Week 12 HW: Building Genomes
HTGAA Week 12 Homework
Part A: The 1,536 Pixel Artwork Canvas | Collective Artwork
Item 1: Pixel Contribution

I contributed to plate #G3, initiating a rose design on April 15. I seeded the concept on Discourse: "#G3 - Starting to build a rose… let’s see what grows!"
Item 2: HTGAA Webpage Notes
2a — What I contributed:
I contributed to plate #G3, planting the seed of a pixel rose and inviting others to build on it — fully expecting it to be overwritten, in the spirit of collaborative design.
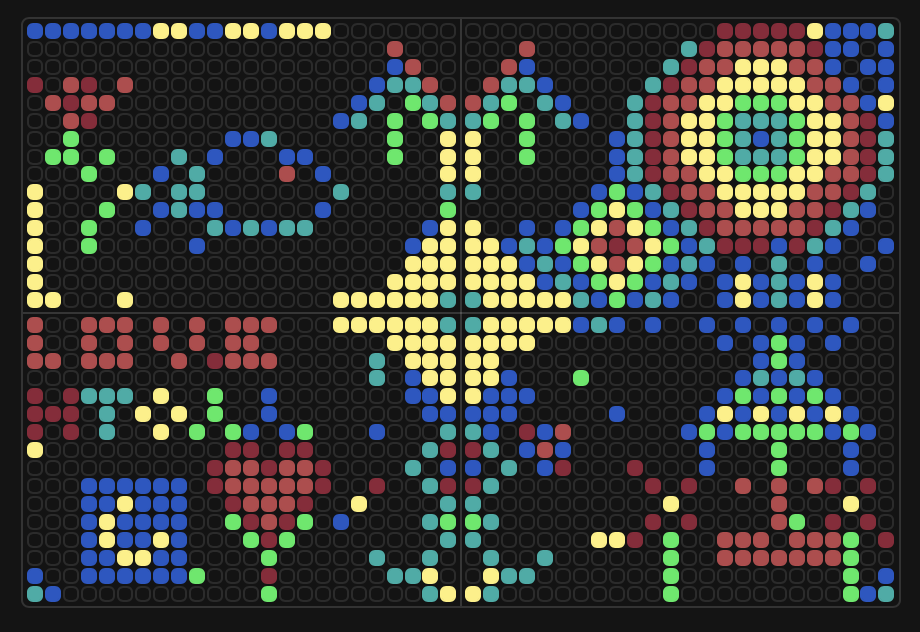



2b — What I liked:
I liked how the canvas was a collaborative activity. It’s interesting how quickly it became an algorithmic scripted design. The timeline feature is a good addition, to be able to see the evolution over time.
2c — What could be made better:
I have been thinking of how to introduce more collaboration and team building by creating a challenge that rewards the player with pixels, or points that can be used to build with. While scripted algorithms are great, there could be a separate or individual board space for that. To take the game concept one step further, imagine the Tron Light Cycle game. The goal is to traverse the board — without crossing over another trail. Pick anywhere on an edge to begin, and see if you can make it to the middle. More earned “points” = more chances to move. Earn points by quiz questions, or scavenger hunt activities. Roll a random # and that’s how many “moves” you get.
Part B: Cell-Free Protein Synthesis Reaction Composition
Component Descriptions
1. E. coli Lysate — BL21 (DE3) Star Lysate (includes T7 RNA Polymerase)
The lysate contains all of the cellular building blocks needed to carry out transcription and translation outside of the cell. T7 polymerase is selected for its known capability of recognizing the T7 promoter sequence in the cell-free system design.
2. Potassium Glutamate
Potassium Glutamate has the most volume of salt in this buffer, and its role is to manage the ionic binding strength.
3. HEPES-KOH pH 7.5
Its main role in the buffer is to stabilize pH.
4. Magnesium Glutamate
Mg²⁺ is an important element in maintaining ribosome structure. It is highly tunable in order to promote ribosome function, without impacting other components.
5. Potassium Phosphate Monobasic & Dibasic
The potassium phosphate is primarily an energy source, that also helps to maintain pH. The key is the ratio between the two to raise/lower pH respectively.
6. Ribose
Ribose provides energy in the transcription process. It is a sugar-based component of fuel.
7. Glucose
Glucose, along with ribose, create an energy system to help drive regeneration of ATP, and works well over long timeframes.
8. AMP, CMP, GMP, UMP (Nucleoside Monophosphates)
These are the consumable building blocks that are considered substrates for RNA synthesis. They are recycled and sustainable over longer periods of time.
9. Guanine
Guanine provides the energy for GTP. It is important in that it prolongs the cycle of ribosome translation.
10. 17 Amino Acid Mix
These are the essential building blocks of protein synthesis, and will allow tRNA to pair with mRNA in the ribosomes, based on transcription information.
