Week 2 HW: DNA Read-Write-Edit
Checklist
- Part 0: Basics of Gel Electrophoresis
- Attend Lecture (2 of 3)
- Attend Recitation
- Review 2025 recording (3 of 3)
- Part 1: Benchling & In-silico Gel Art
- Part 2: Gel Art - Restriction Digests and Gel Electrophoresis (Optional- for those with Lab access)
- Design Simulation
- Part 3: DNA Design Challenge
- 3.1 Choose your Protein
- 3.2 Reverse Translate: Protein (amino acid) sequence to DNA (nucleotide) sequence.
- 3.3 Codon optimization
- 3.4. You have a sequence! Now what?
- 3.5. [Optional] How does it work in nature/biological systems?
- Part 4: Prepare a Twist DNA Synthesis Order
- 4.1. Create a Twist account and a Benchling account
- 4.2. Build Your DNA Insert Sequence
- 4.3. On Twist, Select The “Genes” Option
- 4.4. Select “Clonal Genes” option
- 4.5. Import your sequence
- 4.6. Choose Your Vector
- Part 5: DNA Read/Write/Edit
- 5.1 DNA Read
- (i) What DNA would you want to sequence (e.g., read) and why?
- (ii) In lecture, a variety of sequencing technologies were mentioned. What technology or technologies would you use to perform sequencing on your DNA and why?
- 5.2 DNA Write
- (i) What DNA would you want to synthesize (e.g., write) and why?
- (ii) What technology or technologies would you use to perform this DNA synthesis and why?
- 5.3 DNA Edit
- (i) What DNA would you want to edit and why?
- (ii) What technology or technologies would you use to perform these DNA edits and why?
- 5.1 DNA Read
Part 1: Benchling & In-silico Gel Art
In this section, I was able to successfully sign up for Benchling, request to join HTGAA (pending), and create a new project. I was able to find the Lambda DNA sequence in the FASTA database, which I copied and pasted. I then found the downloadable file in GenBank, which I imported into Benchling. It took me a few tries to get multiple Digests to appear, once I selected multiple restriction enzymes and ordered the tabs before Virtual Digest. I exported the resulting image as a .PNG as well as my NC_001416 Project “Linear Map” and “Sequence Map” as well as the Lambda Map from GenBank, as PDFs for future reference.
Part 2: Gel Art
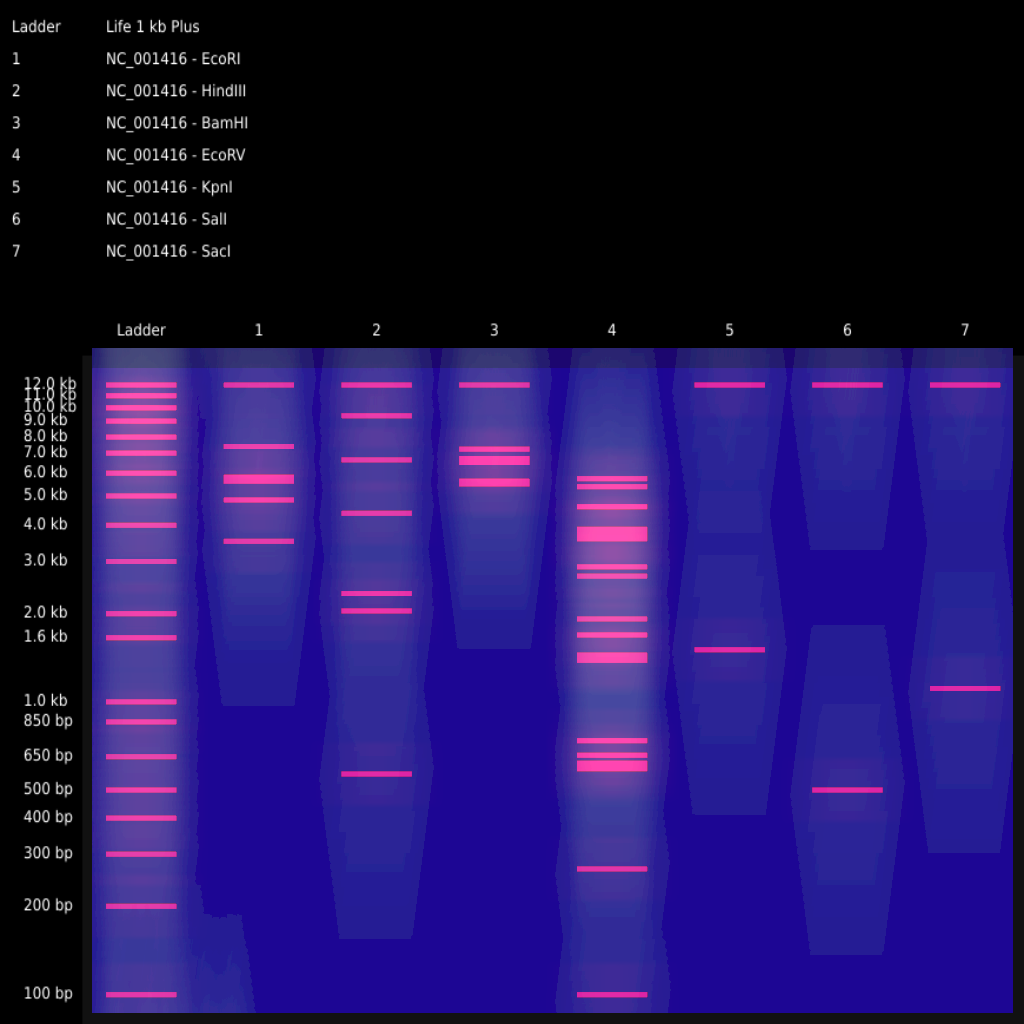

Illustration by Eric Schneider: Simulated Gel Electrophoresis using node based image editing software, “Adobe Substance Designer”
Part 3: DNA Design Challenge
- 3.1 Choose your protien
I chose Candida antarctica Lipase B (CalB) since it has the ability to break down polylactic acid, or PLA, a commonly used filament in 3D printing.
My design intent is to reduce the amount of microplastics that reach our ecosystem. The best place to start remediation may be at the source; the waste stream of PLA that is generated in a MakerSpace. By introducing a biological process that depolymerizes PLA waste, we may contribute to a solution while providing governance and building awareness.
