Week 4 HW: Protein Design Part I
Part A. Conceptual Questions
1. How many molecules of amino acids do you take with a piece of 500 grams of meat? (on average an amino acid is ~100 Daltons)
Meat is about 20% protein, so that is 100 g of protein. There are 6.022e+23 Daltons per gram, so for 100 grams that is 6.022e+25 Daltons. Then if there are 100 Daltons in an average amino acid, we’re back to 6.022e+23 molecules of amino acids.
2. Why do humans eat beef but do not become a cow, eat fish but do not become fish?
Because or food prepartion and digestive systems have broken the cow (and cow DNA), the cow DNA does not mix with human DNA to produce new cells, and our immune systems fight DNA other than our own.
- To be continued … ??? … Will finish later …
Part B: Protein Analysis and Visualization
1. Briefly describe the protein you selected and why you selected it.
I chose the protein “Oxidoreductase cns1” from Cordyceps militaris (strain CM01) (Caterpillar fungus). (https://www.uniprot.org/uniprotkb/G3JF08/entry) I chose it because I am interested in parastic fungi and because it is “part of the gene cluster that mediates the biosynthesis of cordycepin (COR)” and “Cordycepin has antitumor, antibacterial, antifungal, antivirus, and immune regulation properties.”
2. Identify the amino acid sequence of your protein.
The FASTA is at https://www.ncbi.nlm.nih.gov/protein/XP_006669647.1?report=fasta and the amino acid sequence is at https://www.uniprot.org/uniprotkb/G3JF08/entry#sequences which says its length is 792.
XP_006669647.1 oxidoreductase domain-containing protein [Cordyceps militaris CM01] MAMNENAYPTTFPSFERENHRDALRQPFDPAFRRTWSNGVALRQLVDFARPTVANHTMSYALIEYCLSRL PMQHLERLGQLKIPVELHAAPFQYLQKHHRACGFDWVERFVWRTHDLHKPYNFLRPELLLAQESGSQRIV ALLTIMPGEDYIRHYASILEVAQHDGAISSHHGPIRCVLYPHLTQSMMAWTGLTELSLSVEPGDILILGF VAELLPRFASLVPTARVIGRQDAQYYGLVRLELRPGLVFSLIGAKYSYWGNLGGRVVRELAARRPRAICY IAKQGTLLSPGDIHRTIYSPTRYCVFDKGQACWHGDDHSALPINPLSSRFPTFDRGLHVSTPTIVEQDVD FRTQVEAHGASSVDNELAQMARALTDVHEENPSMERVQLLPLMFITDYLRRPEELGMTVPFDLTSRNETV HRNKELFLARSAHLVLEAFNVIERPKAIIVGTGYGVKTILPALQRRGVEVVGLCGGRDRAKTEAAGNKHG