Week 7 HW: Genetic Circuits Part II: Neuromorphic Circuits
This week covers neuromorphic genetic circuits, showing how engineered gene networks can implement neural-network “perceptron”-like computation and learning.
Assignment Part 1: Intracellular Artificial Neural Networks (IANNs)
Q1. What advantages do IANNs have over traditional genetic circuits, whose input/output behaviors are Boolean functions?
Answer: IANNs have many possible responses, reflecting more of a gaussian distribution rather than binary ON/OFF outputs. This allows for gradiated, continuous range or responses versus the step-function behavior of Boolean genetic circuits, making them well-suited for environments with high levels of variability such as changing temperatures, pH, or time.
Q2. Describe a useful application for an IANN; include a detailed description of input/output behavior, as well as any limitations an IANN might face to achieve your goal.
Answer: A useful application would be when there are high levels of variability, such as changing temperatures, pH, or time. For example, if trying to express a large volume of folded proteins and the optimal conditions are changing, an IANN may refine the output by training and tuning the model based on output results. A limitation may be that the system relies on large datasets and is a measure of predictability, which may require additional downstream logic gates for boolean results.
Q3. Draw a diagram for an intracellular multilayer perceptron where layer 1 outputs an endoribonuclease that regulates a fluorescent protein output in layer 2.
Answer: In this dual region circuit:
- X1 DNA binds to a positive receptor in Region 1 (a binding site, not a proton).
- X1 also binds to a positive receptor in Region 2.
- X2 DNA is repressed in Region 1.
- X2 DNA binds in Region 2.
- Region 2 weight is greater than Region 1, producing a favorable maximum output on the Y axis
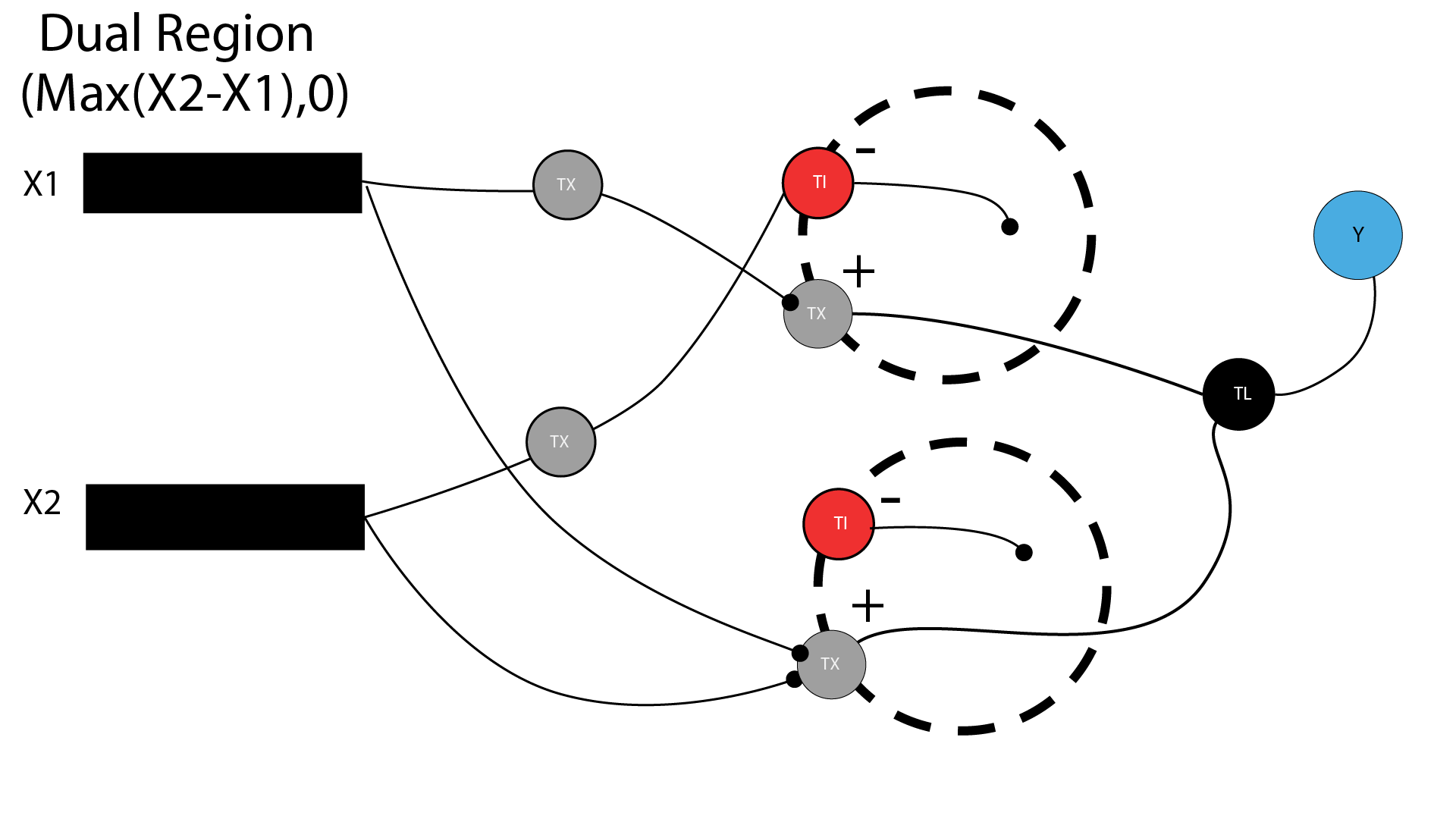
 Figure: Dual region circuit — With primary output of gFp in Layer 2.
