Week 3 HW: Lab Automation
Focus on Lab Automation research, with creative examples of OpenTrans instruction sets using Python. Final project slide to be included in Node deck.
Opentrons Art
This week started witn an exploration of the Opentrons Art web app found at https://opentrons-art.rcdonovan.com
I was able to quickly upload an image and randomize the colors, to generate a point paired data set. I really like the bitmap rasterization and creative expression found in the gallery.
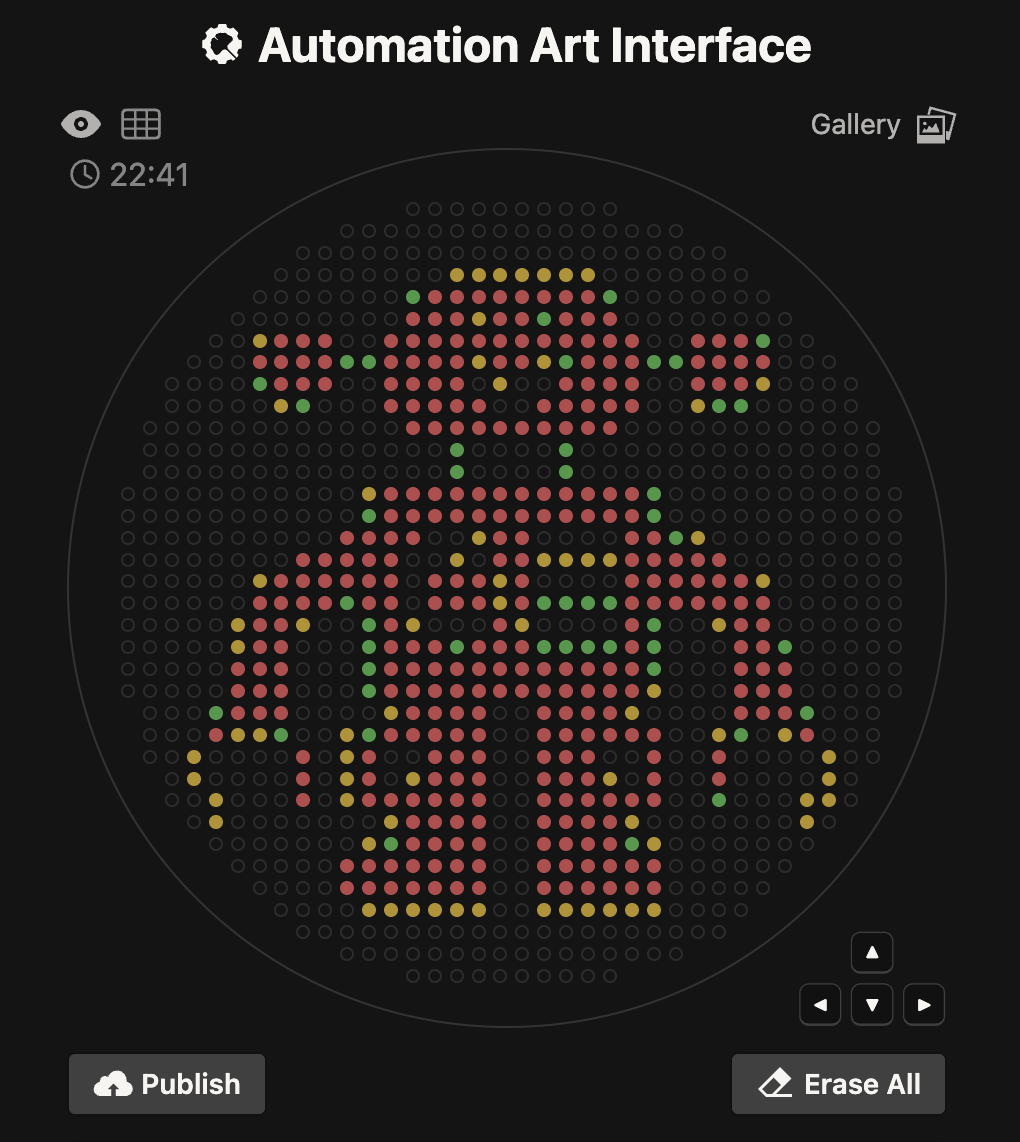

My investigation is based on my background in high resolution digital imaging. I wanted to better understand the pixel to microliter (uL) relationship. I see that with a 200 uL maximum quantity and a 90-100 mm Petrie Dish, it would seem that there are some basic constraints.
I look at that as an opportunity and design challenge to maximize resolution for the purpose of future scientific discovery. Similar to Moore’s law of exponential growth, the imaging industry has experienced the same trends, given today’’s 8K resolution and greater camera sensors.
Another reference point is with Twist labs, who have discovered how to overcome scale and quality limitations through in-silica transformation of a defined lab scale.
My approach was to explore how vector based graphics, defined by a series of points and splines, could be leveraged to create what is considered “infinite resolution” or at the very least, scalable and adjustable to meet the target output.
SVG, or “Scalable Vector Graphics” are the source of my BioArt for this activity. The entire library of icons we use in this Markdown format is a good example of what’s possible!
I used Claude Ai to explore a web-friendly code base that would allow me to generate the key value pairs needed to script a Python function in the Opentrons protocol. The React/JS framework made it possible to design a User Interface (Ui) that allows for selection of any SVG, to render a resolution independent sample to the screen.
Dynamic features include assignment of a Color from an available list, increase in “Pitch” which is the number of points that are spaced along the computed line segments. Most importantly, is “Radius” which includes a value for uL, which relates to the size of a droplet in OpenTrons. The output is a PNG for a quick visual reference, and a JSON file or Text file for future parsing. I chose a simple Copy/Paste Text field to obtain the list of x,y point pairs, for use in Python for Opentrons.
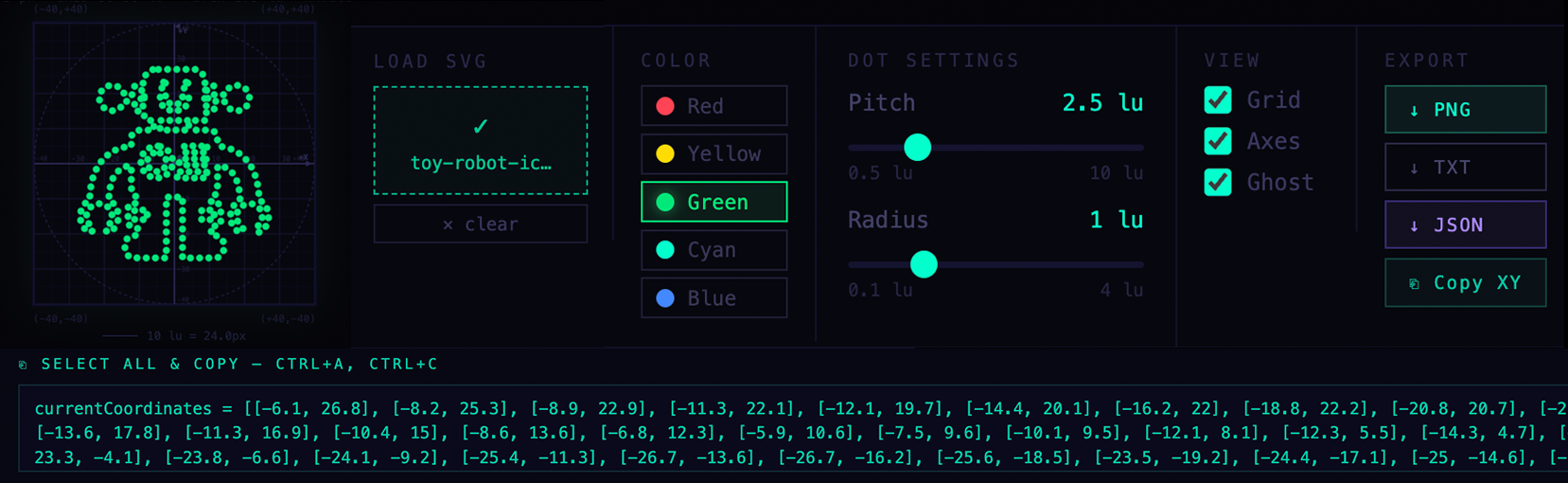
 Screenshot of SVG-to-Opentrons Converter web app by Eric Schneider
Screenshot of SVG-to-Opentrons Converter web app by Eric Schneider
I processed several sample images and ran into a slight issue with how SVG segments are deemed continuous, so I refined the parser to handle each line segment individually. I also introduced GitHub to maintain a sense of version control as a web application can quickly grow, or become corrupt, by Ai agents.
I then focused on ensuring the web application could appear inside of our preferred Colab environment, using Python and iFrame libraries. However, that is “sandboxed” and can’t share data directly. (Which is why the copy-paste is important to expedite). I tried to replicate the solution in Colab, but most things broke.
I moved on to the Opentron Simulator in Colab, with my new Data Set. I have an intermediate understanding of coding, and with the help of Claude Ai, I was able to articulate my need for a recursive list that would not only plot the points needed for pipetting, but also manage aspiration in batches of 20, not exceeding 200 uL.
After some basic Python formatting errors, I was able to preview the results via the Simulation module, and it was a very close match to my design intent.
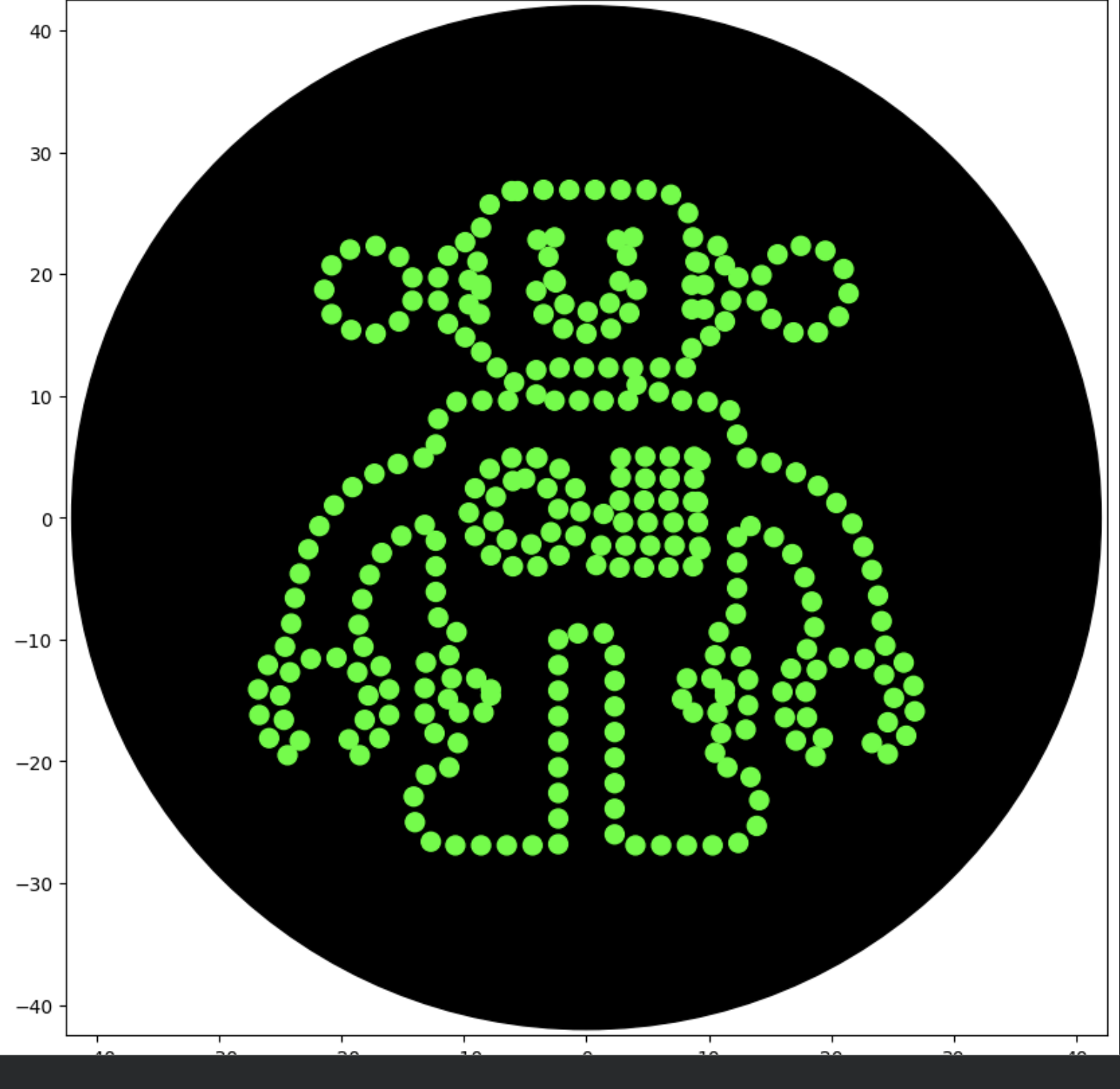

Reflection: I noticed that I was able to control the results of Vector for a high quality line that uses the full range of X, Y to the 10th of a millimeter (1 decimal point). Of course there is still the limitation of 80 mm diameter and 200 uL saturation, but I am encouraged that this technique can be refined for the purpose of high resolution design intent. I’m thinking about:
- BioCircuits that follow continual line traces for current
- BioSensors with defined sizes and shapes that are scalable
- BioArt that mirrors iconography and symbols, with dot-pitch resolution controls.