IPCIDVSLAEVQATHGANLLFVASPHDKHAALVQEALDLGGFDIVCEKPLALDMATMRHFANQSQGSSQL RLMNHPLRFYPPLIQLKAASKEPSNILAIDIQYLTRRLSKLTHWSAGFSKAAGGGMMLAMATHFLDLIEW LTSSSLTPASVQDMSTSNSIGPLPTEDAGATKTPDVESAFQMNGCCGLSTKYSVDCDGAADTELFSVTLR LDNEHELRFIQRKGSPVLLEQRLPGREWLPLKVHWEQRVREGSPWQISFQYFAEELVEAICMGTRSAFAD KATGFSDYARQVGVFGSKVGIA
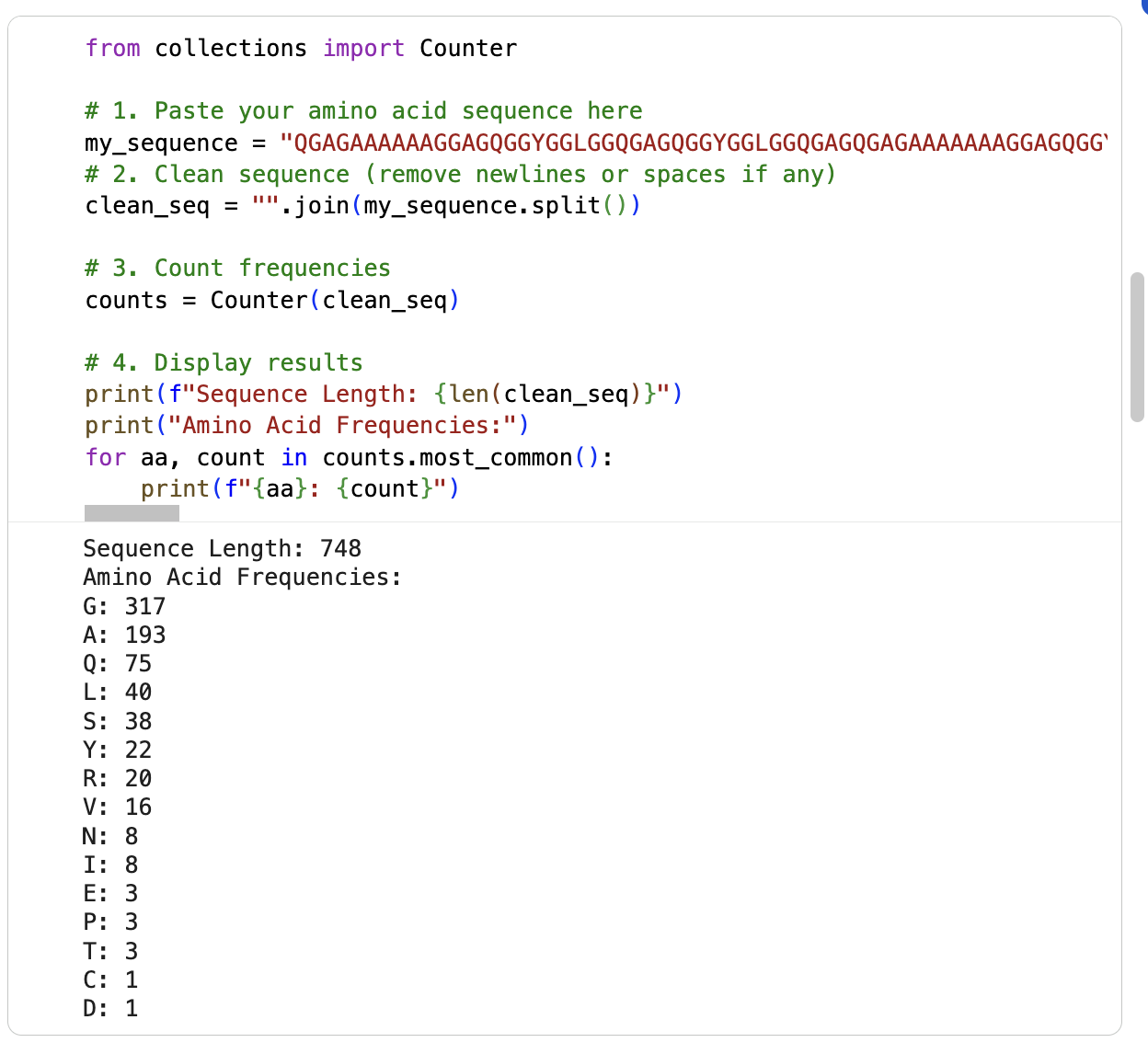
I am running the google collab notebook to count the frequency of amino acids (had some errors) … it says the sequence length is 748 and the most frequent amino acid is glycine (317) followd by alanine (193). See below:

I ran a search for homologs on BLAST. There are six that are in the red, that have an E value of about 0. See below:
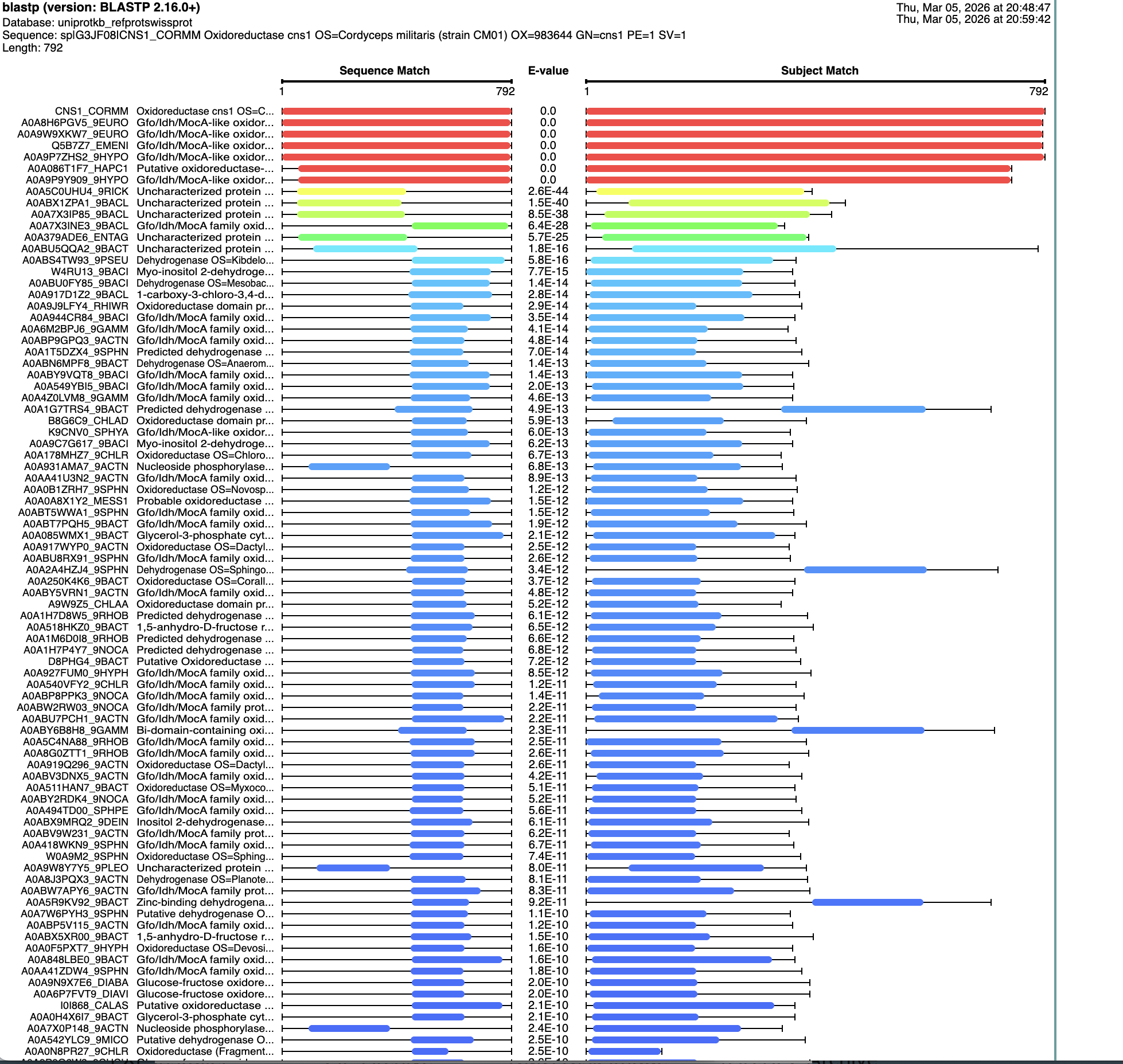

It is reportedly part of the “Gfo/Idh/MocA Oxidoreductase: family according to https://www.ebi.ac.uk/interpro/entry/InterPro/IPR052515/ as well as the “NAD(P)-binding domain superfamily” according to https://www.ebi.ac.uk/interpro/entry/InterPro/IPR036291/ and “Gfo/Idh/MocA-like oxidoreductase, N-terminal” according to https://www.ebi.ac.uk/interpro/entry/InterPro/IPR000683/
