Week 4 HW: Protein Design Part I
Part A: Conceptual Questions
1. How many molecules of amino acids do you take with a piece of 500 grams of meat? (on average an amino acid is ~100 Daltons).
amino acid = ~100 Daltons meat = 25% protein (according to google between 20 to 30% depending on meat)
Daltons = g/mol
So 500g/4 is 125g of protein.
We’re going to have to use Avogadro's number (6.022 × 10²³)
Avogadro’s number helps us to convert Moles to number of molecules.
125 g of amino acids (protein) divided by 100 (g/mol) = 1.25 mol 1.25 mol x AVOGADROS = 7.5 × 10^23 molecules
So you take in approx 7.5 × 10^23 molecules of amino acid by eating 500g of meat.
2. Why do humans eat beef but do not become a cow, eat fish but do not become fish?
Humans can eat beef and fish and not turn into either a cow or fish because when we eat these animals the human digestion system breaks food down into smaller parts before reusing them. The foods break down into proteins, fats, and carbohydrates in our bodies. These nutrients are absorbed into our bloodstream and used by our bodies using human DNA – not the DNA of what was eaten. In essence: eating something doesn’t transfer its DNA to us and therefore doesn’t transfer its identity. The human body just takes the useful parts and repurposes them.
3. Why are there only 20 natural amino acids?
This question is difficult to answer. There are only 20 natural amino acids because that’s just the way life evolved. Part of the limitations are that there are 3-letter codons that code for amino acids. And as we learned, some of the amino acids have multiple codons, and some of the codons are used to signal to the ribosome to stop making the protein. So there is a limited number of combinations possible, but this number exceeds 20. So part of the explanation is just that this is how nature evolved and there wasn’t a need to evolve past this system since it worked.
4. Can you make other non-natural amino acids? Design some new amino acids.
Yes, you can definitely make non-natural amino acids. The 20 natural ones sort of developed by chance. So if you wanted you could design some new amino acids by changing whats called the R-group (or side chain) it’s the only part of the amino acid that is different in each natural amino acid. So you can add atoms that can bond to the carbon base of the amino acid, like: Bromine (-Br) or Iodine (-I). And that way you create a new non-natural amino acid!
5. Where did amino acids come from before enzymes that make them, and before life started?
Amino acids, before life started, were a product of when natural gases (like methand and hydrogen) were hit with sources of energy (like radiation from the sun). This energy was enough to rearrange the atoms in the natural gases and cause amino acids to be formed. This was proven by the Miller-Urey experiment where they triggered the creation of amino acids by zapping natural gases with electricity.
6. If you make an α-helix using D-amino acids, what handedness (right or left) would you expect?
A α-helix using D-amino acids would be left-handed. In D-amino acids, D refers to dextra which means left, which is confusing. But that refers to their position around the carbon in an amino acid. So if you flip it, you get the “handedness” So D-amino acids actually do a right-handed spiral in a shape like the alpha-helix.
7. Why do β-sheets tend to aggregate?
Beta-sheets have a flat shape with zig-zag patterns. They tend to aggregate because this shape makes them very stackable. Also the large surface area of the beta sheets are hydrophobic. So beta sheets tend to “stick” together to avoid water.
8. What is the driving force for β-sheet aggregation?
I answered this in the previous question but the fact that so many of the amino acid chains in the the beta sheet are hydrophobic means that there is a tendency for β-sheets to clump together. They also have hydrogen bonds on the side which allows them to easily attach to other beta-sheets or other proteins
9. Can you use amyloid β-sheets as materials?
Amyloid β-sheets as materials are really useful as materials due to the fact that they aggregate and strongly bind to each other. Spider silk for example contains Beta sheets and it is a very strong natural material. Beta-sheets is also used in some nanowires to conduct electricity.
Part B: Protein Analysis and Visualization
- Briefly describe the protein you selected and why you selected it.
I am choosing pilA from my week 2 homework since I find it’s electron carrying properties fascinating and would love to explore it more:
2. Identify the amino acid sequence of your protein. How long is it? What is the most frequent amino acid? You can use this Colab notebook to count the frequency of amino acids.
Here are the results from using the Colab Notework:
Total Sequence Length: 270
-----------------------------------
Amino Acid | Count | Frequency (%)
-----------------------------------
C | 75 | 27.78%
G | 73 | 27.04%
A | 66 | 24.44%
T | 56 | 20.74%
The most frequent amino acid is: Cysteine.