Figure: Dual region circuit — With primary output of gFp in Layer 2.
Assignment Part 2: Fungal Materials
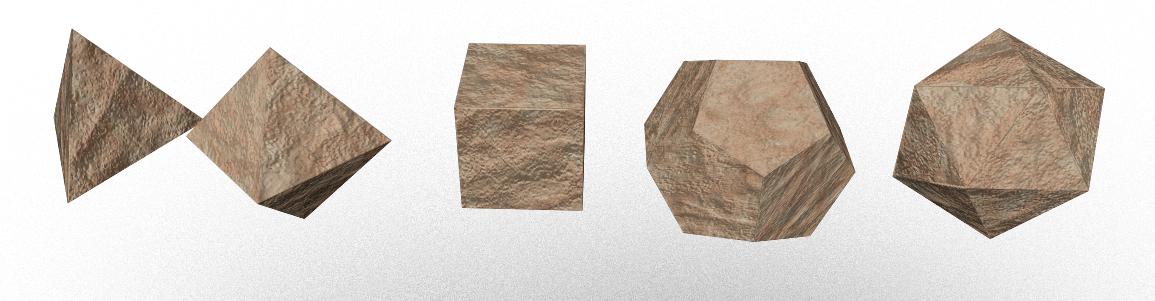

 Figure: Platonic solids cast in Mycelium; Rendering by E. Schneider - Blender 5.01
Figure: Platonic solids cast in Mycelium; Rendering by E. Schneider - Blender 5.01
Q1. What are some examples of existing fungal materials and what are they used for? What are their advantages and disadvantages over traditional counterparts?
Existing fungal materials include shaped mycelium pressed into molds, including bricks that build large structures. There are also packaging solutions, textile designs such as clothing patterns, and thin sheets which have been used as paper, as well as sculptural elements.
Some of the advantages are that the fungi can be mixed with substrates to create different material properties. There are advantages in acoustic dampening, as well as fire resistance. Some of the issues arise from the same properties; structural integrity can be compromised when drying out, causing shrinking, cracking, or breaking. Contamination is an issue when producing raw materials. The materials are inherently biodegradable which is dependent on how the substrate is manufactured and post-processed.
Q2. What might you want to genetically engineer fungi to do and why? What are the advantages of doing synthetic biology in fungi as opposed to bacteria?
I am very interested in genetically engineering fungi to create co-cultures that gain the benefit of both systems. Being able to grow fungi in a controlled environment will accelerate production of selected strains needed for specific design concepts.
The ability to engineer synthetic biology with fungi means that we may introduce attributes like color-changing materials, contamination and anti-fungal resistance, structural and tensile integrity, and novel therapeutic solutions such as drug delivery substrates; all at a larger scale than bacteria alone may provide.

 Figure: 3D Printer preparation for platonic solid forms for Mycelium casting.
Figure: 3D Printer preparation for platonic solid forms for Mycelium casting.
Assignment Part 3: First DNA Twist Order
For my final project, I seek to create a photographic image on a flood plate, using custom built labware to project a high resolution image. My DNA Twist Order for colonal gene is note below (1B), with a parallel track to order a dual plasmid construct from Addgene (1A)
Aim 1(a) has a control experiment, which will use the light projection labware I am designing to produce bacterial photographs. By requesting two existing, well-studied plasmids from Addgene, I may be able to expedite the Design, Build, Test and Analyze(Learn) cycle.
The following genes have been contributed by Tabor Labs, and can be ordered as standard bacterial stabs.
- pSR58.6 (Plasmid #63176) which expresses CcaR constitutively and sfGFP under the PcpcG2-172 promoter
- pSR43.6r (Plasmid #63197) which expresses CcaS, Ho1, and PcyA constitutively.
