduces environmental harm
- Create clear policies for managing waste byproducts safely
- Consideration of biodegradability of materials and assess life-cycles
Goal 3: Prevent dual-use misuse and malicious applications
- Restrict access to strains and enzymes, limit distribution of engineered organisms
- Require transparent documentation of biological materials stored by trusted party
Describe at least three different potential governance ‘actions’ by consider the four aspects below. (Purpose, Design, Assumptions, Risks of Failure & “Success”) Try to outline a mix of actions (e.g. a new requirement/rule, incentive, or technical strategy) pursued by different “actors” (e.g. academic researchers, companies, federal regulators, law enforcement, etc). Draw upon your existing knowledge and a little additional digging, and feel free to use analogies to other domains (e.g. 3D printing, drones, financial systems, etc.).
Actor 1: Academic researchers
- Perform long-term environmental persistence studies with anti-mould/repair building materials
- Require instructions to be paired with reserach publications
- Design with use-case in mind for equitable accessibility
Actor 2: Companies
- Restrict usage such as for indoor use only
- Track product distribution
- Provide recall mechanisms
Actor 3: Governments/ Federal regulators
- Prohibit off-label indoor use
- Require post-market surveillance
- Enforce stop-use orders if unexpected effects emerge
Next, score (from 1-3 with, 1 as the best, or n/a) each of your governance actions against your rubric of policy goals. The following is one framework but feel free to make your own:
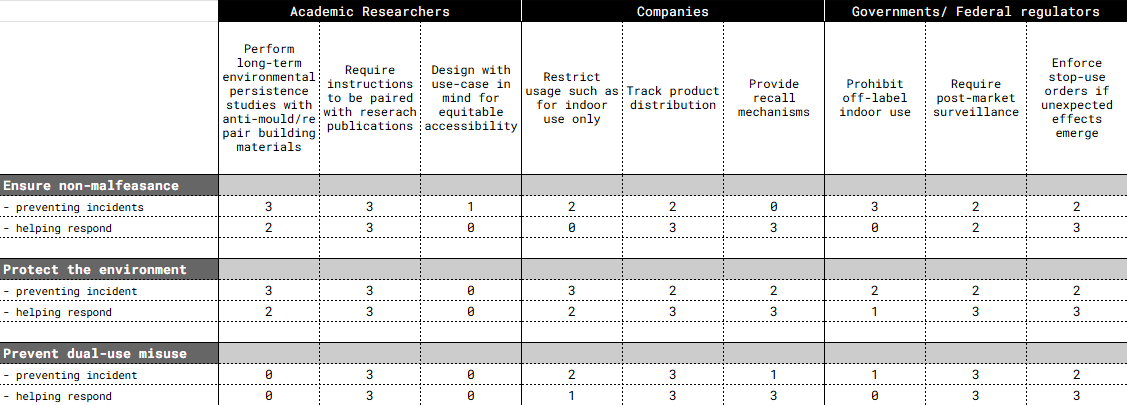

Last, drawing upon this scoring, describe which governance option, or combination of options, you would prioritize, and why. Outline any trade-offs you considered as well as assumptions and uncertainties.
I would prioritize requiring that clear, standardized usage instructions be paired with all research publications, with academic researchers as the primary responsible actors. This exercise highlighted the critical importance of accurate information dissemination in preventing misuse and enabling effective responses to potential harms, thereby supporting my governing goals related to safety, non-maleficence, and responsible deployment. However, this approach assumes that other researchers, companies, and end users will actively engage with and adhere to the guidance provided, which represents a key governance risk if compliance is low.
In addition, I would prioritize tracking product distribution by companies, as this enables restrictions on who may manufacture, sell, and apply the technology, while also establishing accountability mechanisms. Distribution tracking would allow incidents to be traced efficiently, improving response times and regulatory enforcement. Together, these measures strengthen oversight, encourage responsible use, and create a feedback loop between research, deployment, and governance.
Questions from Dr. LeProust: