Week 9 HW: Advanced Imaging & Measurement Technology
Final Project - Measurements
Please identify at least one (ideally many) aspect(s) of your project that you will measure. It could be the mass or sequence of a protein, the presence, absence, or quantity of a biomarker, etc. Please describe all of the elements you would like to measure, and furthermore describe how you will perform these measurements. What are the technologies you will use (e.g., gel electrophoresis, DNA sequencing, mass spectrometry, etc.)? Describe in detail.
I’ll need to a way to check that the Shewanella host actually carry my modified DNA. I’ll also need to measure the electron output/electrical current produced by the Shewanella (modified and unmodified). In a more basic sense, I’ll need to measure the amount of nitrate that I put in the soil for testing, and also a way to test the nitrate in the soil - to check if it correlates with the changes in electron output by the Shewanella.
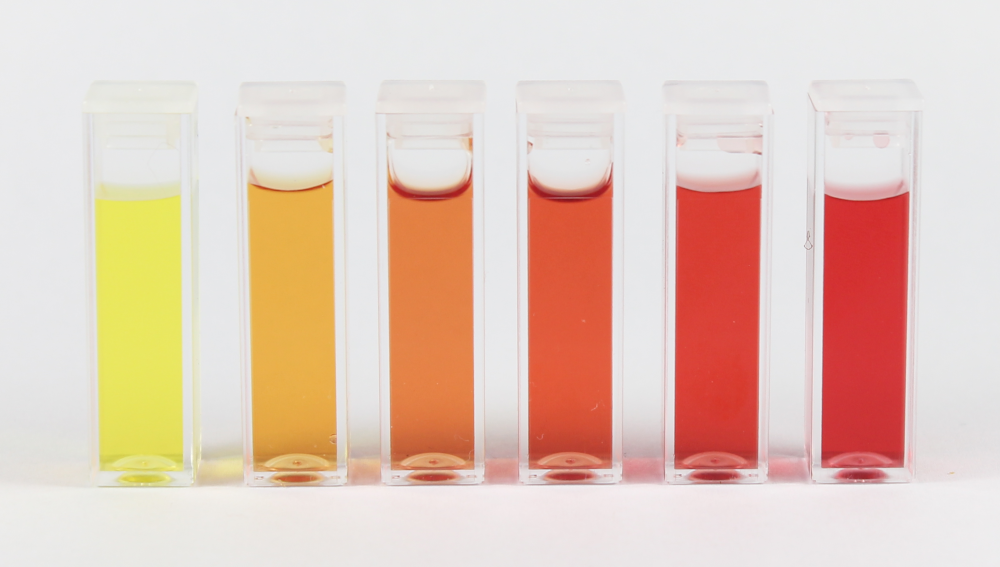
For measuring which Shewanella have been genetically modified I can use Gel Electrophoresis to make sure the DNAs are the lengths they should be, which will tell me that I have the correct DNA. I can also just sequence the DNA using Sanger sequencing to make sure manually that it is thte right DNA. For the electron current measurement I should use a potentiostat which can detect small levels of current. From that I can also calculate the power density. This should give me a good iea of what type of simple circuit could be powered by this. Measuring the nitrate can be done with color-based chemical test kits called “colorimetric nitrate assay kits” that you can easily purchase and is pictured below.


Waters Part I — Molecular Weight
1. Based on the predicted amino acid sequence of eGFP (see below) and any known modifications, what is the calculated molecular weight?
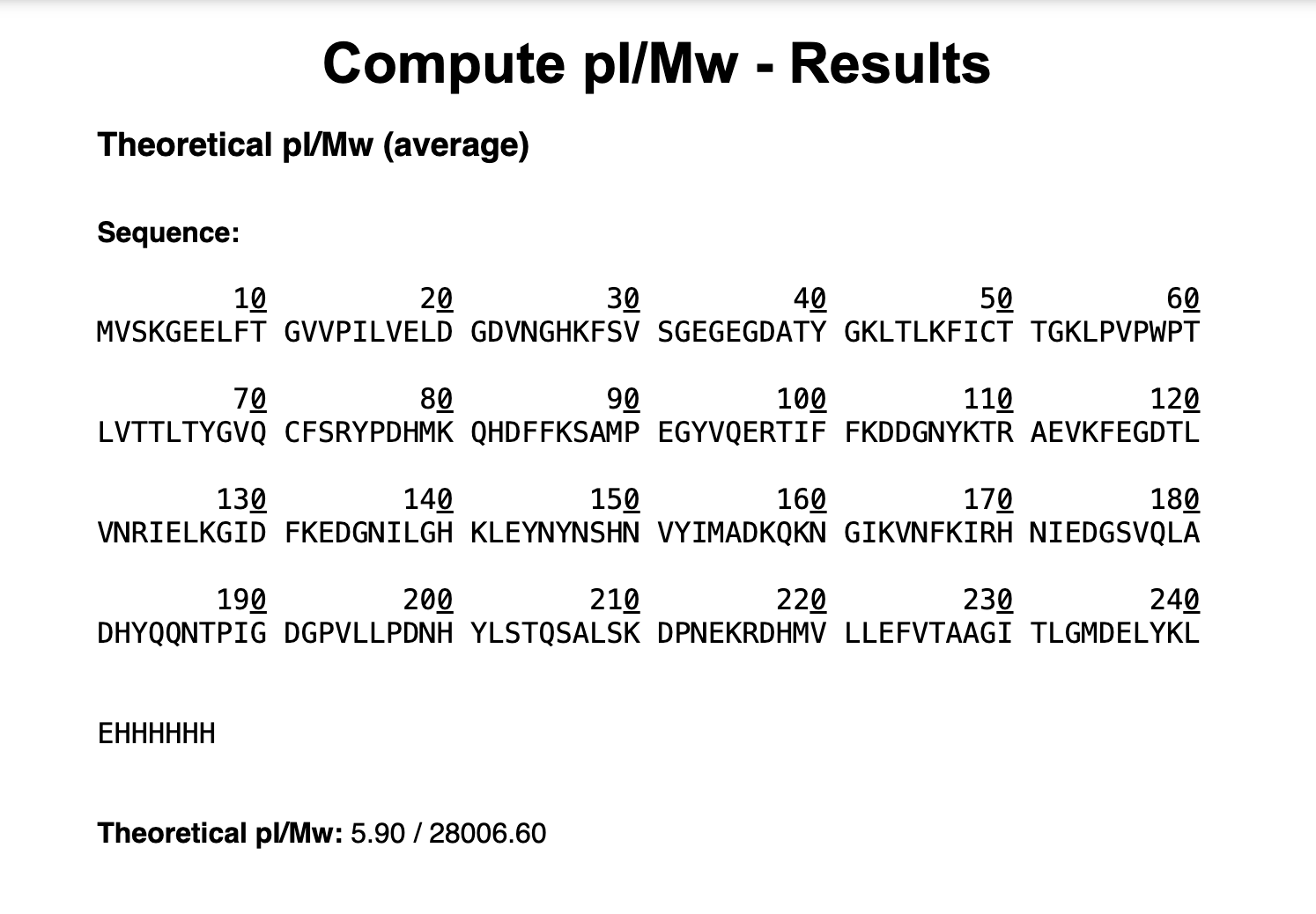
Using the online calculator the theoreatical pI (isoelectric point) is 5.90 and the Theoretical Mw (molecular weight) is 28,006.60 Da.
2. Calculate the molecular weight of the eGFP using the adjacent charge state approach described in the recitation
a. Determine z for each adjacent pair of peaks
I'm going to choose the 2 adjacent peaks:
903.7148 and 933.7349
By plugging that into the formula I get:
z = 903.7148 / (933.7349 − 903.7148)
z = 903.7148 / 30.0201
z = 30.10 ≈ 30
b. Determine the MW of the protein using the relationship between m/zn, MW, and z.
1.008 Da is the mass of a single proton (H+)
So we take this formula: m/zn = (MW + z × 1.008) / z
and then we can rearrange it to get: MW = z × (m/zn) − z × 1.008
30 x 933.7349 - 30 x 1.008MW
= 28,012.05 − 30.24MW
= 27,981.81 Da
c. Calculate the accuracy of the measurement using the deconvoluted MW from 2.2 and the predicted weight of the protein from 2.1 using the formula.
28,006.60 Da from the calculator and 27,981.81 Da from the calculations.
Accuracy = abs(27,981.81 − 28,006.60) / 28,006.60
= 24.79 / 28,006.60
= 0.00089
or 0.089% accuracy.
Waters Part III — Peptide Mapping - primary structure
1. How many Lysines (K) and Arginines (R) are in eGFP? Please circle or highlight them in the eGFP sequence given in Waters Part I question 1 above.
2. How many peptides will be generated from tryptic digestion of eGFP?
a. Navigate to https://web.expasy.org/peptide_mass/ b. Copy/paste the sequence above into the input box in the PeptideMass tool to generate expected list of peptides. c. Use Figure 4 below as a guide for the relevant parameters to predict peptides from eGFP. d. Click “Perform the Cleavage” button in the PeptideMass tool and report the number of peptides generated when using trypsin to perform the digest.
