Week 2 HW: DNA Read/Write and Edit
Part 1: Benchling & In-silico Gel Art
- Make free account at benchling.com
- Import the Lambda DNA
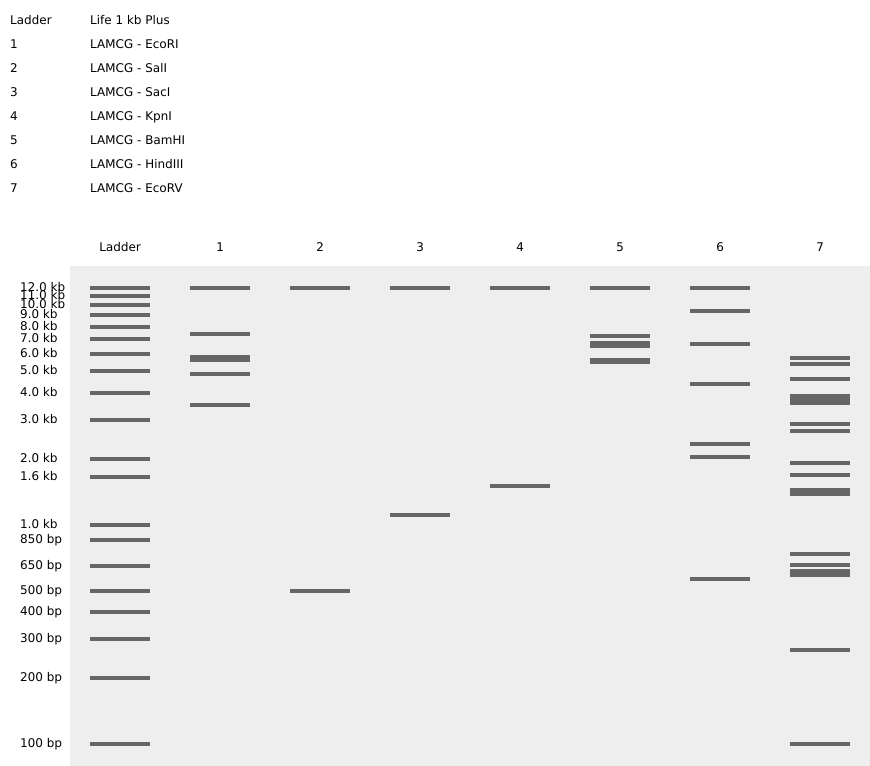
- Simulate Restriction Enzyme Digestion with the following Enzymes:
- Create a pattern/image in the style of Paul Vanouse’s Latent Figure Protocol artworks.
Part 3: DNA Design Challenge
3.1 Choose your protein
- In recitation, we discussed that you will pick a protein for your homework that you find interesting. Which protein have you chosen and why? Using one of the tools described in recitation (NCBI, UniProt, google), obtain the protein sequence for the protein you chose
3.2 Reverse Translate: Protein (amino acid) sequence to DNA (nucleotide) sequence.
- The Central Dogma discussed in class and recitation describes the process in which DNA sequence becomes transcribed and translated into protein. The Central Dogma gives us the framework to work backwards from a given protein sequence and infer the DNA sequence that the protein is derived from. Using one of the tools discussed in class, NCBI or online tools (google “reverse translation tools”), determine the nucleotide sequence that corresponds to the protein sequence you chose above.
3.3 Codon Optimization
- Once a nucleotide sequence of your protein is determined, you need to codon optimize your sequence. You may, once again, utilize google for a “codon optimization tool”. In your own words, describe why you need to optimize codon usage. Which organism have you chosen to optimize the codon sequence for and why?
3.4 You have a sequence! Now what?
- What technologies could be used to produce this protein from your DNA? Describe in your words the DNA sequence can be transcribed and translated into your protein. You may describe either cell-dependent or cell-free methods, or both.
3.5 How does it work in nature/biological systems?
- Describe how a single gene codes for multiple proteins at the transcriptional level.
- Try aligning the DNA sequence, the transcribed RNA, and also the resulting translated Protein!!! See example below.
Part 4: Prepare a Twist DNA Synthesis Order
- 4.1 Create Twist account/ Benchling account
- 4.2 Build your DNA insert Sequence
- 4.3 On Twist, Select The “Genes” Option
- 4.4. Select “Clonal Genes” option
- 4.5. Import your sequence
- 4.6. Choose Your Vector
Part 5: DNA Read/Write/Edit
5.1 DNA Read
- (i) What DNA would you want to sequence (e.g., read) and why?
- (ii) In lecture, a variety of sequencing technologies were mentioned. What technology or technologies would you use to perform sequencing on your DNA and why?
- Is your method first-, second- or third-generation or other? How so?
- What is your input? How do you prepare your input (e.g. fragmentation, adapter ligation, PCR)? List the essential steps.
- What are the essential steps of your chosen sequencing technology, how does it decode the bases of your DNA sample (base calling)?
- What is the output of your chosen sequencing technology?
5.2 DNA Write
- (i) What DNA would you want to synthesize (e.g., write) and why?
- (ii) What technology or technologies would you use to perform this DNA synthesis and why?
- What are the essential steps of your chosen sequencing methods?
- What are the limitations of your sequencing method (if any) in terms of speed, accuracy, scalability?
5.3 DNA Edit
- (i) What DNA would you want to edit and why?
- (ii) What technology or technologies would you use to perform these DNA edits and why?
- How does your technology of choice edit DNA? What are the essential steps?
- What preparation do you need to do (e.g. design steps) and what is the input (e.g. DNA template, enzymes, plasmids, primers, guides, cells) for the editing?
- What are the limitations of your editing methods (if any) in terms of efficiency or precision?