From a BioArt perspective, this is the first step in creating and containing the lactic acid and CO2 that may be generated, for downstream use in feeding and growing colorful algae. In turn, powdered algae pigment can be extracted, showing how PLA can help to create colorful pigments used in painting and other mediums.
3.2 Reverse Translate
I was able to find a suitable Protein for this design challenge by using Ai Prompts and comparing results between ChatGPT and Claude. ChatGPT led me to Proteinase K https://www.ncbi.nlm.nih.gov/nuccore/X14689 which turned out to be very challenging due to complexity of the construct, and actually caused Twist to “freeze” when attempting to synthesize.
I even conducted a rapid experiment where I asked Claude Ai to provide the translation, which it suprisingly did, very confidently. However, I ran into the same complexities when attempting to create a TWIST order.
I went back to the NIH database and found C.antarctica (LF 058) gene for lipase B https://www.ncbi.nlm.nih.gov/nuccore/Z30645 which, according to Claude Ai, would lead to better results with less complexity. I exported a FASTA file for the protein’s genetic structure.
In TWIST, the Lipase B approach fell into the “standard” complexity level, so I am sticking with that translation. Which also validates that the follow-up Claude AI inquiry led to a good result. (see appendix for summary of prompt usage)
- 3.3 Codon Optimization
I used the Twist tool to optimize Codons. It showed me two regions that had repeating sequences that could be optimized.
Question: It seems that the Start and Stop codons were automatically added in, as ATG, and TAA but I want to better understand when and how to ensure they are present manually, with dependency on selected expression. (Note: Answer was found by properly annotating)
I chose e.coli as I learned that it is predictable and suitable for this sequence. Yeast may be used for a higher yield, but with possibly more optimization of repeating codons needed. I completed the Twist optimization, and downloaded the sequence to view in Benchling to learn more about the strucutre.
- 3.4 You have a sequence! Now what?
This protein can be created from DNA from either clonal or strand synthesis. The dna sequence I have identified can be inserted into a host plasmid which is cloned in an industrial-scale lab that can provide quality, speed and editing capabilities. The cell-based method provides more synthetic control and expected outcomes, acting like a factory. The cell-free method may introduce toxins and have lower yield. In advanced industrial production, both may work together for rapid prototyping and scalability.
3.5 [Optional] How does it work in nature/biological systems?
The ability to transcribe from different start points in the sequence leads to diveristy in proteins created.