3. Identify the structure page of your protein in RCSB.
I dod not find the “Oxidoreductase cns1” from Cordyceps militaris (strain CM01) (Caterpillar fungus)” in RCSB. I did find “Crystal Structure of endo-beta-N-acetylglucosaminidase from Cordyceps militaris D154N/E156Q mutant in complex with fucosyl-N-acetylglucosamine” at https://www.rcsb.org/structure/6KPN
I don’t see a solved date. It was Deposited: 2019-08-15. I don’t see it listed as part of any structure classification family at https://www.ebi.ac.uk/pdbe/scop/
4. Open the structure of your protein in any 3D molecule visualization software
Cartoon:
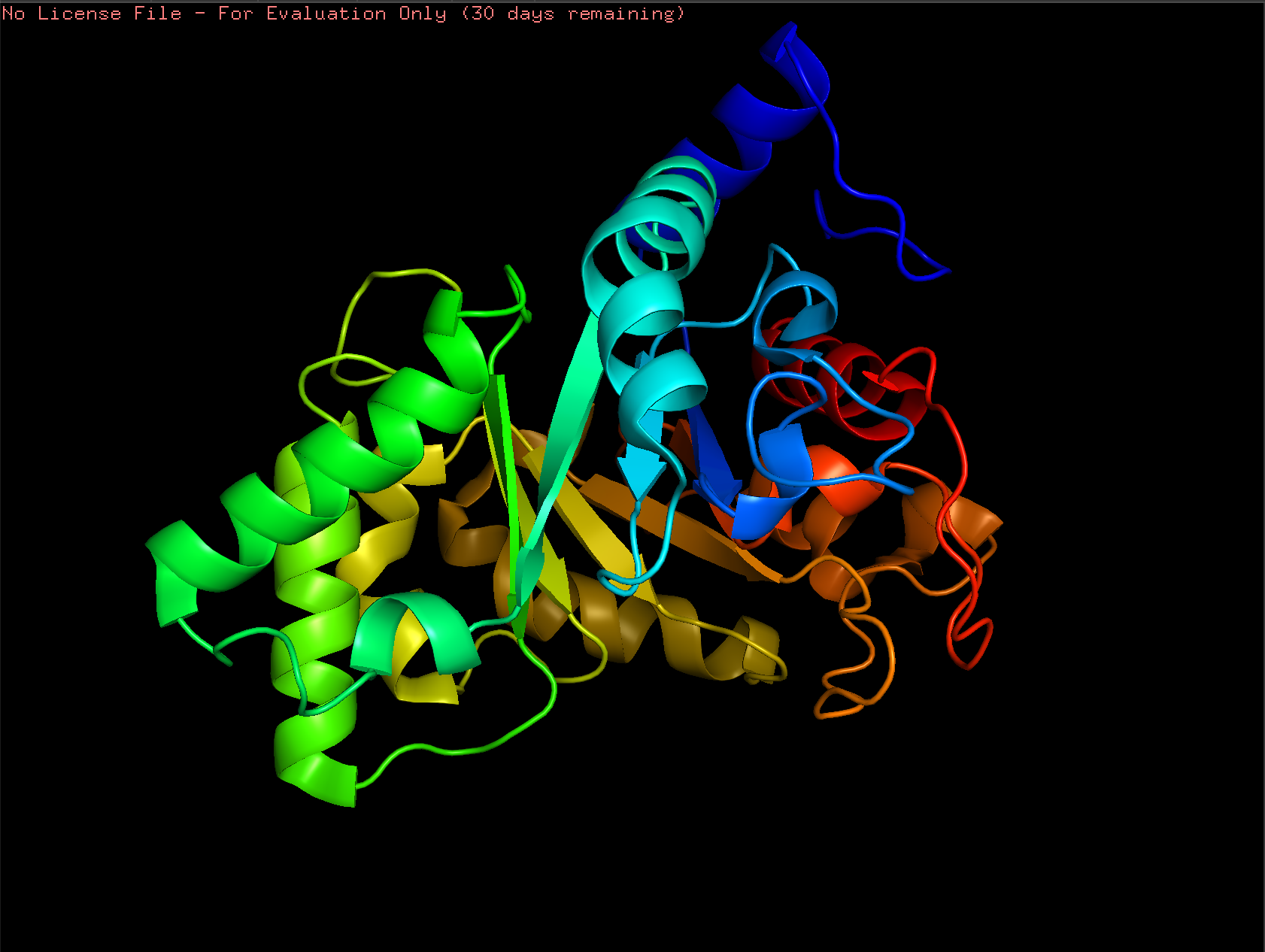

Ribbon:
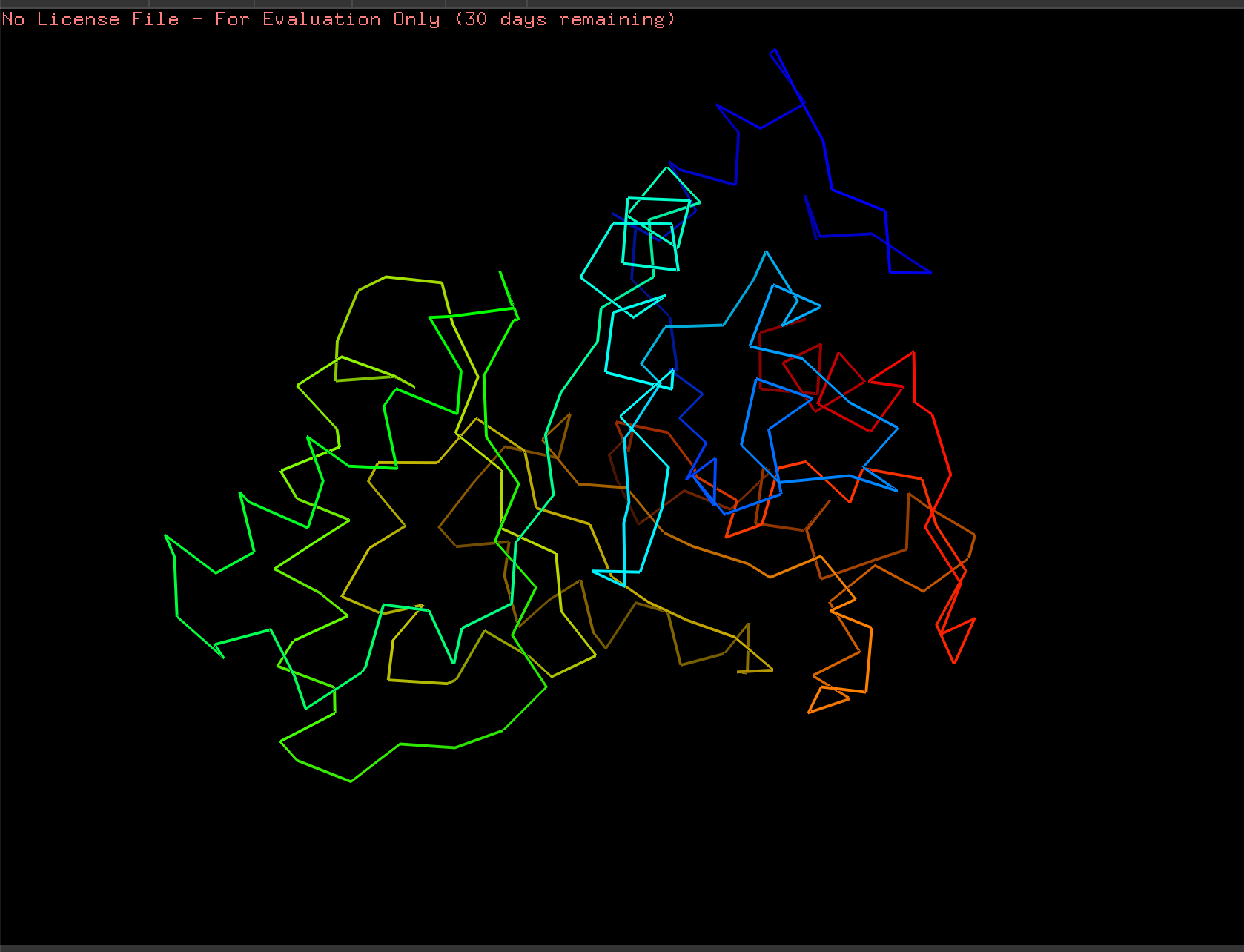

Ball and Stick:

To Do: Color the protein by secondary structure. Does it have more helices or sheets? Color the protein by residue type. What can you tell about the distribution of hydrophobic vs hydrophilic residues? Visualize the surface of the protein. Does it have any “holes” (aka binding pockets)?
