Week 3 HW: Lab Automation
Homework
Assignment: Python Script for Opentrons Artwork
Your task this week is to Create a Python file to run on an Opentrons liquid handling robot.
- Generate an artistic design using the GUI at opentrons-art.rcdonovan.com.
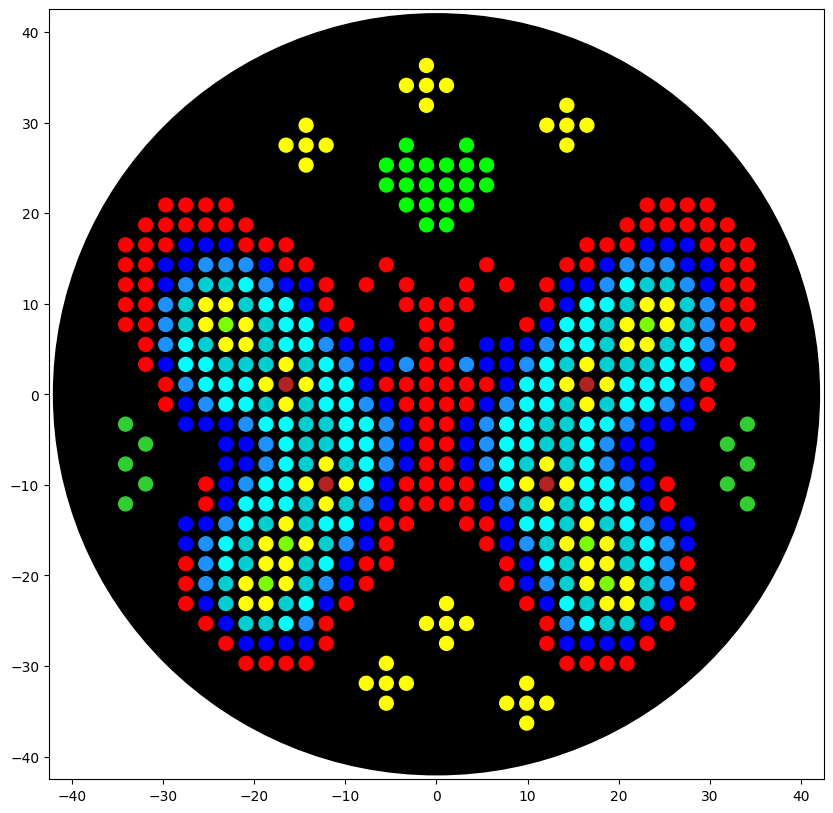
For my first design I made a colorful butterfly! I first used the Opentrons art page to design it by using the upload image option. Initially the design
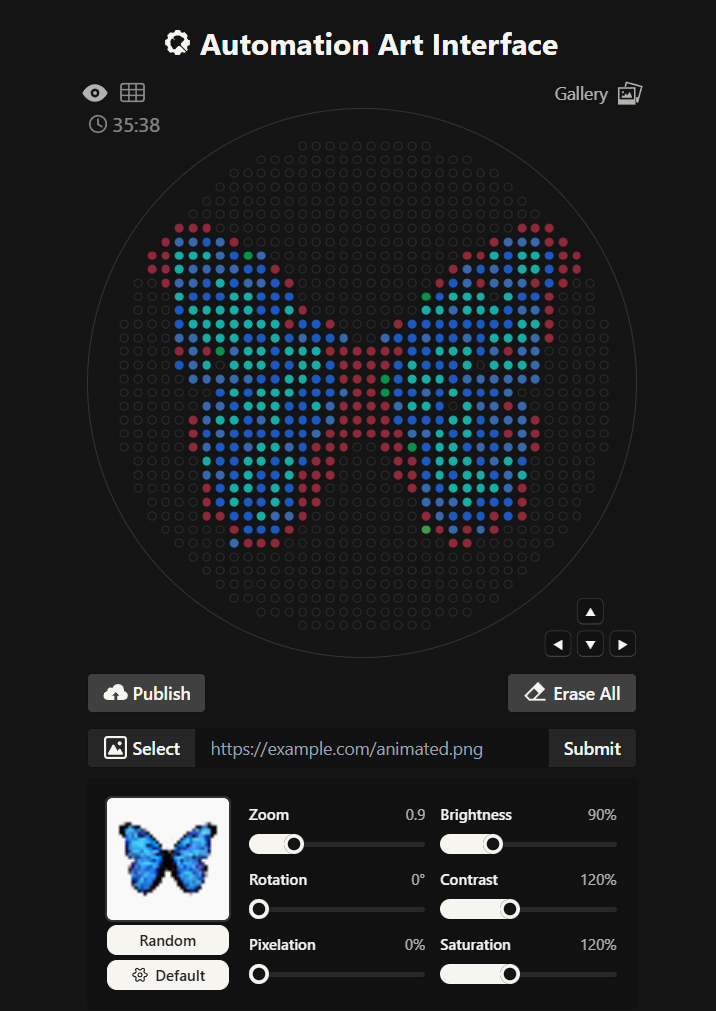

Here you can see the butterfly image that I uploaded and how it generates on the Opentrons art page side by side!
Then after the image upload, I decided to first move the design a bit lower and also change the colors. Lastly I added some fun details like stars and a heart.
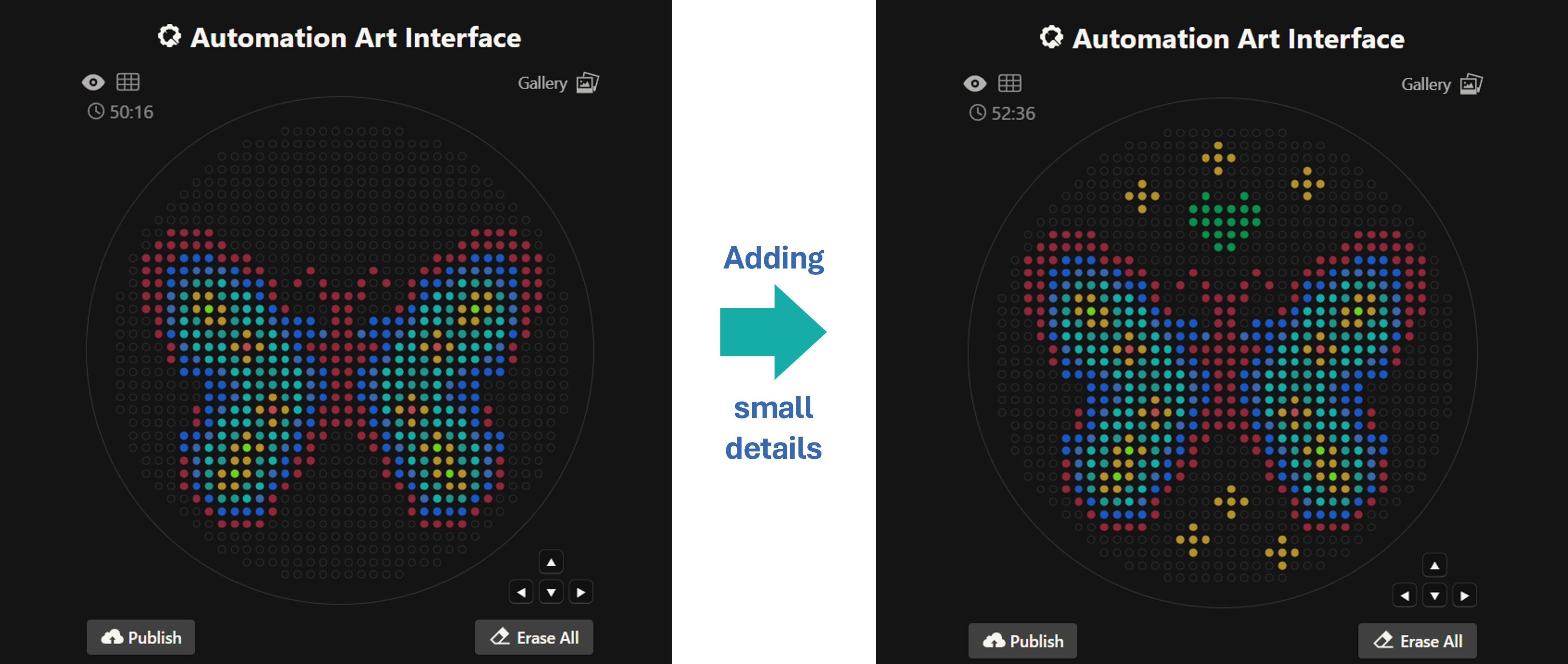

Design process
Here’s the final design!


