Individual Final Project



Final Idea - Symbiotic Circuits

Supervisors: Cale & Shanice
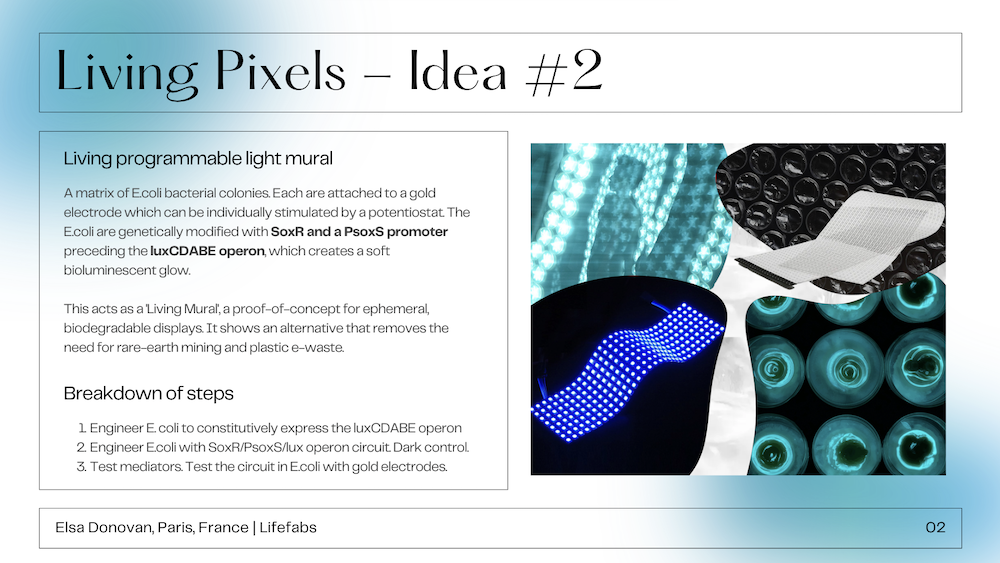
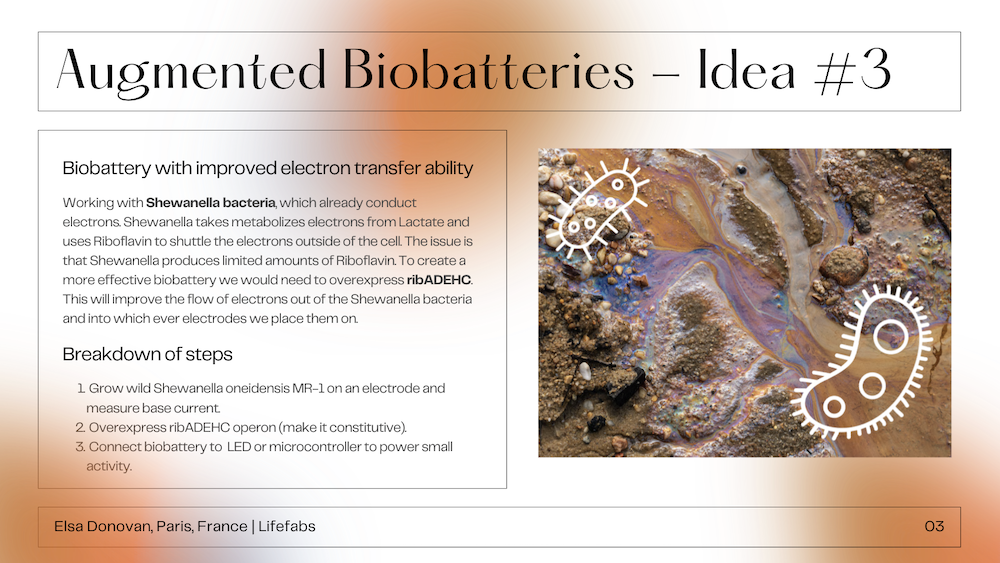
This final project is based on my idea #3 “Augmented Biobatteries” but updated after having taken into consideration the feedback from my mentors. This project will work with Shewanella bacteria to create living biosensors that could monitor soil health in potted house plants. Shewanella are already studied for their ability to test for heavy metals in soil, but with gene editing they can also be used to track and alert for a variety of common soil issues. The ability of Shewanella to be able to create electrical current makes them an extremely useful bacteria when it comes to continuous monitoring. These electrical signals can be collected by a microcontroller and then used to make a variety of outputs - from simple graphs to LED or audio outputs.
