Group Final Project
Proposal: Rational Enhancement of MS2 Lysis Protein Toxicity
Objective
This proposal addresses the subproblem of increasing the toxicity of the L lysis protein from Bacteriophage MS2. Instead of random mutagenesis, toxicity will be approached as a multi-factor optimization problem involving structural stability, membrane insertion, oligomerization efficiency, and expression kinetics in Escherichia coli. The objective is to design L variants that enhance membrane disruption while maintaining proper folding and stability.
Proposed Computational Strategy
First, protein language models (e.g., ESM-2, ProtT5) will be used to perform directed in silico mutagenesis. These models capture evolutionary constraints and residue interactions, enabling the generation of structurally plausible variants while identifying mutation-tolerant and functionally critical positions. This step efficiently reduces the combinatorial search space. Second, predicted variants will be structurally evaluated using AlphaFold2 for monomer folding and AlphaFold - Multimer to assess oligomerization and interaction with host factors such as DnaJ. Variants that preserve global structure and strengthen oligomeric interfaces will be prioritized. Third, membrane compatibility will be analyzed using membrane-aware modeling (RosettaMP) and selected molecular dynamics simulations. These tools estimate insertion stability and transmembrane behavior, key determinants of lytic efficiency. Fourth, ΔΔG prediction tools (e.g., FoldX, Rosetta energy functions) will filter out destabilizing mutations. In parallel, codon optimization algorithms will redesign selected variants for improved expression in E. coli, as toxicity depends on both structure and intracellular concentration.
Rationale
Toxicity emerges from the combination of folding stability, cooperative oligomerization, membrane insertion, and sufficient expression levels. No single tool captures all these dimensions. Integrating sequence-based generative modeling, structural validation, membrane simulation, and stability prediction enables rational prioritization of high-confidence variants and reduces experimental screening burden.
Potential Obstacles
One limitation is the scarcity of quantitative datasets linking specific mutations to measured lysis kinetics, which restricts supervised learning approaches. Additionally, structural prediction tools do not explicitly model lipid bilayers, so oligomeric pore assemblies may require extensive molecular dynamics validation.
Pipeline Overview
Generate ~5,000 variants with protein LLMs, filter by ΔΔG stability, predict structure and oligomers with AlphaFold, evaluate membrane behavior, optimize codons, select top candidates for experimental lysis assays. This streamlined computational - experimental workflow enables targeted optimization of MS2 L protein toxicity through rational design rather than random screening.
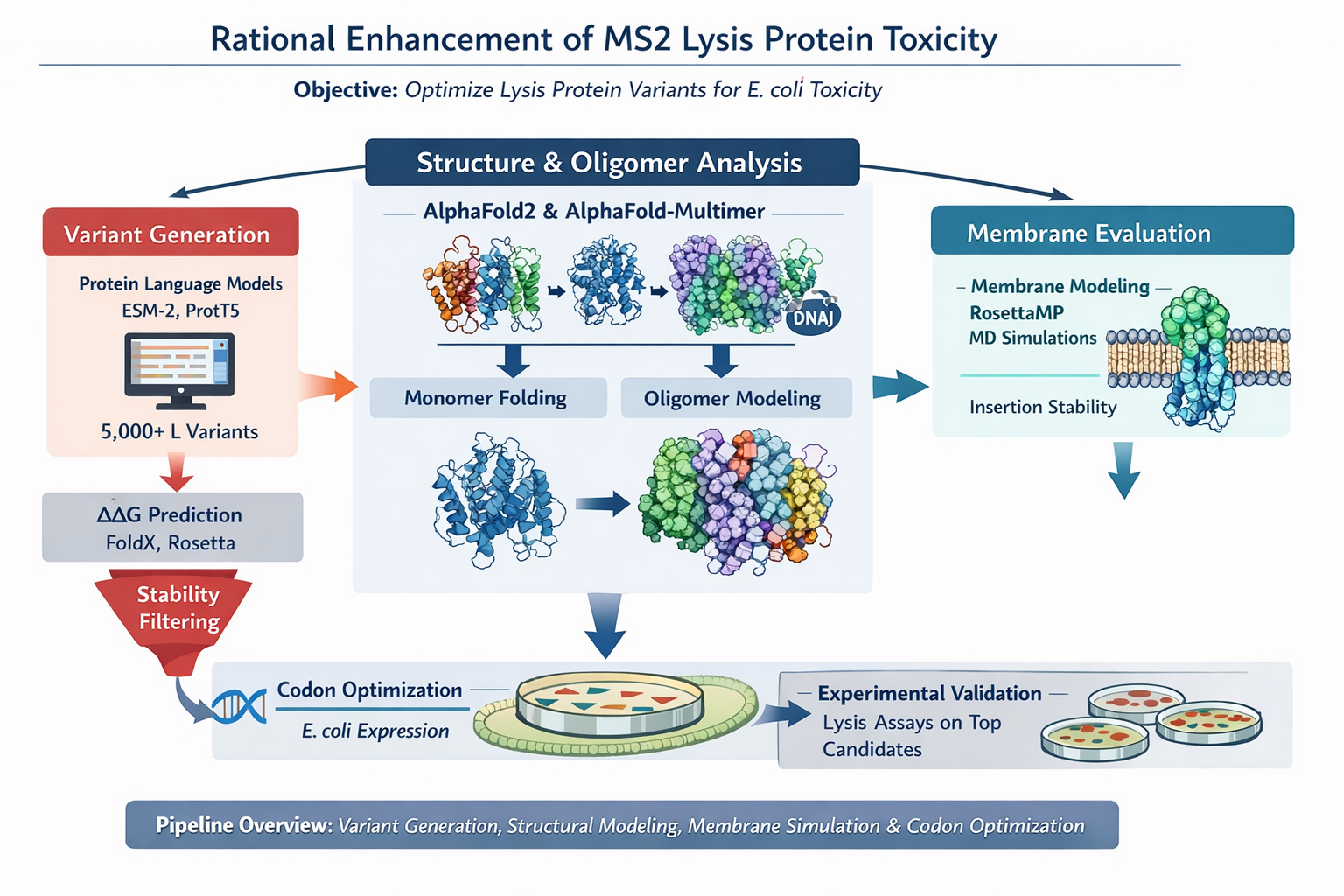
Figure 8. Group project idea. Image generated with AI.