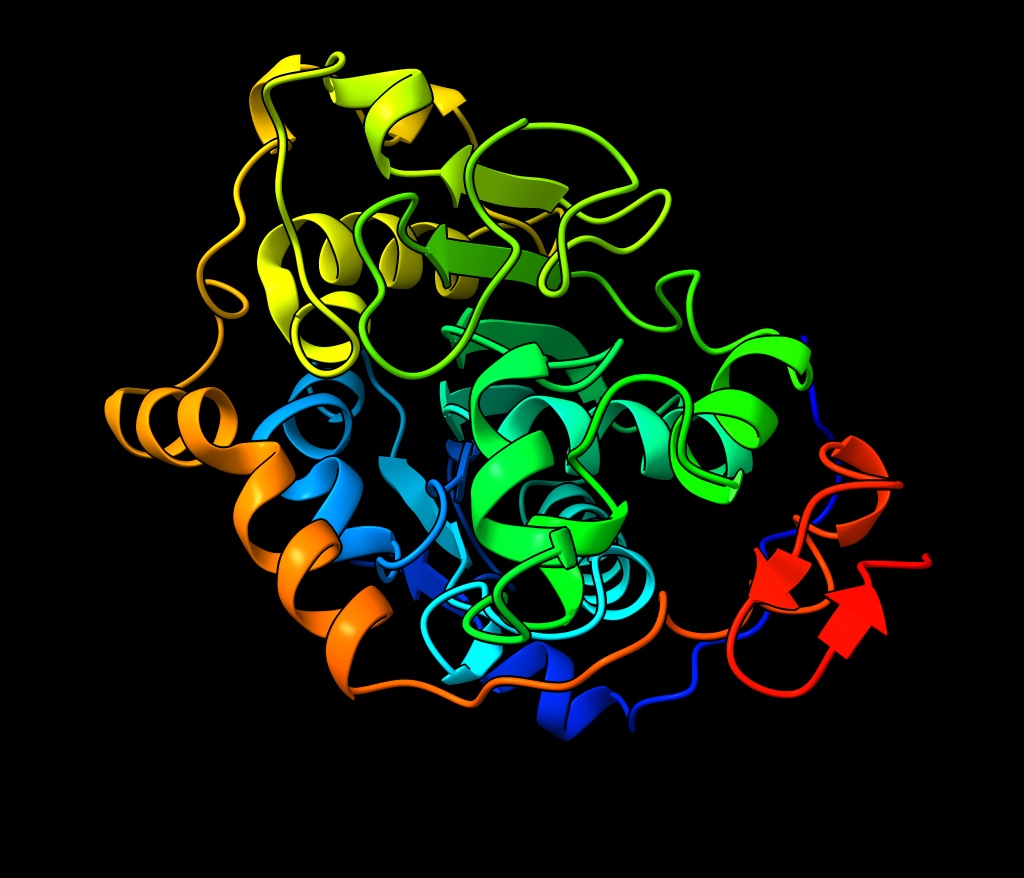

Illustration by Eric Schneider: Lipase B 3D model from UniProt https://www.uniprot.org/uniprotkb/P41365/entry#structure and rendered with ChimeraX https://www.rbvi.ucsf.edu/chimerax
Part 4: Prepare a Twist DNA Synthesis Order
I realized that my prior attempt to create an order was incomplete, as I had not fully optimized or annotated my sequence. I started “from scratch” and optimized my sequence in TWIST, then exported back to Benchling, where I prepared a sequence with the proper annotations. I took this back into Twist and prepared an order. I exported the new Plasmid back to Benchling. This “answered” my initial question related to Annotating start and stop codons, which was a key learning for me.
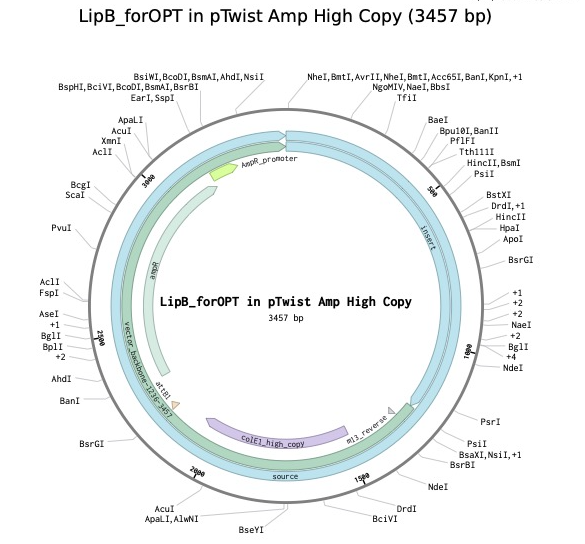

Part 5: DNA Read/Write/Edit
- Part 5: DNA Read/Write/Edit
- 5.1 DNA Read
- (i) What DNA would you want to sequence (e.g., read) and why?