11. Tyrosine
It is another of the essential amino acids for tRNA to mRNA translation. It is handled uniquely due to its poor solubility.
12. Cysteine
Cysteine is an essential amino acid, and it is added separately due to its unstable chemical bonding properties, that prevent it from mixing with other elements. It is important for structural development in protein folding.
13. Nicotinamide
Nicotinamide is a stabilizer that supports the energy system found in metabolic enzymes.
14. Nuclease Free Water (Backfill)
As a backfill, it is free of contamination to allow the defined mix to perform at full potential.
Question 1: 1-Hour PEP-NTP vs. 20-Hour NMP-Ribose-Glucose Master Mix
The main difference between the 1-hour and 20-hour mix is in the composition of the master mix. The 20-hour incubation contains additional Ribose and Glucose to provide energy that allows translation to sustain the longer time frame. The 1-hour mix uses spermidine to help accelerate the translation, with short-term stabilization of nucleic acids. In contrast, the 20-hour contains nicotinamide to stabilize the enzymes. HEPES-KOH has a higher level to mitigate the higher pH of the concentration. In the one-hour mix, PEP (phosphoenolpyruvate) + NTPs is the energy source, which is fast and high-yield but generates inorganic phosphate buildup that inhibits the reaction, limiting it to ~1 hour.
Question 2 (Bonus): How Can Transcription Occur if GMP is Not Included but Guanine Is?
Guanine provides the energy for GTP. It is important in that it prolongs the cycle of ribosome translation. Guanine base can substitute for GMP — the lysate contains the enzymatic machinery to convert it.
Part C: Fluorescent Proteins in Cell-Free Systems
Question 1: Biophysical and Functional Properties
| Protein | Answer | Supplemental Notes |
|---|---|---|
| sfGFP | sfGFP is well suited for cell-free systems as it has a proven high rate of folding ability, and has a fast chromophore response, needed for fast fluorescence | Folds without cellular chaperones; chromophore maturation still requires O₂ |
| mRFP1 | Slowly-maturing monomer with low acid sensitivity; requires more time and has low pH sensitivity which inhibits maturation | Primary limitation is slow maturation + low quantum yield (~0.25); pH is not the limiting factor at pH 7.5 |
| mKO2 | mKO2 has moderate pH sensitivity so it may be inhibited by higher pH; slow to mature which will limit fluorescence yield | Maturation half-time ~180 min vs sfGFP ~30 min; O₂ dependence also limits yield in sealed reactions |
| mTurquoise2 | Rapidly-maturing monomer with very low acid sensitivity; medium expression with high brightness | Brightness driven by exceptionally high quantum yield (~0.93); maturation is moderate not rapid |
| mScarlet-I | Fast maturation and high brightness, with moderate acid sensitivity | Maturation half-time ~40 min; among best red FP choices for CFPS; “I” = intermediate brightness/speed tradeoff |
| Electra2 | Oxygen-independent and requires an exogenous reagent; unique in the low waveform spectrum | Exogenous cofactor is biliverdin, must be added to mastermix; emission is near-infrared (~670nm), high not low wavelength |
Question 2: Mastermix Hypothesis
Protein: sfGFP — selected for its direct relevance to BioLight Aim 2, a cell-free biosensor that expresses sfGFP when exposed to ~470nm blue light.
Reagents: Mg²⁺ availability, adjusted via two mechanisms operating on a single mechanistic axis:
A. Potassium Phosphate Dibasic (Mg²⁺ sink)
Added above the baseline 5.625mM, potassium phosphate dibasic sequesters free Mg²⁺ through phosphate chelation, reducing ribosome assembly efficiency and T7 RNAP cofactor availability — driving sfGFP expression below the control baseline.
B. Magnesium Glutamate (Mg²⁺ source)
Added above the baseline 6.975mM, magnesium glutamate directly increases free Mg²⁺ in solution, stabilizing ribosome subunit assembly and activating Mg-NTP complexes for both transcription and translation — driving sfGFP expression above the control baseline.
Expected effect: Since both reagents interact with ionic affinity to the ribosome, the focus is on a singular variable for fine tuning in both directions from the master mix as control. The relative magnitude of the positive and negative deltas from control, measured by spectrophotometry at excitation 485nm / emission 510nm, will reveal whether the master mix is operating below, at, or above its Mg²⁺ optimum — directly informing optimized reaction conditions for eLightOn CFPS deployment in BioLight x2 Aim 2.