In the scope of this project the Shewanella will be engineered to detect nitrate, which is found in fertilizers. This can help to detect over-fertilization issues in the soil which can be detrimental to house plants.
If this proof of concept works, in the future custom designed Shewanella can be used to detect and monitor a range of soil issues and create an alert system before the poor soil health affects the plant itself.
Aims:
Order traditional Shewanella and genetically engineer Shewanella (where MtrC is behind a nitrate sensitive promoter). Measure the electrons in both strains using potentiostats. For the modified Shewanella measure the power density of the current with and without added nitrate. This is to have a good understanding of the baselines.
For biosecurity reasons an auxotrophy kill-switch will need to be included: the gene that allows Shewanella to make thymidine will be deleted, and as a result thymidine will be supplied externally to the growth medium. Without this external supplementation the Schewanella strain will die.
Transplant the modified Shewanella into a pot with a Pothos plant. Test for the best possible environmental conditions in the modified Shewanella bacteria can thrive (ie. soil humidity, supplement lactate, etc.).
Product design aim: create an aesthetic pot that is fitted with electrodes on the inside which collect and send the fluctuating electron signals. to a microcontroller. This microcontroller can light up the pot with different LEDs that can act as a continuous visual indicator of the plant’s soil health.
Protocol
- Order genetically modified plasmid from Twist.
- Order Shewanella bacteria separately - not sure from where?
- Transform the bacteria
I won’t be in-person in LifeFabs however I will be in London for the last 2 weeks of July if there is a possibility of working on my project there. I am currently based in Montreal and have access to a BSL-1 lab so I might try to order wild type Shewanella to get some baseline measurements as part of my aim 1. I have to check with the lab supervisor first.
[P_napD promoter] → [RBS] → [mtrC coding sequence] → [terminator]
Nitrate promoter in Shewanella has never been done before. Therefore I want to start by overexpressing the mtrC pathway. That way I can test whether this type of activation would work in a potting soil environment before working with a nitrate sensitive promoter.
Shewanella has been used to detect arsenic using an arsenic promoter usually designed for E.coli. I found the plasmid from the Ng et Al 2018 paper: https://www.addgene.org/174615/
References:
Red-Light-Induced Genetic System for Control of Extracellular Electron Transfer https://pubs.acs.org/doi/10.1021/acssynbio.3c00684
This is a paper that shows that
Turn on the Mtr pathway genes under pLacI promoter in Shewanella oneidensis MR-1 Ng et al. (2018) https://link.springer.com/article/10.1186/s40643-018-0221-9
I am creating a DNA sequence that
Benchling Project Link: https://benchling.com/ekdonovan/f_/qNHLaHwB9L-htgaa_final_project/
Steps for Aim 1:
- Receive the plasmid from Twist
- Transform it into E. coli to amplify
- Transform it into Shewanella
What the article shows is that:
the promoter pLacI (R0010) combined with RBS (B0034), which was designed by iGEM, was activated efficiently in S. oneidensis MR-1. iGEM link: https://parts.igem.org/Part:BBa_B0034
pLacI has potential as a common promoter for S. oneidensis MR-1 for exogenous genes.
The new design:
Pnap (or PnrfA) → RBS (B0034) → mtrC → rrnB T1 terminator + T7Te terminator
Nitrate-responsive promoter from: https://journals.plos.org/plosone/article?id=10.1371/journal.pone.0051643
nap and nrfA promoters yeaR promoter (from E.coli - could be used in Schewanella?)
Nitrate promoters in Shewanella are understudied. I was not able to find the right genetic sequence for this.
Updates:
I’ve been spending a lot of time reading relevant research papers, trying to extract gene sequences from their methodology sections. I can’t find a confirmed gene sequence for the innate nitrate-responsive promoter in Shewanella. Or the mtrC sequence for that matter. It’s supposed to be in the addGene here: https://www.addgene.org/174615/ But I tried to annotate the specific mtrC gene in this addGene sequence in Benchling, but I couldn’t externally verify the sequence on uniprot or NCBI or any other database. I just wasn’t able to find it anywhere. It took all day and I came up with nothing.