This is my design: a colorful butterfly! Made using the GUI. You can check it out by yourself here!
Initially, I made one artistic design on a circular petri dish, but after finding out you could make designs on a rectangular plate, I decided to try it out! I ended up making 2 more designs on rectangular plates.
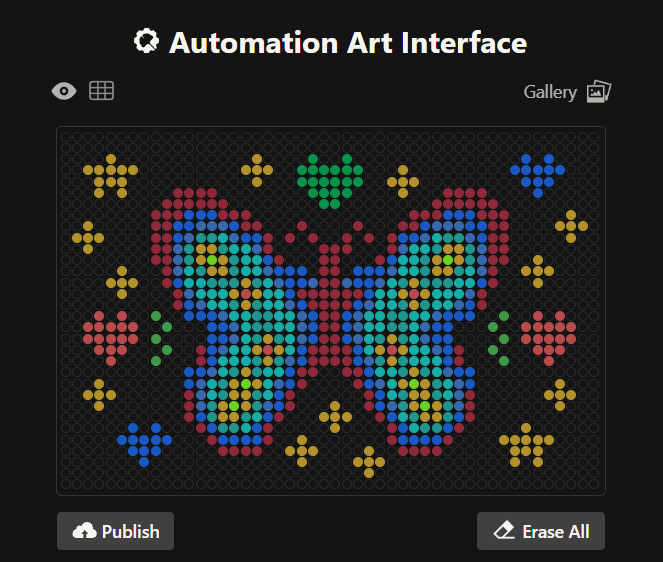

This is my second design which is a readaptation of the colorful butterfly! Made using the GUI. You can check it out by yourself here!
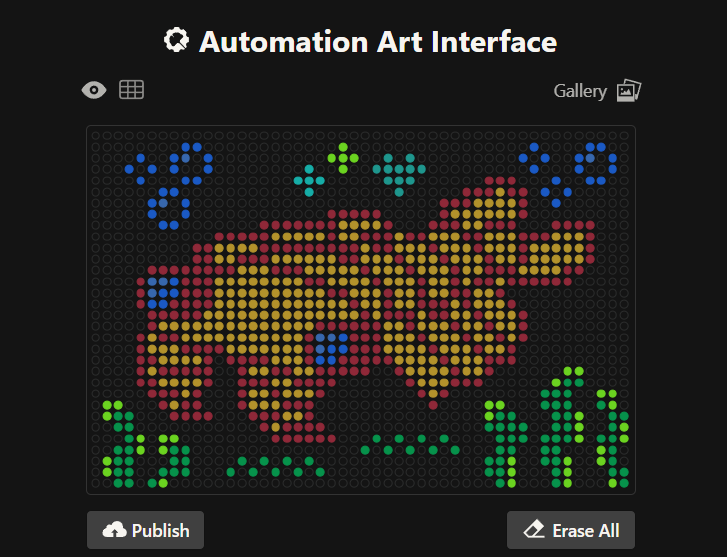

This is my third design: an anomalocaris! Made using the GUI. You can check it out by yourself here!
- Using the coordinates from the GUI, follow the instructions in the HTGAA26 Opentrons Colab to write your own Python script which draws your design using the Opentrons.
- You may use AI assistance for this coding — Google Gemini is integrated into Colab (see the stylized star bottom center); it will do a good job writing functional Python, while you probably need to take charge of the art concept.
- If you’re a proficient programmer and you’d rather code something mathematical or algorithmic instead of using your GUI coordinates, you may do that instead.