Waters Part IV — Oligomers
We will determine Keyhole Limpet Hemocyanin (KLH)’s oligomeric states using charge detection mass spectrometry (CDMS). CDMS single-particle measurements of KLH allow us to make direct mass measurements to determine what oligomeric states (that is, how many protein subunits combine) are present in solution. Using the known masses of the polypeptide subunits (Table 1) for KLH, identify where the following oligomeric species are on the spectrum shown below from the CDMS (Figure 7):
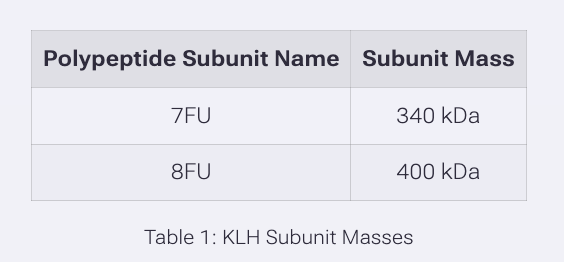
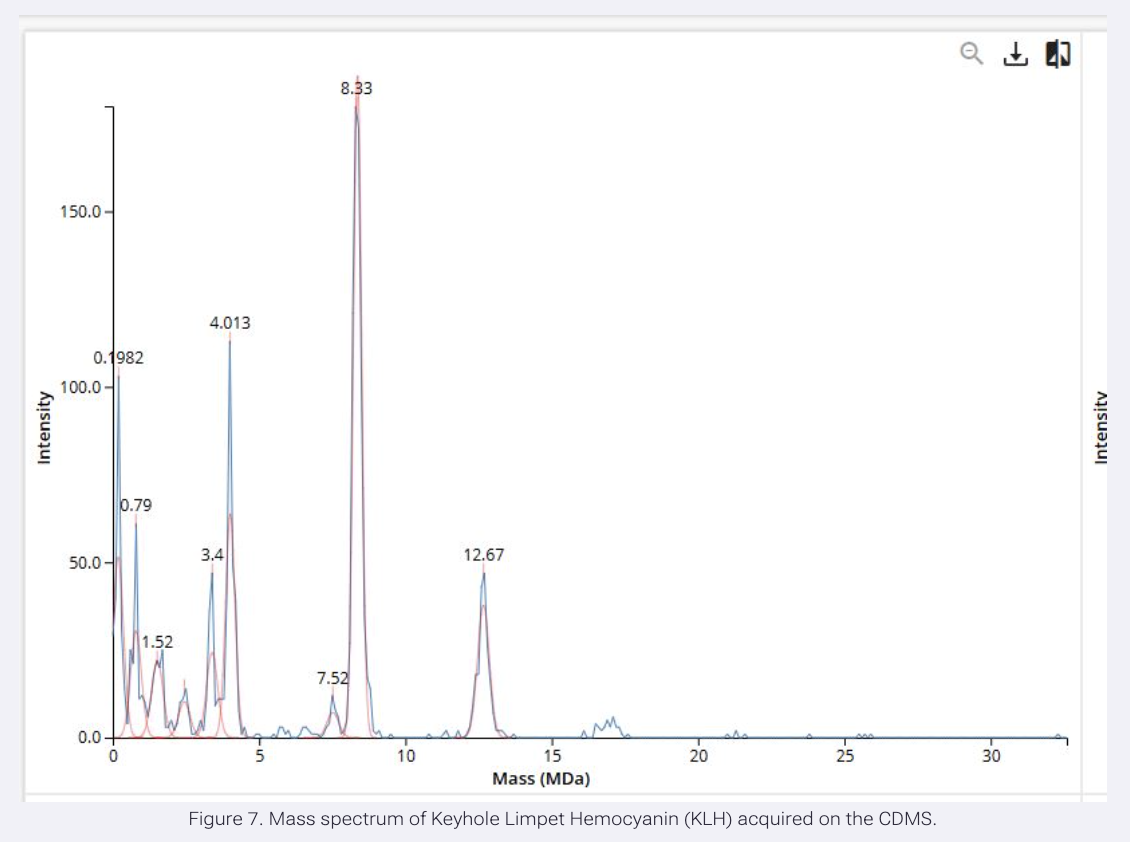
7FU Decamer
The 7FU subunit has a mass of 340 kDa Decamer refers to 10 units so you multiple 340 kDa by 10. Which is 3,400 kDa or 3.4 MDa.
So it’s at intensity 50 on the graph.
8FU Didecamer
The 8FU has a mass of 400 kDa Didecamer refers to 20 units so multiply 20 x 400kDA 20 × 400 kDa = 8,000 kDa = 8.0 MDa
Which roughly correspond to the tallest peak in the graph
8FU 3-Decamer
400 kDA x 30 = 12,000 kDa = 12.0 MDa
Which corresponds to the 12.67 peak one of the last ones on the graph.
8FU 4-Decamer
400 kDA x 40 16,000 kDa = 16.0 MDa
It’s not labeled on the graph but I’m pretty sure you can see this on the graph it’s the last visible peak.