How many protein sequence homologs are there for your protein? Hint: Use Uniprot’s BLAST tool to search for homologs.
Does your protein belong to any protein family?
Identify the structure page of your protein in RCSB When was the structure solved? Is it a good quality structure? Good quality structure is the one with good resolution. Smaller the better (Resolution: 2.70 Å)
Are there any other molecules in the solved structure apart from protein?
Does your protein belong to any structure classification family?
Part C: Using ML-Based Protein Design Tools
I’m choosing this protein from the PDB. It’s called MtrC from Shewanella oneidensis and it transports electrons to the surface of the membrane of the conductive bacteria. https://www.rcsb.org/structure/4LM8

Protein Language Modeling
1. Deep Mutational Scans
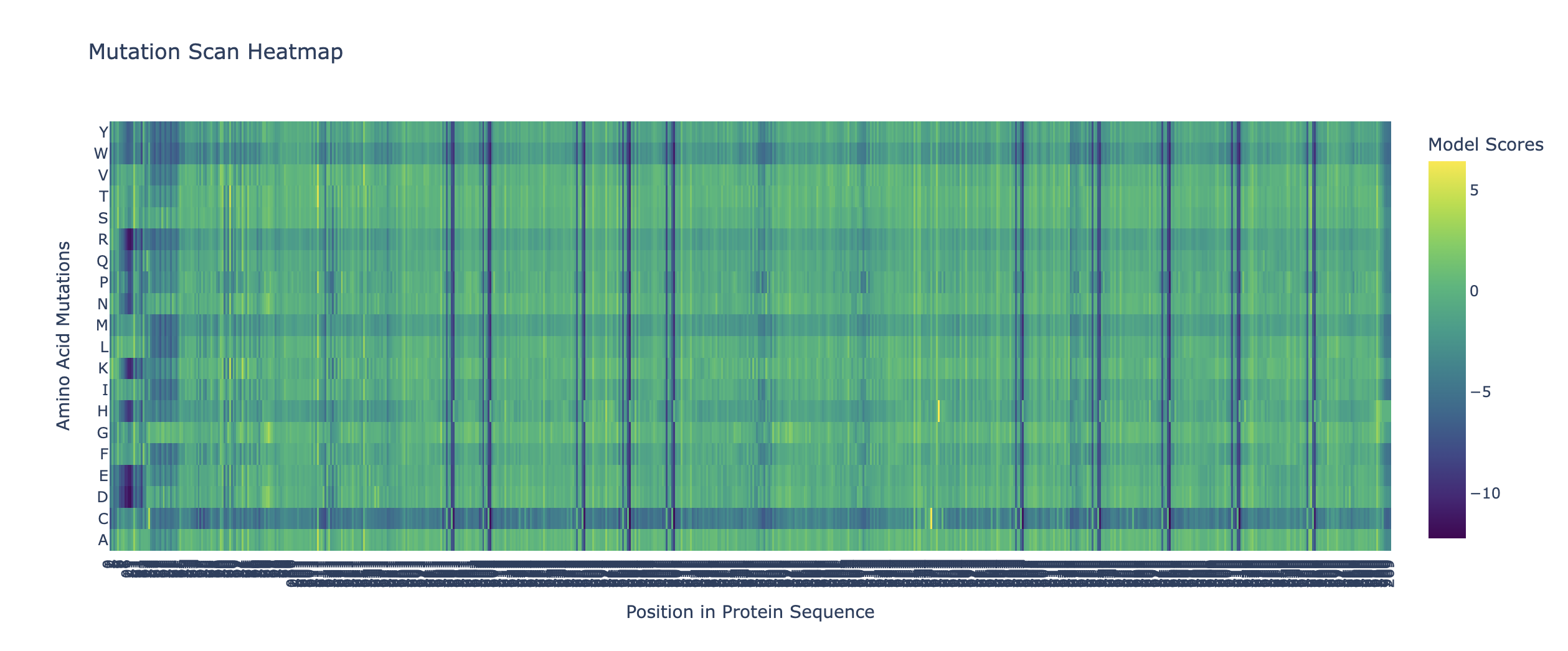
What I can tell from this heatmap is that there are multiple positions in my protein where every single mutation is harmful. These can be identified by the blue/purple vertical stripes. Interestingly there is also a horizontal blue/purple band towards the bottom. This is for the amino acid C or Cysteine. In essence this means that almost any mutation involving Cysteine would be a bad idea.