Here’s the Opentrons Lab Simulation in Google Colab for the first design. You can check it out by yourself here!
Post-Lab Questions — DUE BY START OF FEB 24 LECTURE
One of the great parts about having an automated robot is being able to precisely mix, deposit, and run reactions without much intervention, and design and deploy experiments remotely.
For this week, we’d like for you to do the following:
- Find and describe a published paper that utilizes the Opentrons or an automation tool to achieve novel biological applications.
Paper: Automation of protein crystallization scale-up via Opentrons-2 liquid handling.
This paper explores the use of the Opentrons OT-2 machine to automate protein crystallization! The researchers developed three Python scripts using the Opentrons Python module to control the robot for mixing and setting up 24-well sitting drop plates using model proteins like lysozyme and a periplasmic protein from Campylobacter jejuni.
The study achieved the desired scale-up goals after minimal trial and error. By automating the liquid handling, the researchers were able to test a wider range of crystallization conditions (reagents, concentrations, and pH) with higher reproducibility than manual pipetting. Although the setup time was around 35 to 40 minutes, it greatly reduces plate variability from person to person. This is a novel application because it makes high-quality structural biology workflows accessible and low-cost, allowing labs to screen protein conditions at a much higher throughput, which is essential for understanding protein function and drug design.
Reference: DeRoo, J. B., Jones, A. A., Slaughter, C. K., Ahr, T. W., Stroup, S. M., Thompson, G. B., & Snow, C. D. (2025). Automation of protein crystallization scaleup via Opentrons-2 liquid handling. SLAS Technology, 32, 100268. https://doi.org/10.1016/j.slast.2025.100268
- Write a description about what you intend to do with automation tools for your final project. You may include example pseudocode, Python scripts, 3D printed holders, a plan for how to use Ginkgo Nebula, and more. You may reference this week’s recitation slide deck for lab automation details.
While your description/project idea doesn’t need to be set in stone, we would like to see core details of what you would automate. This is due at the start of lecture and does not need to be tested on the Opentrons yet.
For instance, my first idea is an interactive 3D bio-art installation that translates a person’s biological data into a living and blooming sculpture. This idea uses genetically engineered bacteria to create a visual representation of a user’s unique microbial/DNA fingerprint. The process starts when a user interacts with a sensor that captures basic biological data, which is then processed by a script to assign specific colors using fluorescent proteins like GFP, RFP, and BFP. In this case, an Opentrons OT-2 acts as a high-precision bio-printer to deposit these living bio-inks into a 3D-printed scaffold made of agar or hydrogel, allowing the sculpture to grow and glow over time to reveal the user’s identity.
Additionally, I will need to design 3D-printed holders with micro-channels and a specialized needle adapter so the OT-2 can deposit the bacteria without breaking the hydrogel/agar structure. I will use capacitive touch sensors to generate the initial data that determines the bacterial/DNA distribution throughout the sculpture. Moreover, I plan to use cloud laboratories like Ginkgo Nebula to synthesize the custom DNA circuits needed to ensure the bacteria express the exact colors and intensity required for the piece.
Here’s a rough python pseudocode for this 3D sculpture idea.
Final Project Ideas — DUE BY START OF FEB 24 LECTURE
As explained in this week’s recitation, add a slide in your Node’s section of this slide deck with an idea you have for an Individual Final Project. Be sure to put your name on your slide!
Here are my three individual final project ideas!
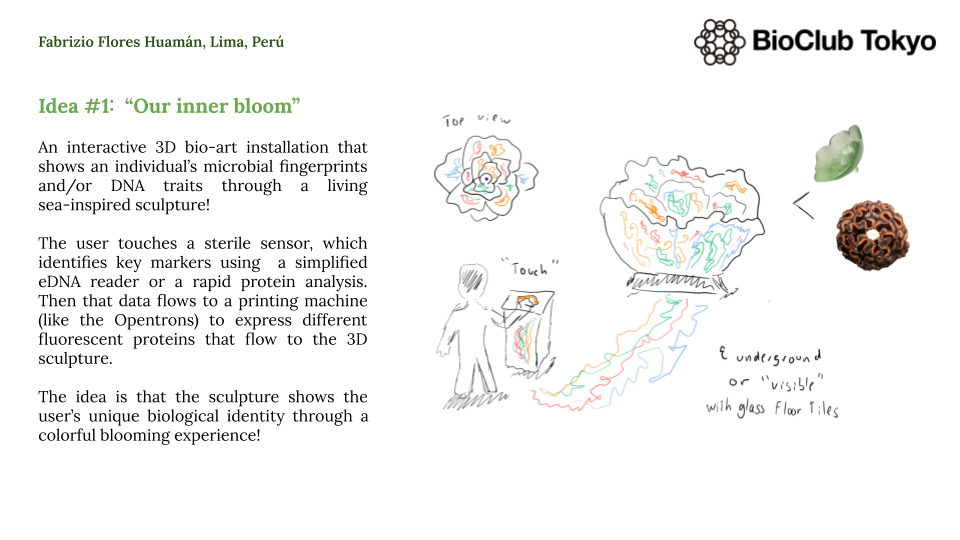

An interactive 3D bio-art sculpture
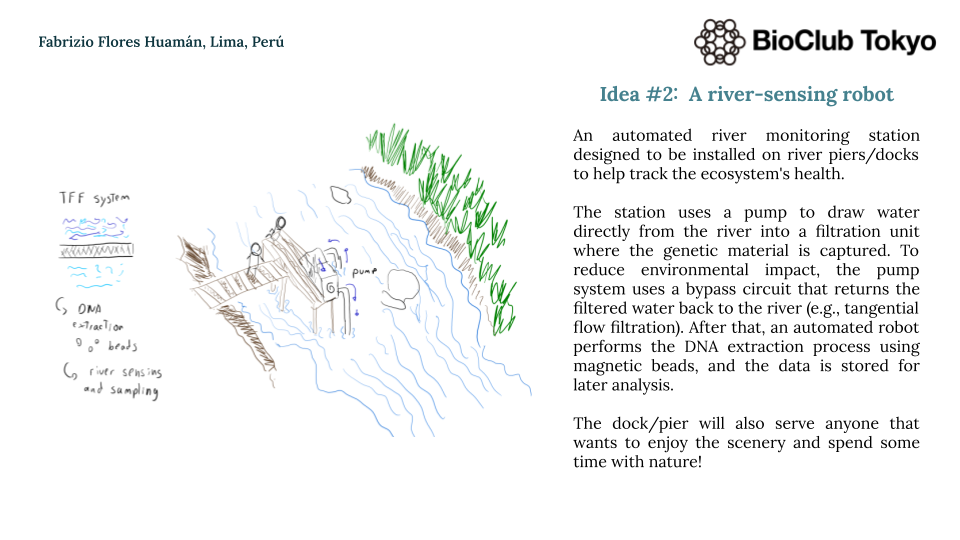


A river-sensing automated robot system
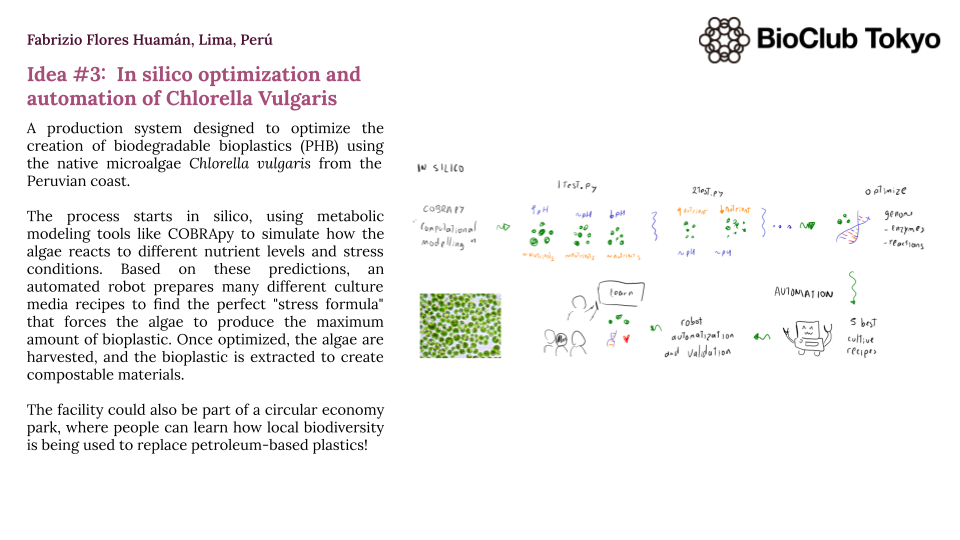


Chlorella vulgaris in silico optimization and automation