Week 6 HW: Genetic Circuits I
DNA Assembly
1. What are some components in the Phusion High-Fidelity PCR Master Mix and what is their purpose?
According to the New England Biolabs product page, Phusion High-Fidelity PCR Master Mix consists of:
- Phusion DNA Polymerase
- deoxynucleotides
- a reaction buffer that has been optimized and includes MgCl2
The Phusion DNA polymerase is able to synthesize new DNA strands. It is able to “proofread” itself which means it produces much less errors than other polymerase (hence why it is “high fidelity”). Deoxynucleotides refers to the molecules that makeup DNA (ie. A,T,G,C). These building blocks are going to be used in the new DNA strands. The reaction buffer creates the right environment for this process to take place. It ensure there is the right pH and the right amount of ions.
2. What are some factors that determine primer annealing temperature during PCR?
The factors that determine primer annealing temperature during PCR are: primer length (meaning the number of bonds - the more bonds, the higher the Tm), the amount of G-C base pairs (they form 3 hydrogen bonds as opposed to 2 which means you need a higher Tm, which means higher annealing temp). In the lab we are doing some intentional mismatches, so there is a lack of hydrogen bonds which means the strands are less “sticky” and therefore we may need a lower Tm and subsequently a lower annealing temperature. That’s another factor.
3. There are two methods from this class that create linear fragments of DNA: PCR, and restriction enzyme digests. Compare and contrast these two methods, both in terms of protocol as well as when one may be preferable to use over the other.
PCR: PCR is more of a refined process than restriction enzyme digests because you can design primers, and introduce mutations as you exponentially amplify DNA.
Restriction enzyme digests: This process is a bit more “rudimentary” than PCR because restriction enzymes can only cut DNA at certain sections. You can’t add new sequences or mutations, then enzymes just cut where they are told to. Restriction enzymes just cut existing DNA they don’t amplify anything. PCR can do it all.
PCR is a powerful process that can be used if you want to make exponential copies of the DNA and introduce mutations or add Gibson overlaps for Gibson assemply. There can be polymerase errors though. Restriction enzyme digests is great if you know exactly where you want to cut and you want to avoid polymerase errors.
4. How can you ensure that the DNA sequences that you have digested and PCR-ed will be appropriate for Gibson cloning?
To ensure DNA sequences are appropriate for Gibson cloning you have to have the Gibson overlaps I mentioned above. As mentioned in the lab the overlaps need to be “typically 20–40 bp of sequence identity between adjoining fragments” to have them connect. And these, of course, need to matcht the cloning vector. You can also check the sequences to make sure the restriction enzymes have not cut any necessary part of the DNA that will be used in the Gibson assembly, like the Gibson overlaps.
5. How does the plasmid DNA enter the E. coli cells during transformation?
The plasmid DNA can enter the E.coli using different transformation methods. The most common are heat shock and electroporation. The electric or heat shock causes the cell to “open up” and the plasmid DNA is able to enter the cell through diffusion. Once the shocks stop the E.coli cells are able to “reseal” themselves and in that way trap the plasmid DNA inside.
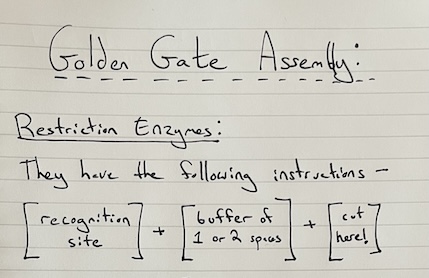
6. Describe another assembly method in detail (such as Golden Gate Assembly)
a. Explain the other method in 5 - 7 sentences plus diagrams (either handmade or online).
Golden Gate Assembly is another method that uses restriction enzymes, such as Type IIS, which cut the DNA that is next to the spot that they recognize. This creates “sticky ends” which allows DNA sequences to attach together.

For a visual:
[Type IIS site] — [4-bp overhang A] — [Gene of your Choice] — [4-bp overhang B] — [Type IIS site]
After the restriction enzymes do their job you’re left with:
[sticky end A] — [Gene of your Choice] — [sticky end B]
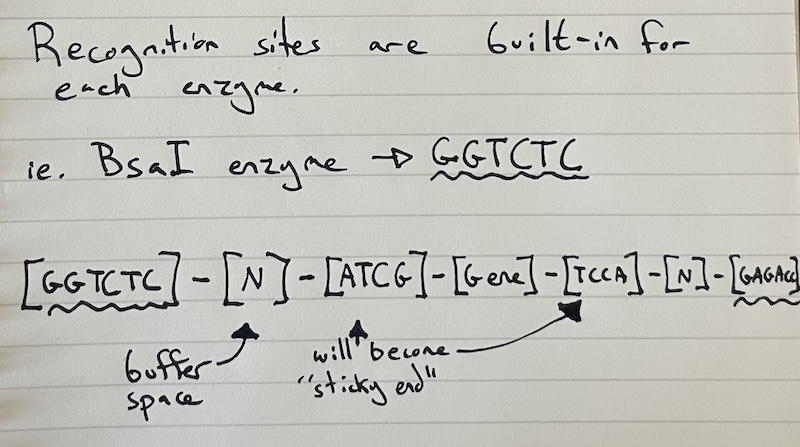
To make sure they sticky ends of your choice properly bond with each other you have to use a ligase enzyme. This isn’t related to the hydrogen bonds that keep the sticky ends together, but rather it helps to rebuild the “spine” of the DNA that was cut by the enzyme (also called the sugar phosphate backbone).
Asimov Kernel
Note: Waiting for Asimov Kernel login