2. Latent Space Analysis
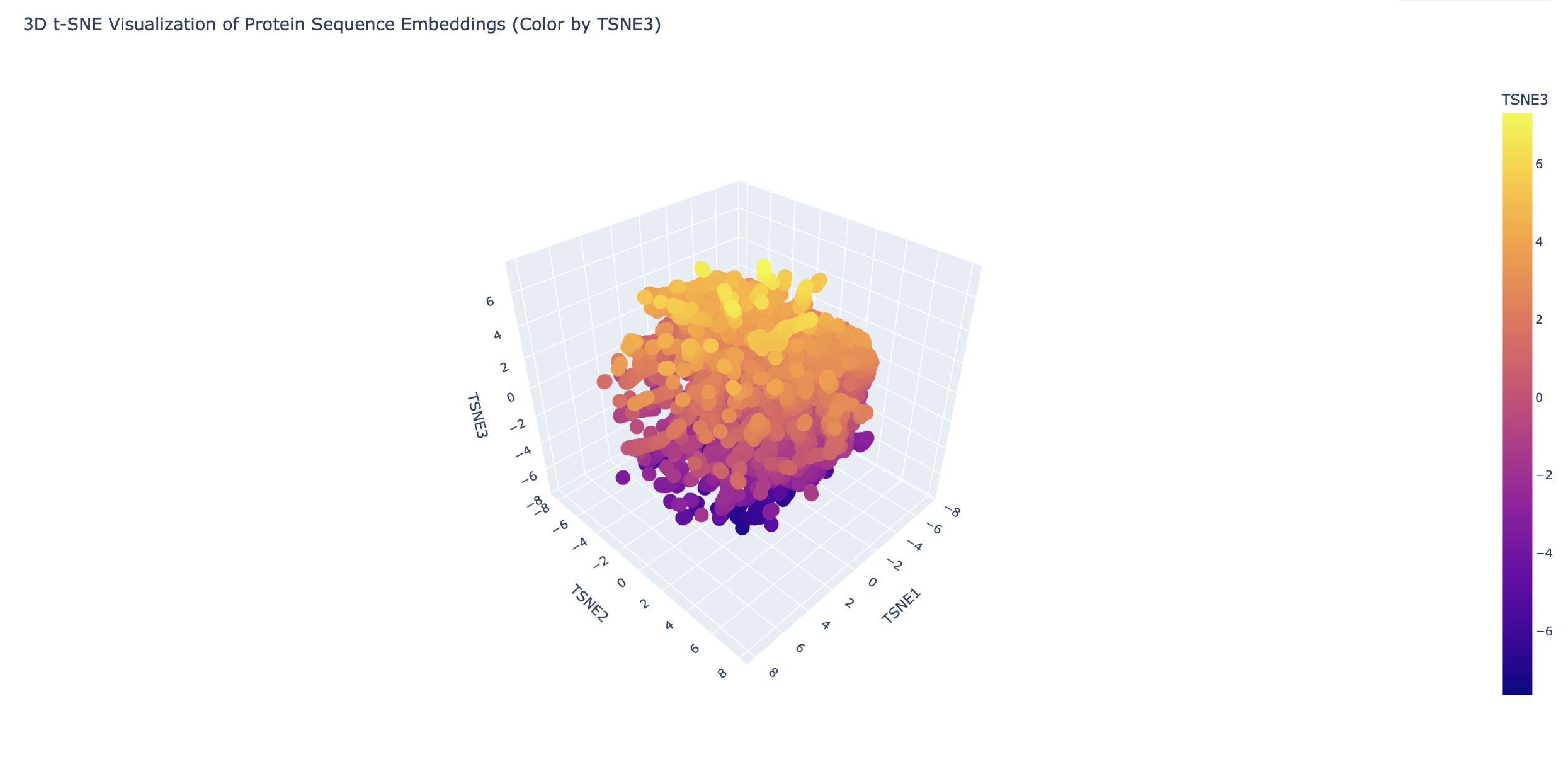
3. Protein Folding
Unfortunately I crashed my RAM trying to run ESMFold 😨. So I will revisit this homework tomorrow to complete it!