Question 3: Reagent Concentrations
Protein: sfGFP Hypothesis: v2.0 — Mg²⁺ single-axis variable luminosity construct Measurement: Spectrophotometry at excitation 485nm / emission 510nm
| Well | Reagent | Adjusted Final Concentration | Predicted RFU | Delta from Control |
|---|---|---|---|---|
| W1 | KPO₄ dibasic | 15mM | ~800 | −2,700 |
| W2 | KPO₄ dibasic | 11mM | ~1,400 | −2,100 |
| W3 | KPO₄ dibasic | 8mM | ~2,200 | −1,300 |
| W4 | Control | 6.975mM | ~3,500 | — |
| W5 | Control | 6.975mM | ~3,500 | — |
| W6 | MgGlu | 9mM | ~4,800 | +1,300 |
| W7 | MgGlu | 11mM | ~6,200 | +2,700 |
| W8 | MgGlu | 13mM | ~7,800 | +4,300 |
Predicted dynamic range: ~10× from W1 to W8. The asymmetry between negative delta (−2,700 max) and positive delta (+4,300 max) suggests the master mix is operating below its Mg²⁺ optimum — more headroom above baseline than below.
Connection to BioLight x2:
| Aim | Title | Dependency |
|---|---|---|
| Aim Zero | CFPS calibration | This experiment |
| Aim 1 | Wetlab validation | Independent — running in parallel |
| Aim 2 | Cell-free + imaging platform | Inherits Aim Zero predictive model |
| Aim 3 | Makerspace deployment | Inherits Aim 2 validated protocol |
Question 4: Fluorescence Data Analysis
Pending — due one week after data is returned (date TBD).
Part D: Build-A-Cloud-Lab (Bonus)


What I Built
For the Cloud Lab render, I used the default assembly, but focused on an idea for a circular floor plan, to introduce the idea of a circular manufacturing layout. While linear production lines are expected, there may be some spatial advantages to a circular layout, to fit more equipment in less space, with a central pivot point for automated transfer. I added a visualization of a data wall, where all activity and data is monitored from a command center or virtual reality heads-up display (which would correspond well with a circular layout — no linear navigation required).
What I Liked
I liked the idea of a visual configurator that leads the way towards a “digital twin” of the lab, where protocols can be designed, built, and simulated.
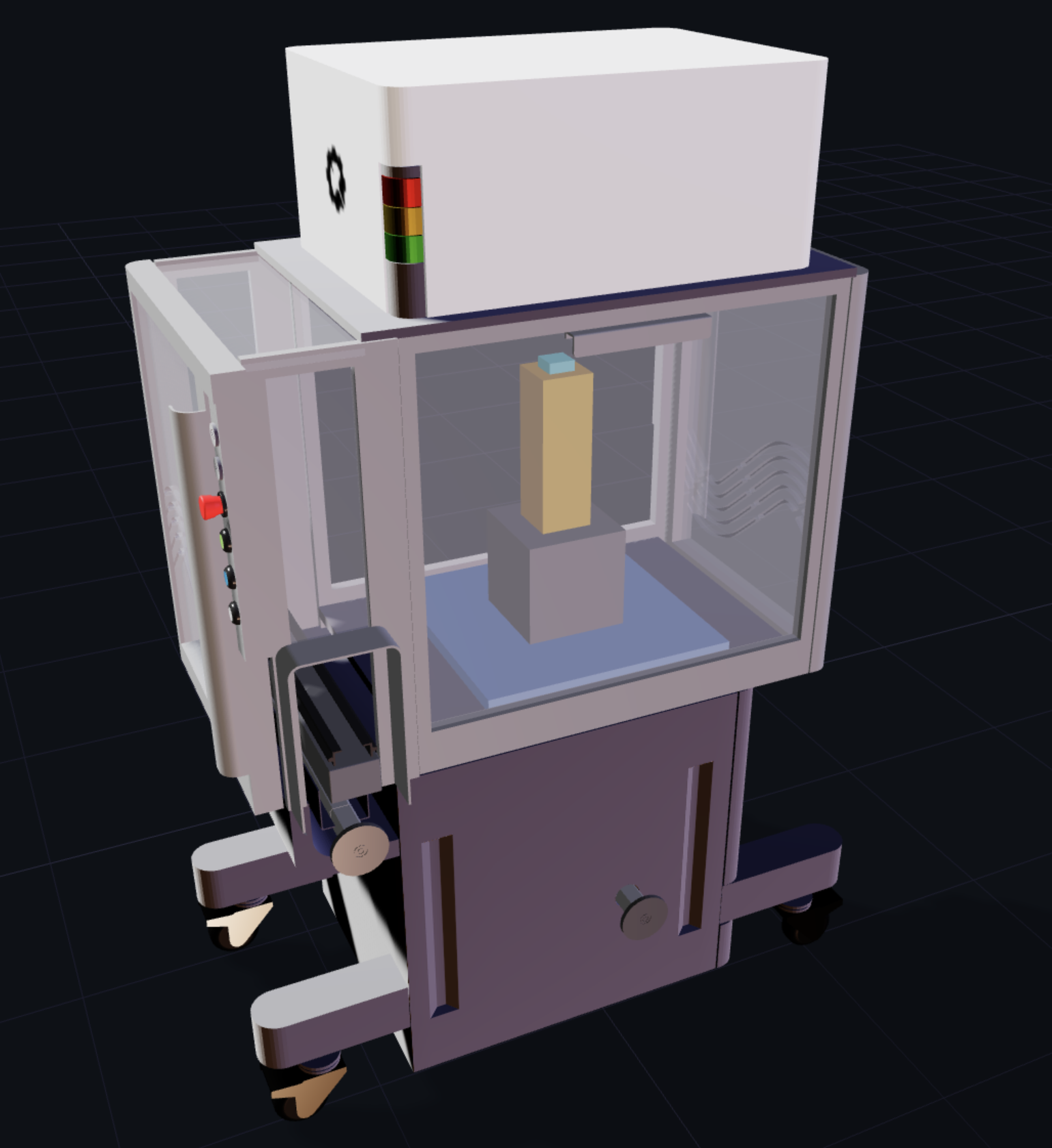


Design Ideas
I would like to be able to import my own custom labware, and apply transforms such as x, y, z position and scale within a unit. That would lead to connectivity with the API to perform animated sequences, true to the intent of remote automation.
