Week 5 HW: Protein Design - Part II
Homework
Part A: SOD1 Binder Peptide Design (From Pranam)
Superoxide dismutase 1 (SOD1) is a cytosolic antioxidant enzyme that converts superoxide radicals into hydrogen peroxide and oxygen. In its native state, it forms a stable homodimer and binds copper and zinc.
Mutations in SOD1 cause familial Amyotrophic Lateral Sclerosis (ALS). Among them, the A4V mutation (Alanine → Valine at residue 4) leads to one of the most aggressive forms of the disease. The mutation subtly destabilizes the N-terminus, perturbs folding energetics, and promotes toxic aggregation.
Your challenge:
- Design short peptides that bind mutant SOD1.
- Then decide which ones are worth advancing toward therapy.
You will use three models developed in our lab:
- PepMLM: target sequence-conditioned peptide generation via masked language modeling
- PeptiVerse: therapeutic property prediction
- moPPIt: motif-specific multi-objective peptide design using Multi-Objective Guided Discrete Flow Matching (MOG-DFM)
Part 1: Generate Binders with PepMLM
- Begin by retrieving the human SOD1 sequence from UniProt (P00441) and introducing the A4V mutation.
Here’s the FASTA file of the human SOD1 sequence (you can also check it out here):
>sp|P00441|SODC_HUMAN Superoxide dismutase [Cu-Zn] OS=Homo sapiens OX=9606 GN=SOD1 PE=1 SV=2 MATKAVCVLKGDGPVQGIINFEQKESNGPVKVWGSIKGLTEGLHGFHVHEFGDNTAGCTS AGPHFNPLSRKHGGPKDEERHVGDLGNVTADKDGVADVSIEDSVISLSGDHCIIGRTLVV HEKADDLGKGGNEESTKTGNAGSRLACGVIGIAQ
Also, here’s a screenshot of the original SOD1 protein!
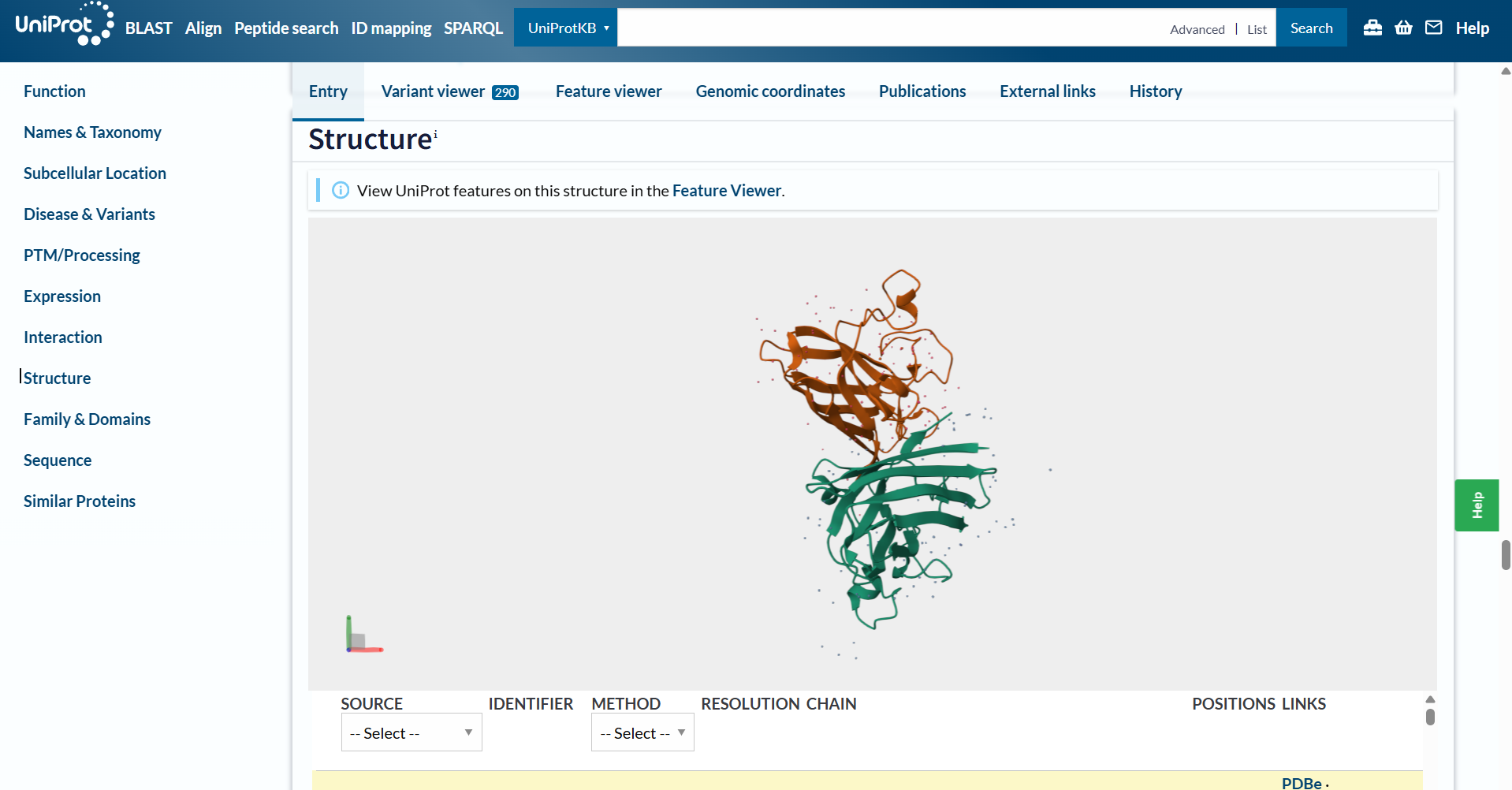

The A4V mutation would be the following:
>SOD1 A4V mutation MATKVVCVLKGDGPVQGIINFEQKESNGPVKVWGSIKGLTEGLHGFHVHEFGDNTAGCTS AGPHFNPLSRKHGGPKDEERHVGDLGNVTADKDGVADVSIEDSVISLSGDHCIIGRTLVV HEKADDLGKGGNEESTKTGNAGSRLACGVIGIAQ
- Using the PepMLM Colab linked from the HuggingFace PepMLM-650M model card:
I made my own copy of the PepMLM Colab; you can access it here: Fabrizio_Flores_PepMLM-650M.ipynb!
- Generate four peptides of length 12 amino acids conditioned on the mutant SOD1 sequence.
Using the PepMLM-650M model, I generated four potential binders of length 12 AA’s. The model’s confidence is reflected in the pseudo-perplexity scores, where lower values suggest a more plausible binding interaction.
| PepMLM Generated Binder |
|---|
WRYGVAGVRHWX |
WLYPPAVVEHKE |
HRYYPTAVRWKX |
WHYGVVGLAHKK |
Here’s also a screenshot of the binders generated using the PepMLM colab:
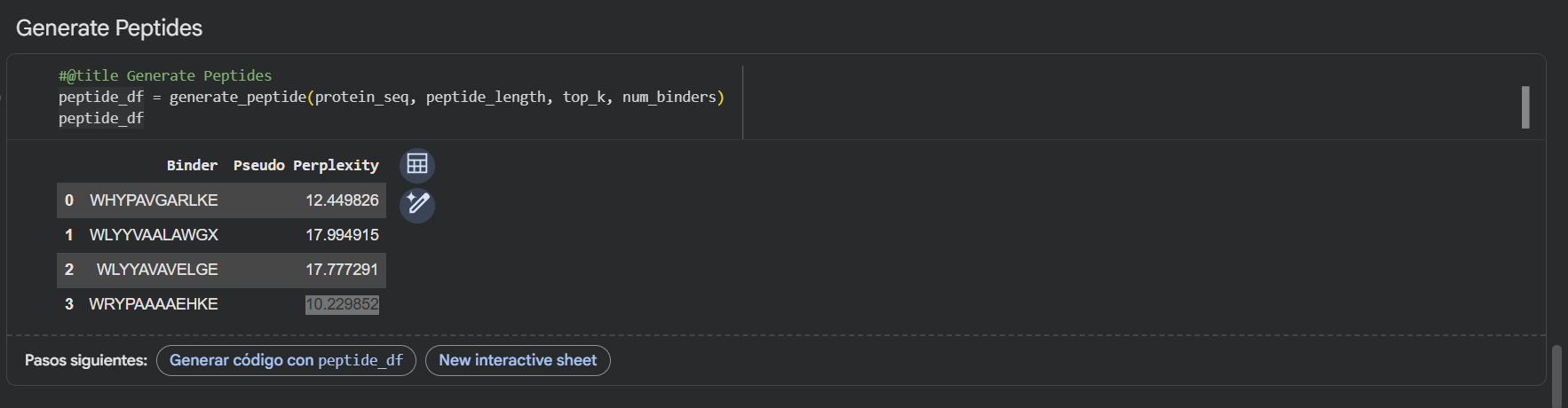

- To your generated list, add the known SOD1-binding peptide FLYRWLPSRRGG for comparison.
Here’s the updated list with the known SOD1-binding peptide:
| Binder |
|---|
WRYGVAGVRHWX |
WLYPPAVVEHKE |
HRYYPTAVRWKX |
WHYGVVGLAHKK |
FLYRWLPSRRGG |
- Record the perplexity scores that indicate PepMLM’s confidence in the binders.
Here’s the final list of binders with their respective perplexity scores
| Binder | Pseudo Perplexity |
|---|---|
WRYGVAGVRHWX | 13.614870 |
WLYPPAVVEHKE | 20.275341 |
HRYYPTAVRWKX | 10.113044 |
WHYGVVGLAHKK | 12.192786 |
FLYRWLPSRRGG | - |
My top-ranked candidate after looking at the generated binders is the third binder
WLYYAVAVELGE(perplexity score: 10.11) because of its low perplexity score. That indicates high model confidence, so it should generate the best results out of the four binders.
Part 2: Evaluate Binders with AlphaFold3
- Navigate to the AlphaFold Server: alphafoldserver.com
- For each peptide, submit the mutant SOD1 sequence followed by the peptide sequence as separate chains to model the protein-peptide complex.
Side note: since AlphaFold doesn’t support the X’s, I decided to use the neutral amino acid Alanine (A)
Peptide N°1: WRYGVAGVRHWX Seed: 1418500094
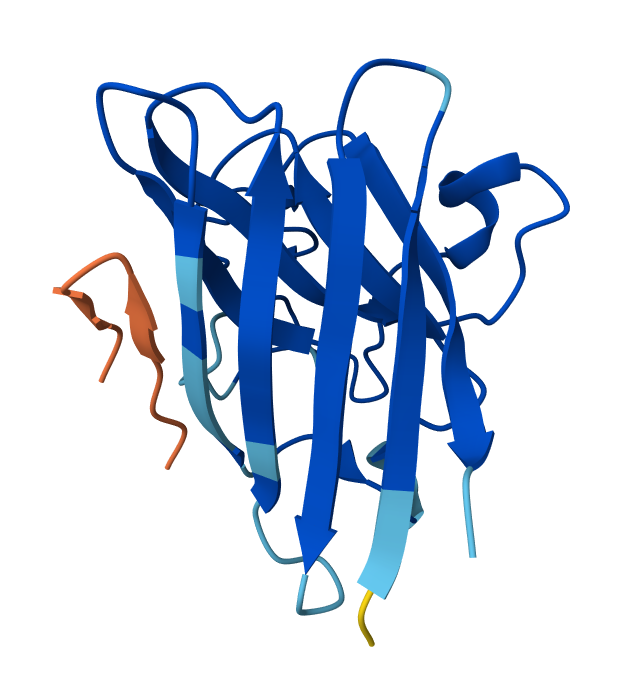

Peptide N°2: WLYPPAVVEHKE Seed: 1181188013
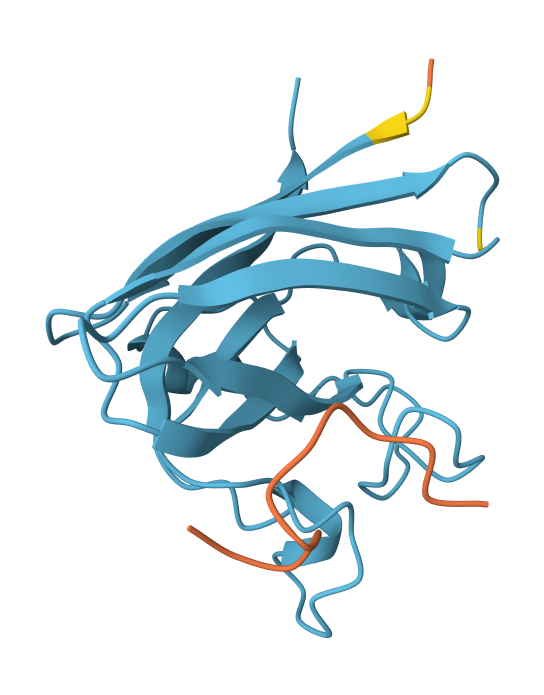

Peptide N°3: HRYYPTAVRWKX Seed: 826762887
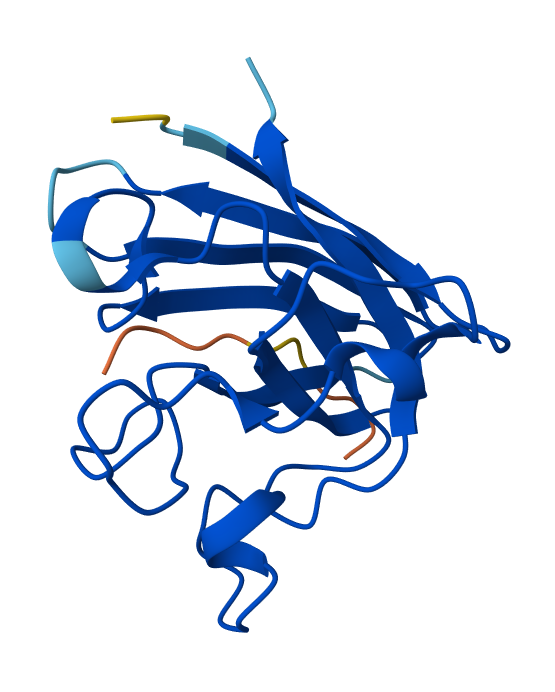

Peptide N°4: WHYGVVGLAHKK Seed: 1427381627
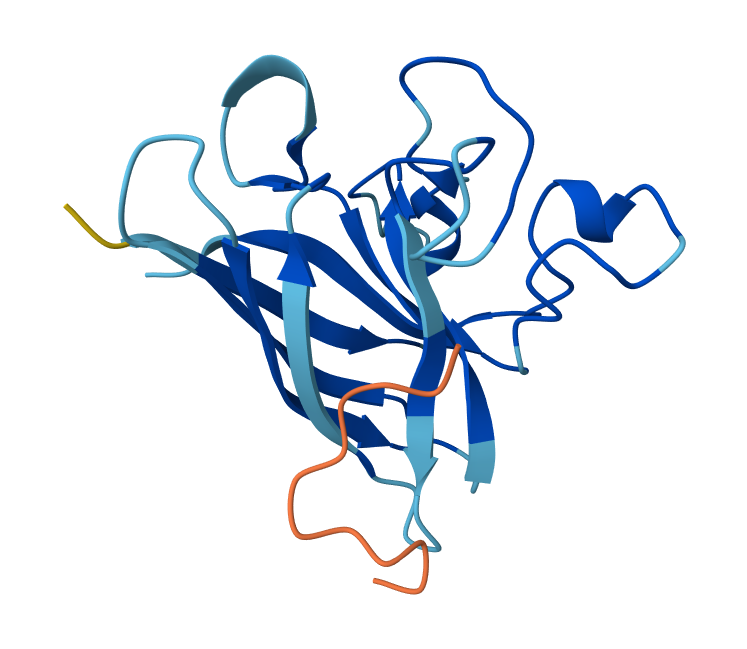

Peptide N°5: FLYRWLPSRRGG Seed: 449653589
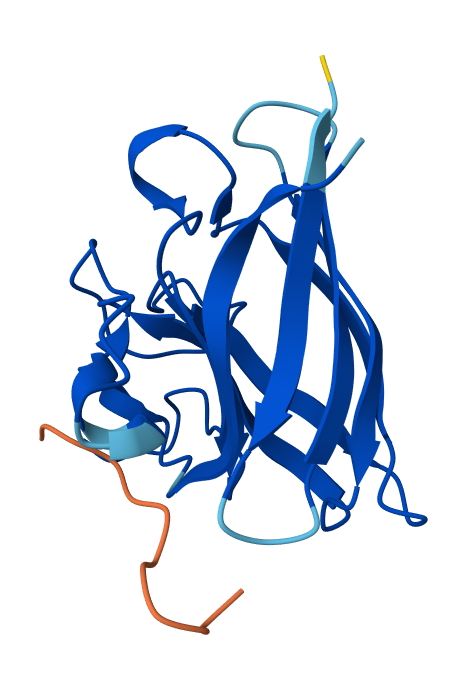

- Record the ipTM score and briefly describe where the peptide appears to bind. Does it localize near the N-terminus where A4V sits? Does it engage the β-barrel region or approach the dimer interface? Does it appear surface-bound or partially buried?
| Binder | Pseudo Perplexity | ipTM score |
|---|---|---|
WRYGVAGVRHWX | 13.614870 | 0.42 |
WLYPPAVVEHKE | 20.275341 | 0.25 |
HRYYPTAVRWKX | 10.113044 | 0.41 |
WHYGVVGLAHKK | 12.192786 | 0.34 |
FLYRWLPSRRGG | - | 0.31 |
- Peptide N°1: This one binds predominantly near the β-barrel region and partially engages the dimer interface. It appears partially buried within a surface groove, suggesting strong structural complementarity.
- Peptide N°2: This candidate localizes near the N-terminus, specifically approaching the A4V mutation site. However, it remains mostly surface-bound with lower structural confidence.
- Peptide N°3: Similar to the first binder, this peptide anchors itself against the β-barrel, showing a stable orientation that is partially buried against the protein core.
- Peptide N°4: It localizes at the edge of the dimer interface, appearing as a surface-bound “cap” rather than a buried ligand.
- Peptide N°5 (reference): The known binder shows a moderate ipTM, localizing primarily at the dimer interface of the SOD1 mutant.
- In a short paragraph, describe the ipTM values you observe and whether any PepMLM-generated peptide matches or exceeds the known binder.
The observed ipTM scores are moderate and lower than 0.5, but two of my peptides, mutant_peptide_1 and mutant_peptide_3, significantly exceeded the known reference binder (0.31). This confirms that PepMLM identified novel sequence patterns with better structural affinity for the mutant surface than the reference. While these scores aren’t yet at “drug-level” affinity, they provide a much better starting point for optimization than the current benchmark.
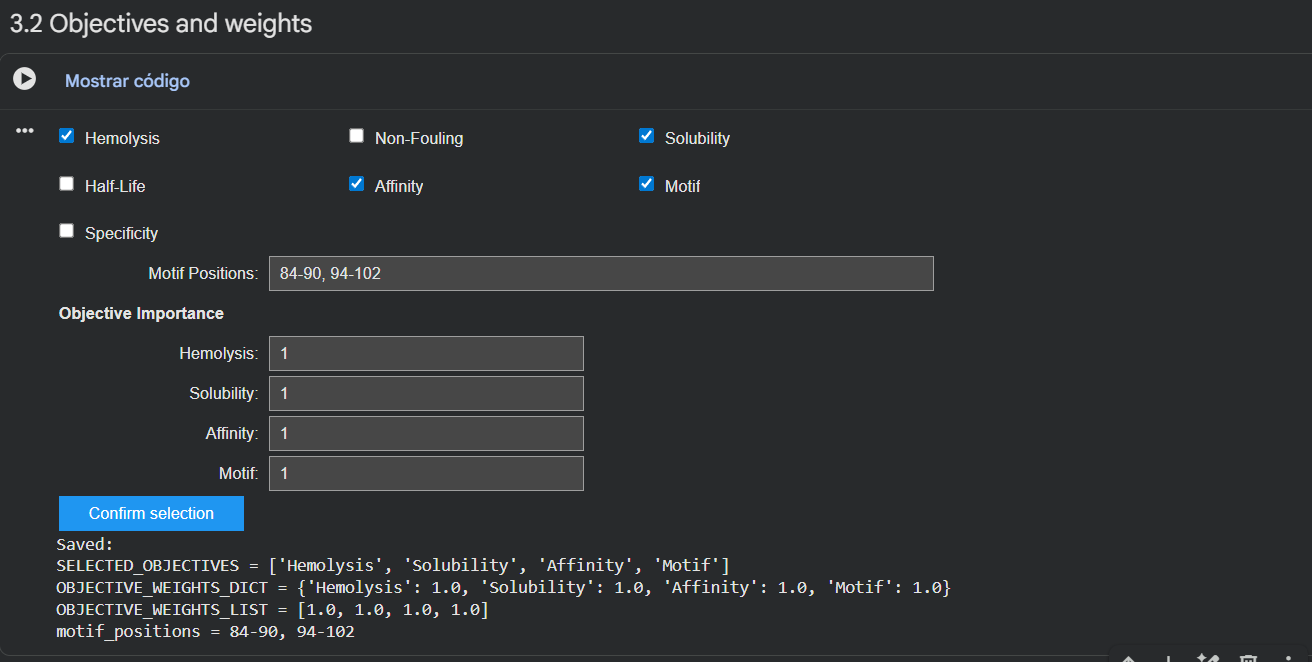
Part 3: Evaluate Properties of Generated Peptides in the PeptiVerse
Structural confidence alone is insufficient for therapeutic development. Using PeptiVerse, let’s evaluate the therapeutic properties of your peptide! For each PepMLM-generated peptide:
- Paste the peptide sequence.
- Paste the A4V mutant SOD1 sequence in the target field.
- Check the boxes
- Predicted binding affinity
- Solubility
- Hemolysis probability
- Net charge (pH 7)
- Molecular weight
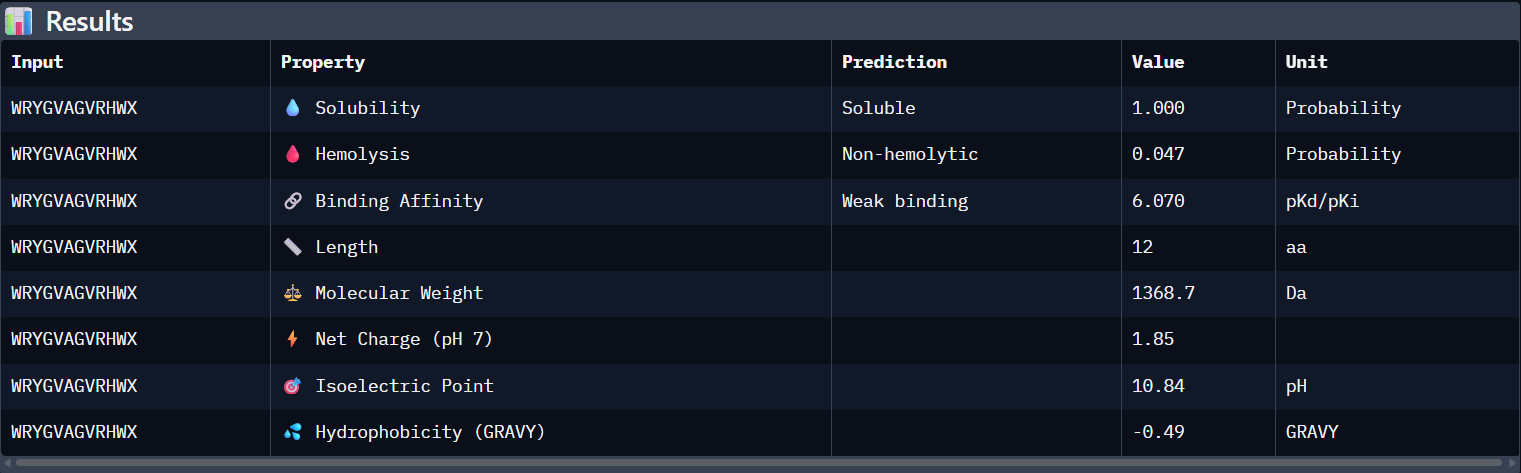
Peptide N°1:

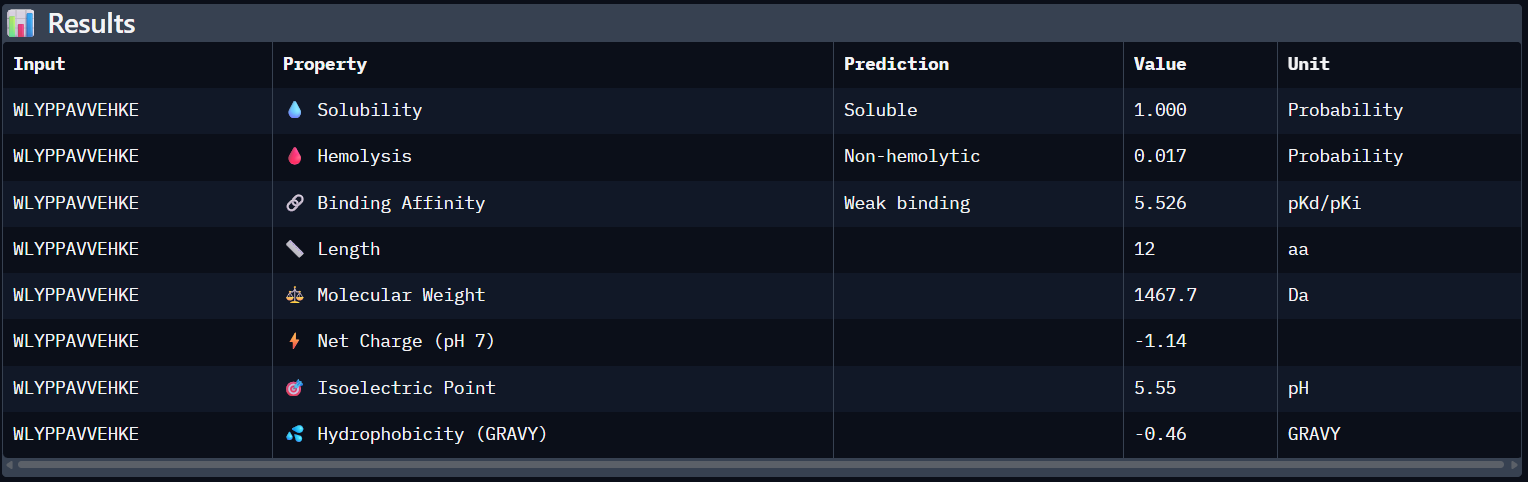
Peptide N°2:

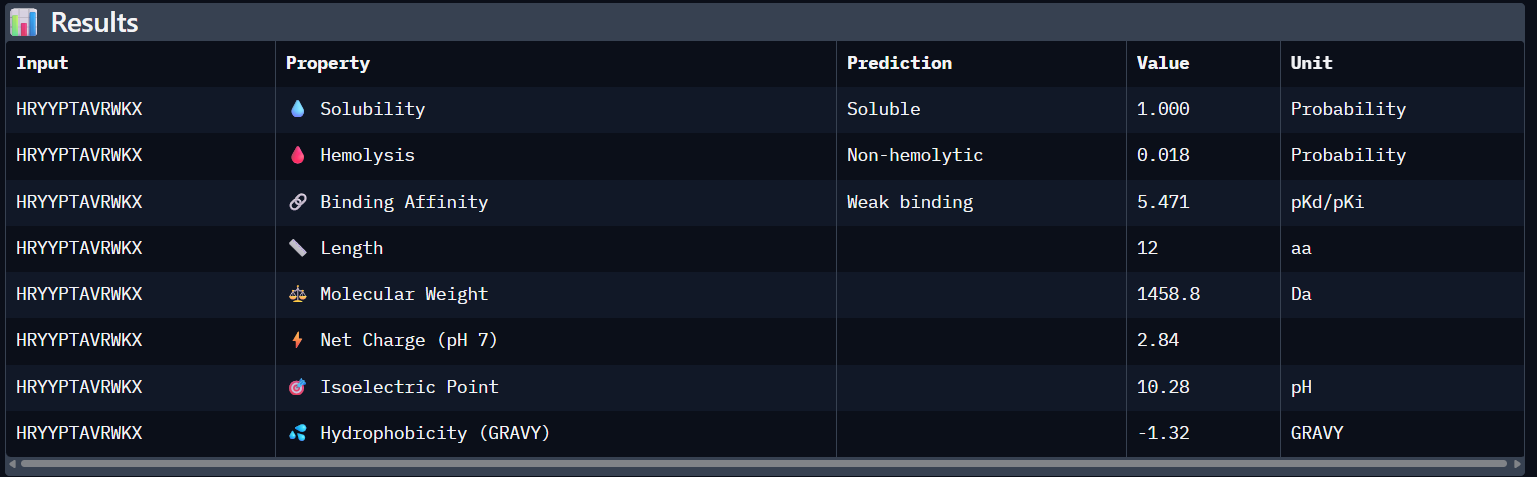
Peptide N°3:

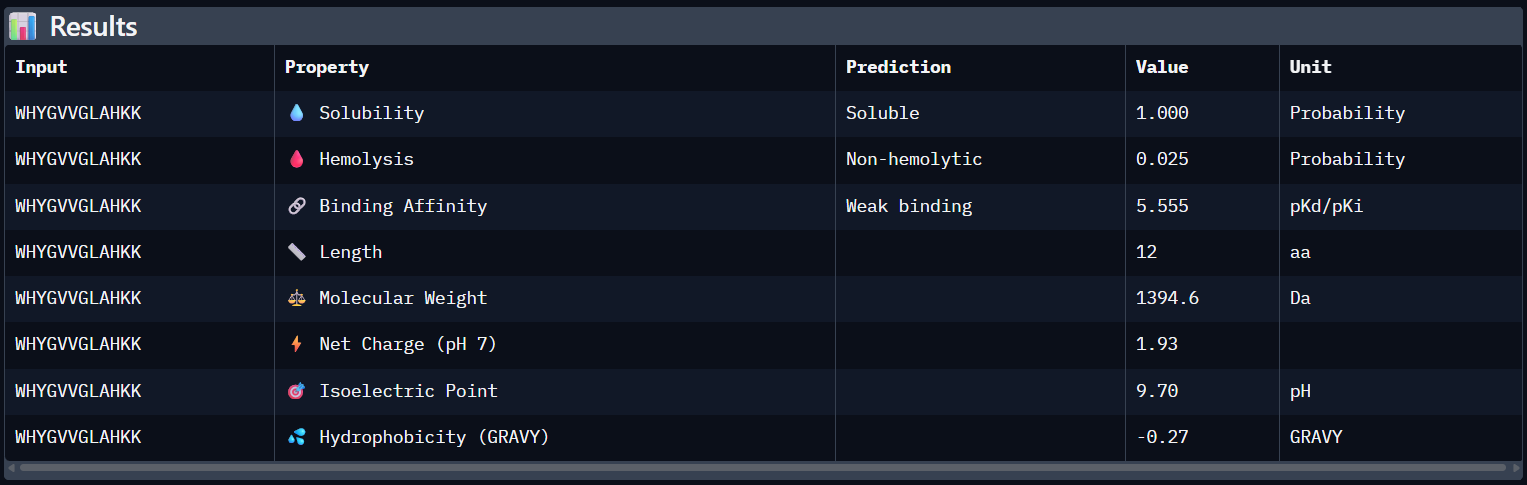
Peptide N°4:

Peptide N°5:
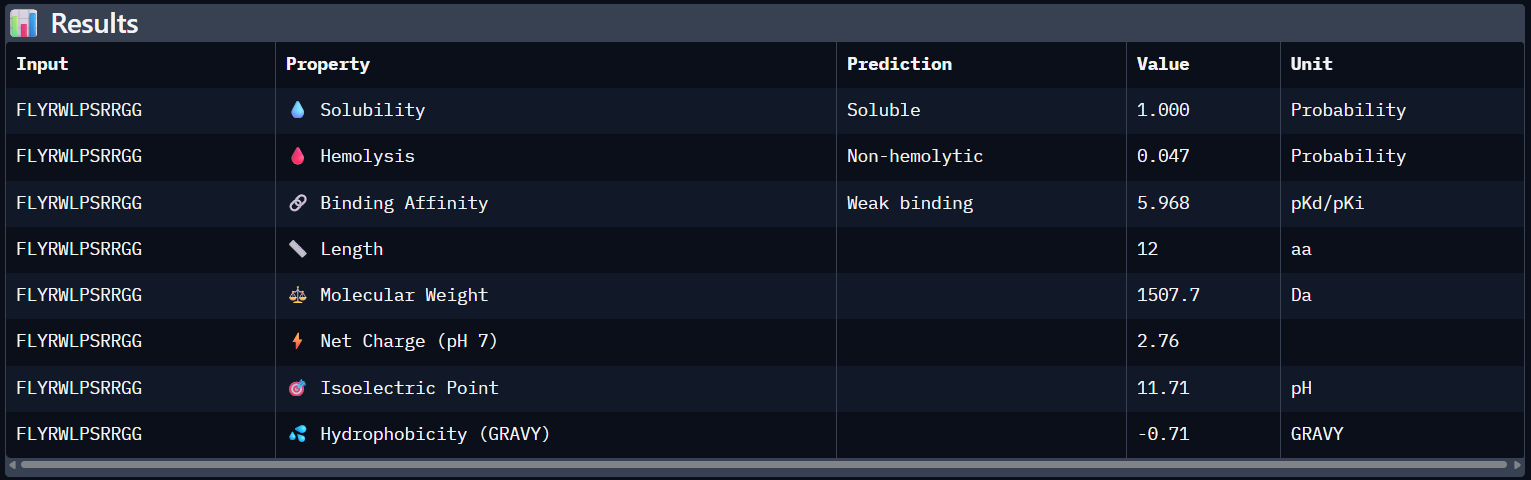

Analysis of the PeptiVerse Results:
| Peptide | ipTM (AF3) | Binding Affinity (pKd) | Solubility | Hemolysis (Prob) | Net Charge |
|---|---|---|---|---|---|
WRYGVAGVRHWA | 0.42 | 6.070 | Soluble (1.00) | 0.047 | +1.85 |
WLYPPAVVEHKE | 0.25 | 5.526 | Soluble (1.00) | 0.017 | -1.14 |
HRYYPTAVRWKA | 0.41 | 5.471 | Soluble (1.00) | 0.018 | +2.84 |
WHYGVVGLAHKK | 0.34 | 5.555 | Soluble (1.00) | 0.025 | +1.93 |
FLYRWLPSRRGG | 0.31 | 5.968 | Soluble (1.00) | 0.047 | +2.76 |
Compare these predictions to what you observed structurally with AlphaFold3. In a short paragraph, describe what you see. Do peptides with higher ipTM also show stronger predicted affinity? Are any strong binders predicted to be hemolytic or poorly soluble? Which peptide best balances predicted binding and therapeutic properties?
Comparing AlphaFold3’s structural confidence with PeptiVerse predictions shows that structural docking alone isn’t enough for drug design. My first and third PepMLM binders, WRYGVAGVRHWX and HRYYPTAVRWKX, both exceeded the known reference binder in ipTM score (0.42 and 0.41 vs. 0.31). Crucially, PeptiVerse shows these new designs are less hemolytic than the reference (0.018 vs. 0.047), which is a significant safety improvement.
Choose one peptide you would advance and justify your decision briefly.
I would choose to advance with my third peptide (HRYYPTAVRWKX) for therapeutic development because it strikes the best balance between structural fit and safety. It has the lowest chance of hemolysis (0.018) and is expected to be completely soluble (1.000). Its low pseudo-perplexity (10.11) also reflects high model confidence. Aditionally, the positive net charge (+2.84) should favor its interaction with the mutant SOD1 surface in the cytosolic environment.
Part 4: Generate Optimized Peptides with moPPIt
Now, move from sampling to controlled design. moPPIt uses Multi-Objective Guided Discrete Flow Matching (MOG-DFM) to steer peptide generation toward specific residues and optimize binding and therapeutic properties simultaneously. Unlike PepMLM, which samples plausible binders conditioned on just the target sequence, moPPIt lets you choose where you want to bind and optimize multiple objectives at once.
- Open the moPPit Colab linked from the HuggingFace moPPIt model card
- Make a copy and switch to a GPU runtime.