Note: This order selection will be reviewed and validated to confirm if this is the optimal candidate for this project. There are several versions, each with increasing on/off folding range, reaching >100X for high contrast, high quality visualization
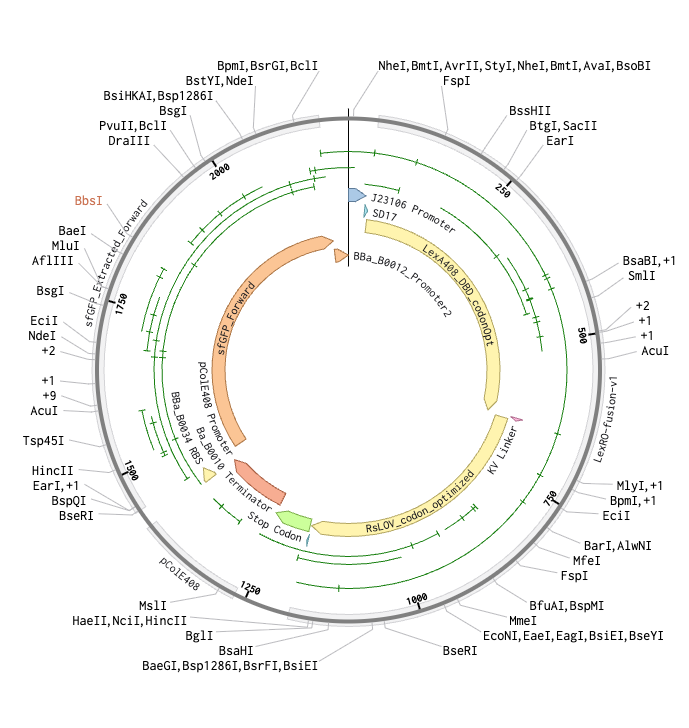
Aim 1(b) is to submit a TWIST order of an engineered E. coli with a single-plasmid optogenetic system ( derived from eLightOn) that activates sfGFP expression in response to 470 nm blue light, producing a photographic image in green fluorescence on a flood plate using my projection labware.
- The link to my shared Benchling file is found in the class submittal via Google Form.

Citations
Aim 1A — CcaSR Green Light System (pSR43.6r + pSR58.6)
Primary system citation: Schmidl SR, Sheth RU, Wu A, Tabor JJ. Refactoring and optimization of light-switchable Escherichia coli two-component systems. ACS Synthetic Biology. 2014 Nov 21;3(11):820–31. doi: 10.1021/sb500273n. PMID: 25305428.
Plasmid citations (Addgene):
- pSR43.6r was a gift from Jeffrey Tabor (Addgene plasmid #63197; http://n2t.net/addgene:63197; RRID:Addgene_63197)
- pSR58.6 was a gift from Jeffrey Tabor (Addgene plasmid #63176; http://n2t.net/addgene:63176; RRID:Addgene_63176)
Foundational bacterial photography citation: Levskaya A, Chevalier AA, Tabor JJ, Simpson ZB, Lavery LA, Levy M, Davidson EA, Scouras A, Ellington AD, Marcotte EM, Voigt CA. Synthetic biology: engineering Escherichia coli to see light. Nature. 2005 Nov 24;438(7067):441–2. doi: 10.1038/nature04405. PMID: 16306981.
Aim 1B — eLightOn Blue Light System (pBioLight-1B-eLightOn-v1)
Primary eLightOn system citation: Li X, Zhang C, Xu X, Miao J, Yao J, Liu R, Zhao Y, Chen X, Yang Y. A single-component light sensor system allows highly tunable and direct activation of gene expression in bacterial cells. Nucleic Acids Research. 2020 Apr 6;48(6):e33. doi: 10.1093/nar/gkaa044. PMID: 31989175. PMC: PMC7102963.
sfGFP sequence source (extracted from pJT119b for E. coli codon-optimized sequence only): Olson EJ, Hartsough LA, Landry BP, Shroff R, Tabor JJ. Characterizing bacterial gene circuit dynamics with optically programmed gene expression signals. Nature Methods. 2014 Mar 9;11(4):449–55. doi: 10.1038/nmeth.2884. PMID: 24608181.
- pJT119b was a gift from Jeffrey Tabor (Addgene plasmid #50551; http://n2t.net/addgene:50551; RRID:Addgene_50551)
RsLOV structural reference: Conrad KS, Bilwes AM, Crane BR. Light-induced subunit dissociation by a light-oxygen-voltage domain photoreceptor from Rhodobacter sphaeroides. Biochemistry. 2013 Jan 15;52(2):378–91. doi: 10.1021/bi3015373. PMID: 23252338. PMC: PMC3582384. PDB: 4HJ6.
LexA408 mutation reference: Little JW, Edmiston SH, Pacelli LZ, Mount DW. Cleavage of the Escherichia coli LexA protein by the RecA protease. Proceedings of the National Academy of Sciences. 1980;77(6):3225–9. doi: 10.1073/pnas.77.6.3225. PMID: 6251456.
iGEM Registry Parts
- BBa_J23106 — Constitutive promoter (Anderson family). iGEM Registry. https://parts.igem.org/Part:BBa_J23106
- BBa_B0034 — Ribosome binding site (Elowitz 1999 repressilator). iGEM Registry. https://parts.igem.org/Part:BBa_B0034
- BBa_B0010 — rrnB T1 transcription terminator. iGEM Registry. https://parts.igem.org/Part:BBa_B0010
- BBa_B0012 — rrnB T2 transcription terminator. iGEM Registry. https://parts.igem.org/Part:BBa_B0012
Destination Backbone
pUC19 backbone — Addgene #50005. NCBI accession: L09137. Specified as clonal plasmid destination vector for Twist Bioscience synthesis order.