Appendix: Scoring Summary
Part A
| Item | Score |
|---|---|
| A1 — Pixel contribution | 9/10 |
| A2a — What I contributed | 9/10 |
| A2b — What I liked | 8/10 |
| A2c — What could be improved | 9/10 |
| Part A overall | 8.75/10 |
Confidence: ± 6%
Part B — Component Scores
| Component | Initial | Final | Confidence |
|---|---|---|---|
| E. coli Lysate | 6/10 | 8/10 | 8/10 |
| Potassium Glutamate | 5/10 | 7/10 | 8/10 |
| HEPES-KOH | 8/10 | 8/10 | 8/10 |
| Magnesium Glutamate | 5/10 | 7/10 | 8/10 |
| Potassium Phosphate | 3/10 | 7/10 | 6/10 |
| Ribose | 4/10 | 6/10 | 7/10 |
| Glucose | 6/10 | 8/10 | 8/10 |
| AMP/CMP/GMP/UMP | 4/10 | 7/10 | 7/10 |
| Guanine | 4/10 | 7/10 | 8/10 |
| 17 AA Mix | 7/10 | 8/10 | 9/10 |
| Tyrosine | 3/10 | 8/10 | 6/10 |
| Cysteine | 6/10 | 9/10 | 8/10 |
| Nicotinamide | 3/10 | 7/10 | 8/10 |
| Nuclease Free Water | 6/10 | 7/10 | 7/10 |
| B-Q1: PEP vs NMP | 7/10 | 9/10 | 9/10 |
| B-Q2: Bonus | 5/10 | 7/10 | 6/10 |
| Average | 5.3/10 | 7.6/10 | 7.5/10 |
Part C — Scores
| Item | Initial | Final | Confidence |
|---|---|---|---|
| sfGFP | 5/10 | 8/10 | 8/10 |
| mRFP1 | 6/10 | 6/10 | 8/10 |
| mKO2 | 4/10 | 6/10 | 6/10 |
| mTurquoise2 | 6/10 | 6/10 | 9/10 |
| mScarlet-I | 4/10 | 8/10 | 7/10 |
| Electra2 | 3/10 | 7/10 | 4/10 |
| Q2 Hypothesis | 8/10 | 9/10 | 9/10 |
| Average | 5.1/10 | 7.1/10 | 7.3/10 |
Part D — Scores
| Item | Score |
|---|---|
| Participation | 10/10 |
| Design rationale | 9/10 |
| What I liked | 9/10 |
| Design Ideas | 9/10 |
| Overall | 9.25/10 |
Overall Scorecard (All Parts)
| Section | Final Accuracy |
|---|---|
| Part A | 8.75/10 |
| Part B | 7.6/10 |
| Part C | 7.1/10 |
| Part D | 9.25/10 |
| Overall | 8.2/10 |
| Metric | Score |
|---|---|
| Final accuracy | 8.2 / 10 |
| Competency score | 8.4 / 10 |
| Average improvement (initial → final) | +2.2 pts |
| Confidence calibration gap | ± 8% |
| Self-correction rate | 88% |
Strengths: Part D design thinking, Part B Q1, Cysteine, mScarlet-I, sfGFP Recurring gap: pH framing consistently inverted; molecular mechanism specificity (enzyme names, pathways, quantum yield)
Score Visualizations
Chart 1 — accuracy & confidence progress
Chart 2 — accuracy improvement delta per item
Chart 3 — overall scorecard
Document complete pending Part C Q4 data return.