- I would like to sequence the DNA of a Lipase as it appears to be well suited for the depolymerization of PLA. I would also like to sequence a Cutinase as it has similar properties, as well as Proteinase K which may be best for industrial-scale applications. I am intrigued by the potential for a hybrid solution . I am also interested in harnessing any CO2 emissions for downstream processing or pigmented algae growth.
- (ii) In lecture, a variety of sequencing technologies were mentioned. What technology or technologies would you use to perform sequencing on your DNA and why?
- I would use UniProt to locate Proteins with sequences.
- I would use TWIST or other standalone optimization tools to minimize repeats in a sequence
- I would use Benchling as the primary method of visualizing sequences to be able to annotate and construct sequences with better probability of success when ordering Clones or Strands.
- I would use TWIST for the speed, quality, and configuration capabilities when building Plasmids.
- I would again use Benchling to visualize Plasmids once constructed.
- I also learned about ChimeraX to 3D visualize Nucleotides and molecular bonds
- (i) What DNA would you want to sequence (e.g., read) and why?
- 5.2 DNA Write
- (i) What DNA would you want to synthesize (e.g., write) and why?
- In support of my PLA depolymerization design, I would want to manage and control the throughput through synthetic means, in contrast of depending on natural biodegradation, which may happen only under the most optimal conditions such as heat, sunlight/UV and presence of enzyme producing organisms.
- (ii) What technology or technologies would you use to perform this DNA synthesis and why?
- Using a technology like TWIST as well as a safe and operational synthetic biology lab, I feel that a repeatable solution can be designed that can scale to the global use case of 3D printed PLA filament sources of microplastic waste reduction
- (i) What DNA would you want to synthesize (e.g., write) and why?
- 5.3 DNA Edit
- (i) What DNA would you want to edit and why?
- I would like to edit the DNA of enzymes that biodegrade PLA to create higher yield, lower temperature requirements, and safe industrial processing to ensure the production is accessible to the quickly growing market segment. This may lead to greater awareness of the growing problem of microplastics through educational Makerspace activities that demonstrate this concept.
- (ii) What technology or technologies would you use to perform these DNA edits and why?
- I would start with well-tested and proven enzymes such as LipaseB to ensure a baseline for any future experimentation. I would follow well-defined procedures of synthesizing DNA. For example, eColi is deemed a good vector, and yeast is also compatible. Once I have validated that a sequence can be synthesized, I would like to order via Twist, and collaborate with a Node Lab to conduct a PLA experiment with a control group, and measure PH, Co2 emissions, and weight delta, as well as temperature monitoring.
- (i) What DNA would you want to edit and why?
- 5.1 DNA Read
Appendix
Ai Prompts
Chat GPT was used to explore the environmenal and ecological impact of microplastics, which led me to the idea of capturing waste at the source.
Here is a condensed list of prompt themes used:
- What biological system (enzyme) can depolymerize PLA into lactic acid?
- What environmental problem does PLA create, especially regarding microplastic persistence in oceans?
- How can PLA waste be prevented from entering mixed waste streams through source segregation?
- What experimental conditions are required to depolymerize PLA at small scale?
- How can successful depolymerization be quantitatively measured (mass balance and lactic acid detection)?
- How can the experiment avoid generating microplastic through mechanical fragmentation?
- What happens to PLA in marine environments or when ingested by sea life?
- How can lactic acid or derived CO₂ be reused in biological systems (plants, algae)?
- How can algae-derived pigment serve as a material outcome of the carbon loop?
Claude Ai seemed to better undertand the boiengineering context:
- What are some examples of polyester hydrolase
- what (enzymes) cuts down PLA the best
- Confirm which are considered synthetic and effective
- what is proteinase K derived from
- what enzyme will work best for DNA replication
- is eColi or yeast better
- clarify cell-dependent or cell-free methods, of synthetic biology