This whole thing is becoming very complicated. I think it’s because Shewanella is still understudied, compared to E.coli and the nitrate-responsive promoters in Shewanella are also poorly understood. So I can’t find the genetic sequence for mtrC or the nitrate-responsive promoters.
So instead I’m going to use E.coli for this project instead beacuse the gene sequences I’m looking for are better documented. I’ll take genes from Shewanella, like the electron transfer pathway (mtrC) and implement them in e.coli and put them behind a e.coli-friendly nitrate responsive promoter.
In order to make this more effective I’m going to base myself off this iGEM project: https://parts.igem.org/Part:BBa_K1316012
I’d have to make a Soil Microbial Fuel Cell to test this.
Graphite or carbon cloth as the anode and an electrode on the surface for the cathode.
Pyear promoter for E.coli: https://parts.igem.org/Part:BBa_K216005
This project is becoming very complicated, but I guess that’s par for the course when working with biology.
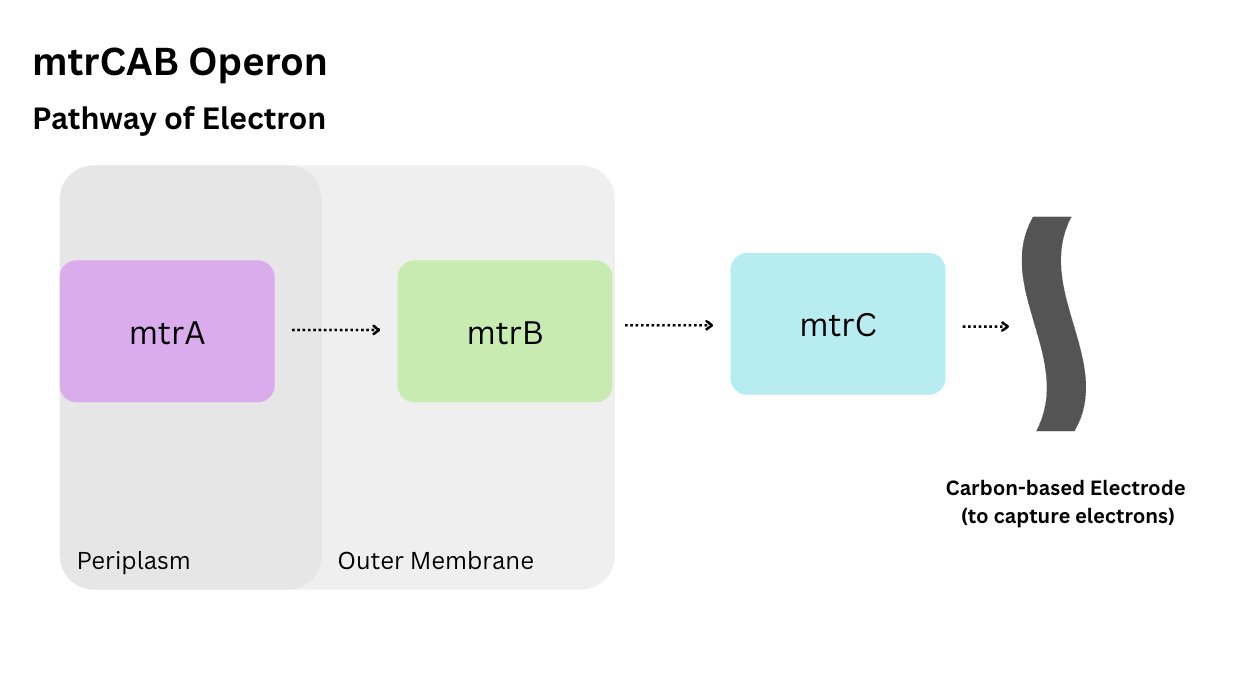
I’m going to need to use the full mtrCAB operon as they did in the iGEM project mentioned above if I want to create a “conductive bacteria”. These proteins work together as a “chain” to transport the electrons from the inside of the E.coli membrane to the outside. I created the diagram below to illustrate this process: