Week 10 HW: Bioproduction & Cloud Labs
Part A: The 1,536 Pixel Artwork Canvas | Collective Artwork
So I forgot to take a photo but I submitted one green pixel that was above a red heart in the bottom left quadrant. This was early on, when I first received the link to the Pixel Artwork Canvas.
However I’m pretty sure the whole heart got erased and replaced multiple times over by the time the deadline ended! haha
Here’s the final output (maybe my pixel ended up being used as some of the green background for “Love HTGAA”?):
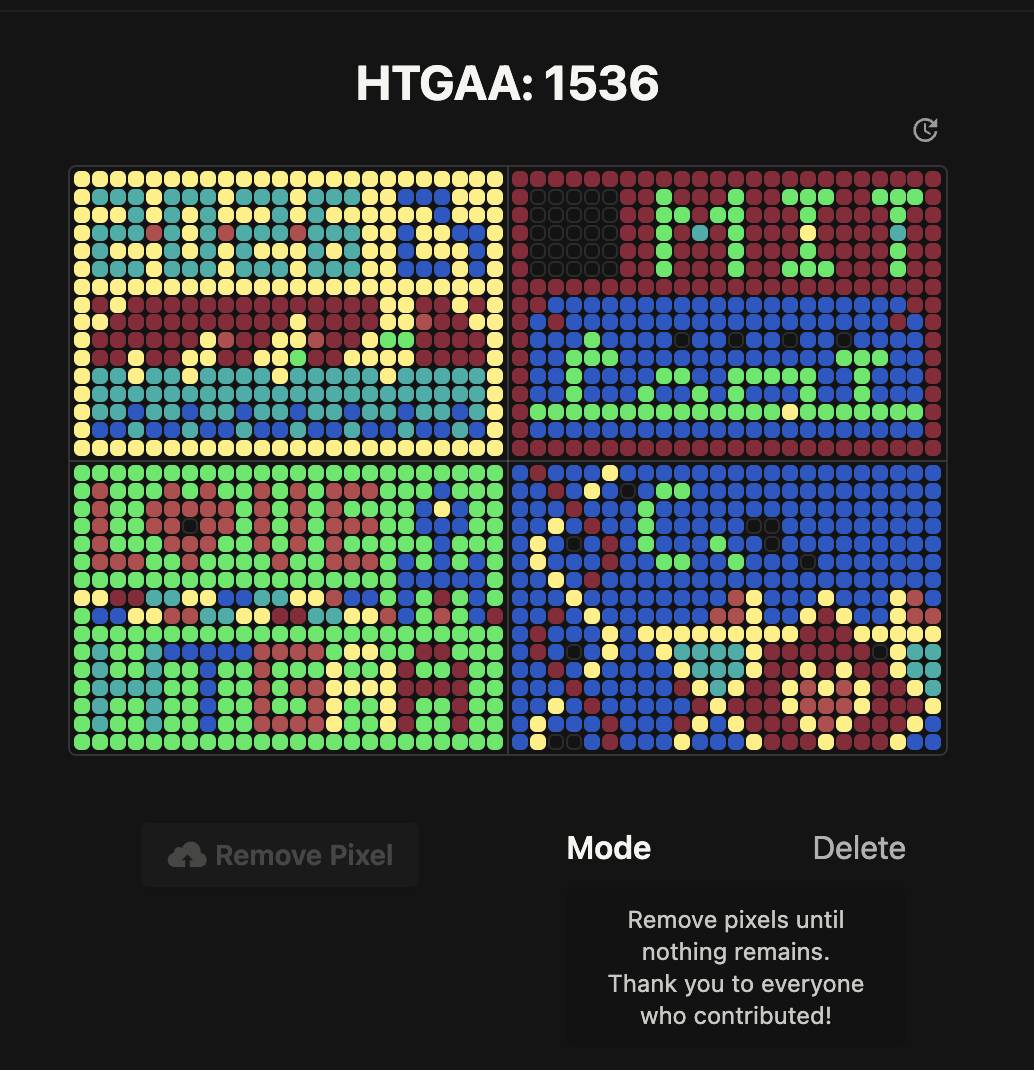

It’s a cool idea but I think you should only be able to modify one pixel one time. Some people wrote scripts to try to get a certain image they want and that kind of defeats the purpose of collaboratively making art.
Part B: Cell-Free Protein Synthesis | Cell-Free Reagents
Referencing the cell-free protein synthesis reaction composition (the middle box outlined in yellow on the image above, also listed below), provide a 1-2 sentence description of what each component’s role is in the cell-free reaction.
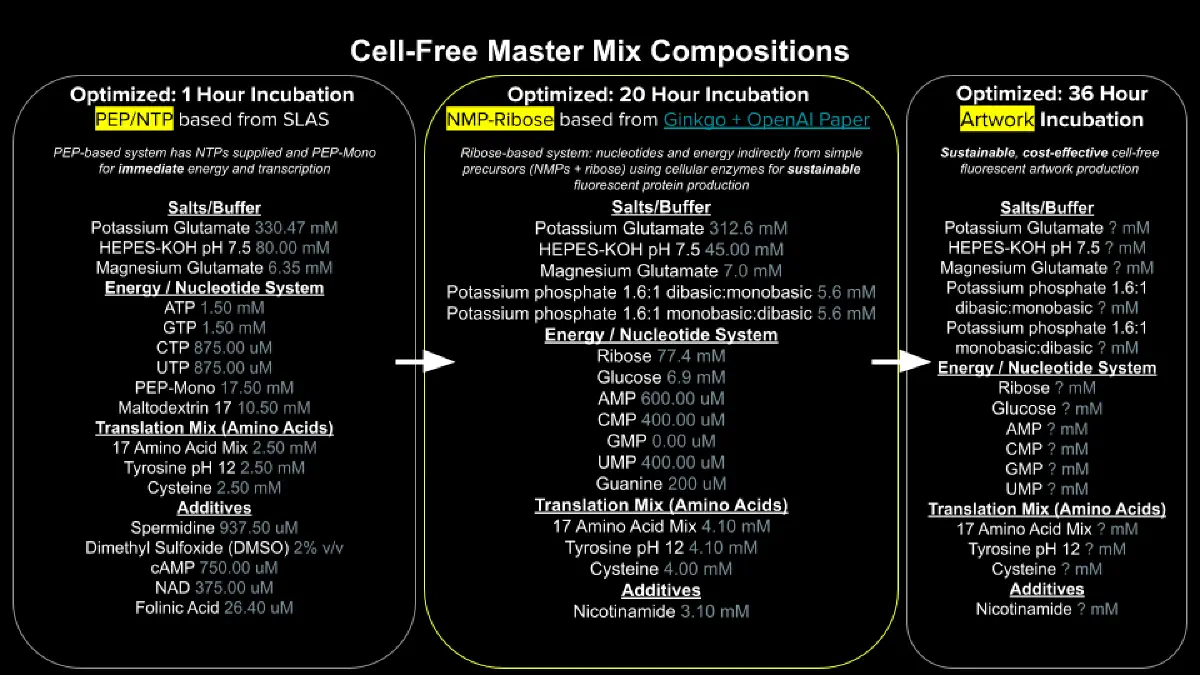
Component Roles
E. coli Lysate
- BL21 (DE3) Star lysates - will contain the protein synthesizing components of E. coli cells such as ribosomes, tRNA, elongation factors and T7 RNA Polymerase.
Salts & Buffers
- Potassium glutamate - is used to control ionic strength and stabilize ribosome complexes.
- HEPES-KOH buffer - maintains optimal pH conditions for enzyme activity.
- Magnesium glutamate - is an ion source for magnesium which serves as a cofactor in both ribosomal activity and enzyme activity.
- Phosphoric acids (monobasic/phosphate) - serve as additional pH buffers as well as contribute phosphor atoms to enable regenerative steps of energy production.
Energy/Nucleotide Regeneration Systems
- Ribose - is fed into the pentose phosphate pathway to generate new NTP’s from existing NMP’s allowing longer time frames with this system.
- Glucose - can be converted by glycolytic pathways to produce ATP for the overall process.
- AMP, CMP, GMP, and UMP - are available for conversion to their corresponding diphosphate or triphosphate forms to support transcription.
- Guanine - is also provided in its free form to help circumvent limitations on generating GMP during the reaction.
Components of Translation Mixture (Amino Acids)
- The 17 amino acid mix plus tyrosine and cysteine make up all of the necessary materials required for peptide bond formation during translation.
- Separate additions of tyrosine and cysteine were made because they have limited solubility in aqueous solutions and cysteine was found to oxidize upon standing.
Additional Components
- Nicotinic acid - has been included to maintain adequate levels of NAD+ in order to keep all redox dependent metabolic reactions proceeding as needed.
Backfilling
- Nuclease-free water - used to bring the reaction mixture to its desired final volume
Part C: Planning the Global Experiment | Cell-Free Master Mix Design
Given the 6 fluorescent proteins we used for our collaborative painting, identify and explain at least one biophysical or functional property of each protein that affects expression or readout in cell-free systems. (Hint: options include maturation time, acid sensitivity, folding, oxygen dependence, etc) (1-2 sentences each)
sfGFP - The sfGFP was created to better fold and have less tendency to aggregate than normal GFP. This creates an advantage in using it in cell free systems, since the folding of mis-folded proteins are aided by chaperone enzymes in cells but do not exist in a cell free system.
mRFP1 is a slow-maturing FP, therefore many of the translated proteins may never achieve their fluorescent state during a cell free reaction due to the consumption of O2 as the reaction progresses.
mKO2 is very sensitive to oxygen levels required to complete chromophore maturation. Therefore, in longer cell free reactions as the available oxygen levels decrease, this can lead to incomplete maturation of chromophores and reduced fluorescence levels.
mTurquoise2 - Due to a slow forming chromophore reaction with oxygen, mTurquoise2 matures slowly and therefore, some portion of the expressed protein may not exhibit fluorescence when measured at the end of the reaction.
mScarlet- I - The T74I mutation in mScarlet-I allows it to mature significantly faster than wild-type mScarlett. However, this comes at the cost of a lower quantum yield. While this is somewhat of a trade-off, in cell free systems, where rapid maturation is advantageous, it is likely still a beneficial mutation.
Electra2 - Since blue FPs such as Electra2 produce inherently less light than green or red FPs, the fluorescence produced by electra2 will be weaker, even if it is well-expressed relative to the other FPs on the plate.
Create a hypothesis for how adjusting one or more reagents in the cell-free mastermix could improve a specific biophysical or functional property you identified above, in order to maximize fluorescence over a 36-hour incubation. Clearly state the protein, the reagent(s), and the expected effect.
My hypothesis is that if we lower the glucose content of the master mix it will slow the rate at which our lysates consume oxygen because less will be required for glycolysis; therefore there should be a greater availability of oxygen to support continued growth and fluorescence of mKO2 during the 36 hour incubation.