- BioPhotos that strive for incremental bitmap resolution at the microscopic level.
Imaging App- Future enhancement ideas:
- Z depth may impact Radius.
- Multiple SVG Layers, for multi-color assignments.
- Save/load to a repository
- Data sharing with Colab workspaces.
- Integration of JSON for data sharing
- Replicate application in Python in Colab natively.
- Integrate color selection into color location.
- Branching existing Automation Art code and exploring how to contribute to codebase.
OpenTrons Lab:
I was able to coordinate a working session with an OpenTrons OT-2, with Karen Ingram at the Charlotte Makerspace “BioArt Studio” which is an emerging destination for bioscience and art.
We attempted to load my protocol with vectorized points, but we encountered errors partially due to some code bugs which were quickly resolved. However, my Labware profiles were not defined for this platform configuration.
We deferred additional debugging in favor of using a known working Protocol for this session, which led to the output shown here. This is a good test since it shows the current state of functionality.
I learned how to launch and calibrate the equipment for an automated production run. I also observed an opportunity to 3D print a calibration target that would make centering the gantry over a printable art medium like watercolor paper inside of a petrie dish. We discussed a custom hold-down to keep the paper flat for more control over quality.
Our BioArt Studio session concluded with a request for a copy of a working Protocol file, so I could “reverse engineer” and configure my Protocol with the correct Labware settings. I installed a local copy of OpenTrons controller app, and was able to edit the script to include available Labware, as well as suppress the Thermal plate as it is not used in this model, and required adjustments to handling of the Z axis. Our next working session will fine-tune and test the Automation & Design protocol.
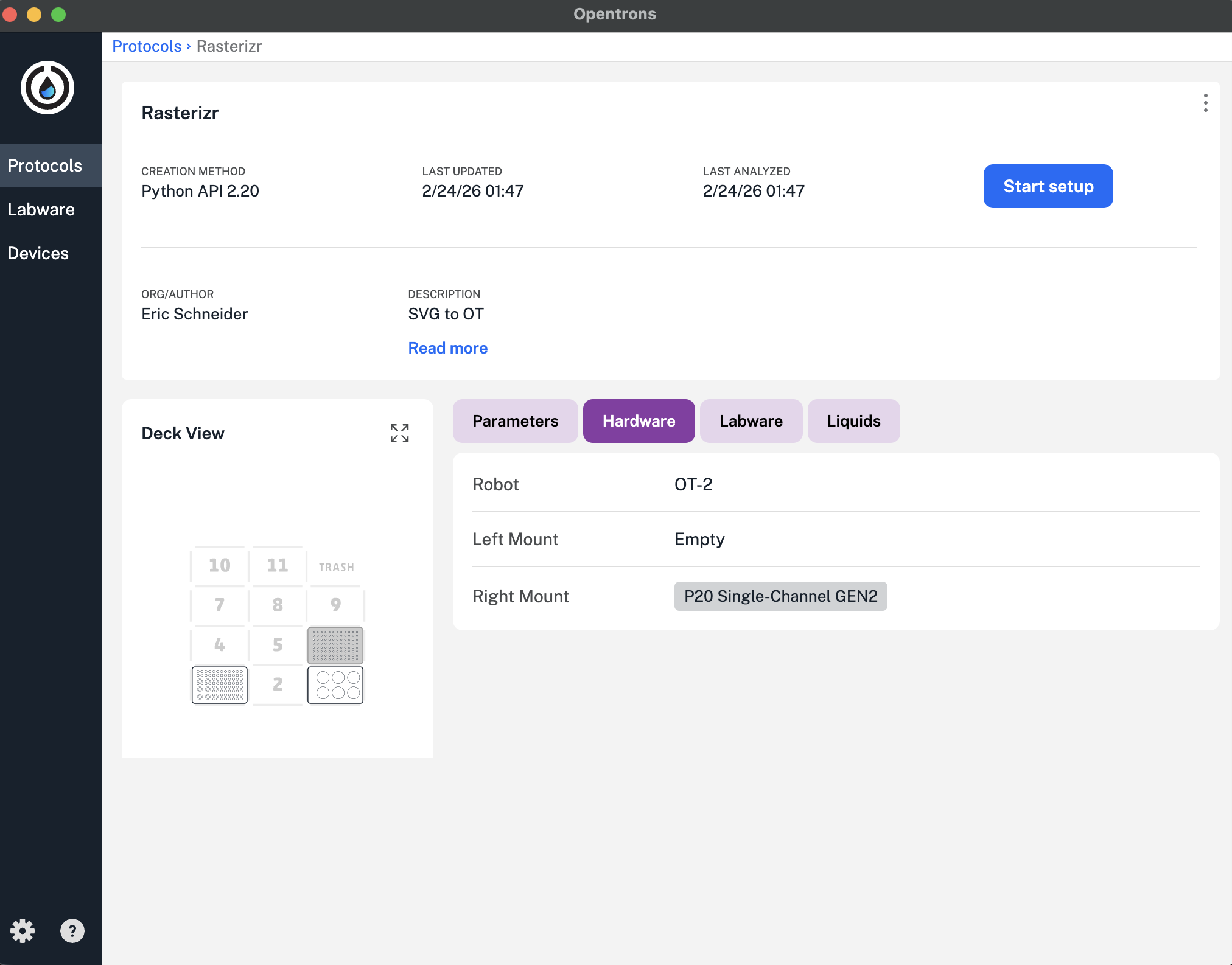

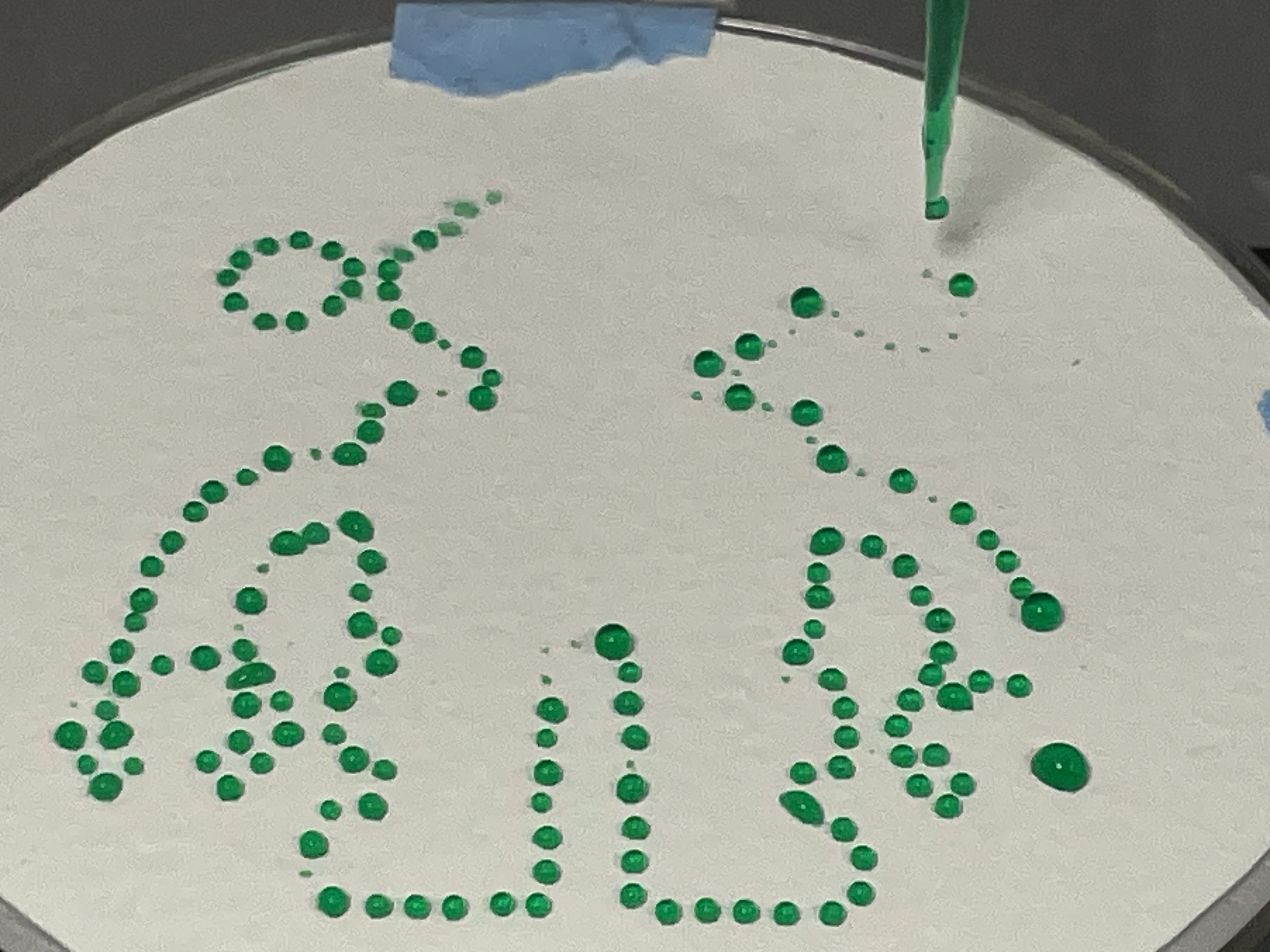
Update: 4/25/26 - The Protocol file was updated with the reassigned Labware, and was able to run the following design at 0.5uL with success:


Research Paper
I am sharing a link to an essay written by Karen Ingram, that illustrates the influence of automation on BioArt, including OpenTrons Ot-2 renderings.
https://biogeneticblooms.substack.com/p/the-blue-rose-of-metaphor-and-mystery
Be sure to browse the essays and artwork in this collection!
To learn more about the intersection of Biotechnology & Art , I am citing a research topic published by Cambridge Press. https://www.cambridge.org/engage/coe/article-details/660b2e409138d23161e8ebdf?show=item
I am excited about the field of synthetic Bioscience and Art as a result of our recent collaboration. I am grateful for the knowledge sharing and access to the BioLab.
Final Project
My Final project has been positively influenced by this week’s automation activity, as it validates that I can strive to achieve some specific lab results using the automated OpenTrons OT-2 as a tool in the process.
The path I will take for my final project starts with the identification of a Protein that can be synthesized to ensure my work is based on biotechnology best practices. The use of TWIST as a provider of automated creation of a Plasmid is the 1st step in the automation workflow.
Once I have a product, I expect to use the OpenTrons automation platform to construct a series of experiments in a host medium that will Grow into Art.
I plan on 3D printing supporting assemblies that will allow me to grow a photographic “film negative” plate, which could be a modified petrie dish that acts as a film back on a customized camera body and lighting rig.
I plan on creating a unique “exposure calibration” plate that will assist in lab test cases.
My long-range goal is to achieve a sustainable, repeatable solution that leverages automation and can scale up based on future demand for a BioPhoto “Lab” experience. I believe we are at pivotal moment in science and automation similar to when George Eastman revolutionized the photography industry through film and camera development for mass consumption. Many other industrial design solutions surround this theme.
My Final Project will reflect (and develop) artifacts of biotechnology and photography.


Checklist:
- Review this week’s recitation and this week’s lab for details on the Opentrons and programming it.
- Generate an artistic design using the GUI at https://opentrons-art.rcdonovan.com
- Write your own Python script which draws your design using the Opentrons.
- If you use AI to help complete this homework or lab, document how you used AI and which models made contributions.
- Sign up for a robot time slot if you are at MIT/Harvard/Wellesley or at a Node offering Opentrons automation.(Alt:MakerspaceCharlotte)
- Find and describe a published paper that utilizes the Opentrons or an automation tool to achieve novel biological applications.
- Write a description about what you intend to do with automation tools for your final project.
- Final Project Ideas - Submit one slide to Node