Part C1. Using ML-Based Protein Design Tools: Protein Language Modeling
My Google Collab is at https://colab.research.google.com/drive/1xcDY5dXbRKVTG2YmNOSjDgwPw2_QO5cB#scrollTo=ySOWXRjTja9D …
1. Deep Mutational Scans
Here is my mutation scan heatmap:
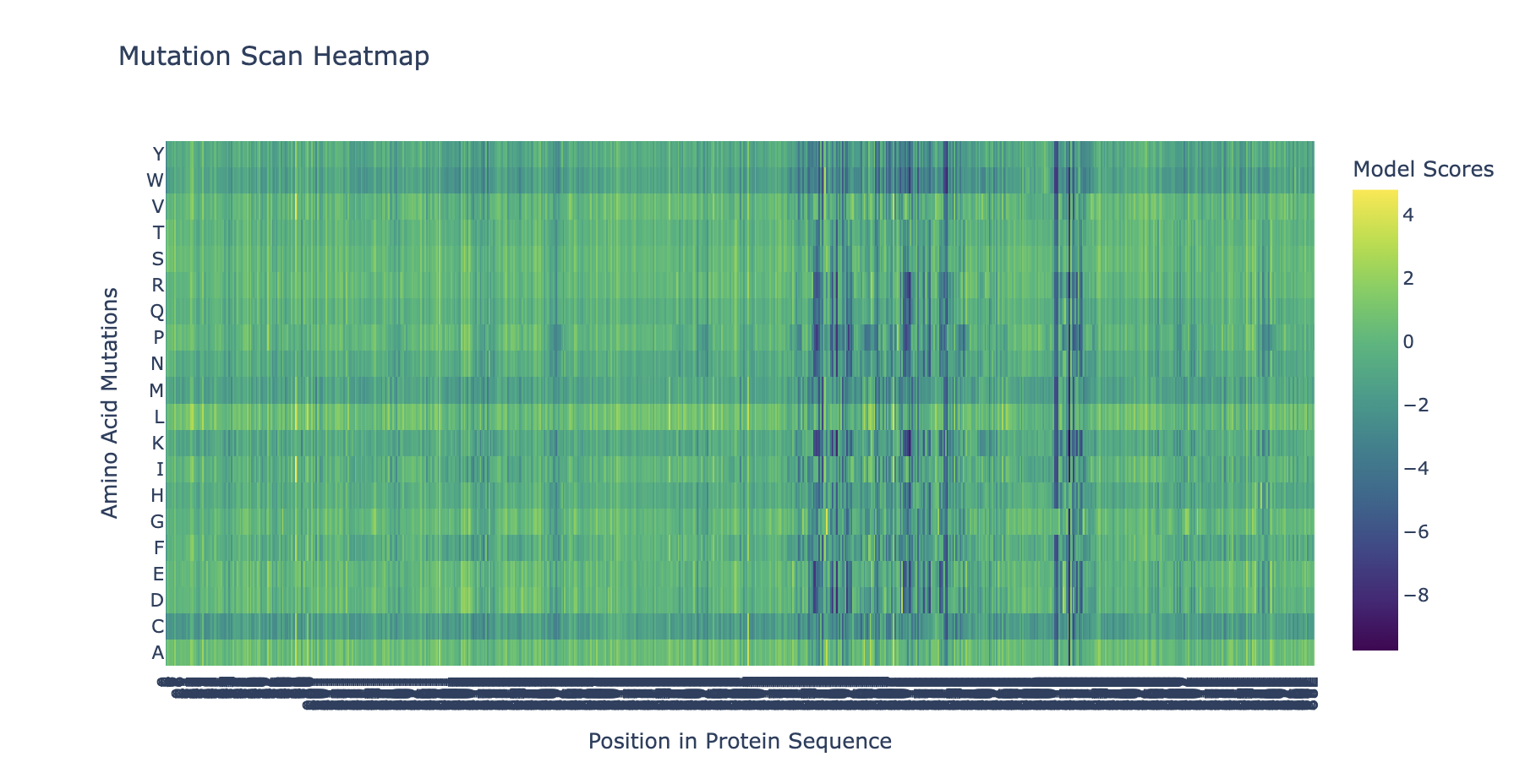

To do: Can you explain any particular pattern? (choose a residue and a mutation that stands out) and (Bonus) Find sequences for which we have experimental scans, and compare the prediction of the language model to experiment.
2. Latent Space Analysis
To do: Use the provided sequence dataset to embed proteins in reduced dimensionality. Analyze the different formed neighborhoods: do they approximate similar proteins? Place your protein in the resulting map and explain its position and similarity to its neighbors.