You can access my Colab moPPit copy here: Fabrizio_Flores_moPPIt-v3
In the notebook:
- Paste your A4V mutant SOD1 sequence.
- Choose specific residue indices on SOD1 that you want your peptide to bind (for example, residues near position 4, the dimer interface, or another surface patch).
I selected residues 84-90 and 94-102 from two adjacent β-strands (ending at Asp84 and Asp102) to define a broad binding pocket on the β-barrel surface.
- Set peptide length to 12 amino acids.
- Enable motif and affinity guidance (and solubility/hemolysis guidance if available). Generate peptides.
After generation, briefly describe how these moPPit peptides differ from your PepMLM peptides. How would you evaluate these peptides before advancing them to clinical studies?
These are the binders moPPit generated:
RVRTYKRTQKEMKCYSLKLKKKYYYEYYKKKTCQKH
Additionally, I wanted to see how AlphaFold3 evaluated the new optimized binders, so here are the results:
Optimized Peptide N°1:
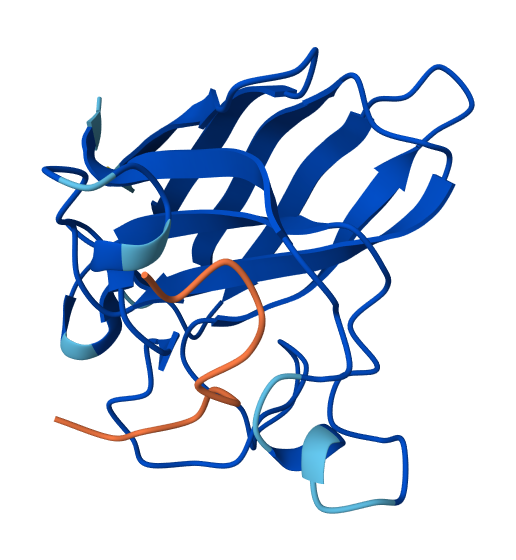

Optimized Peptide N°2:
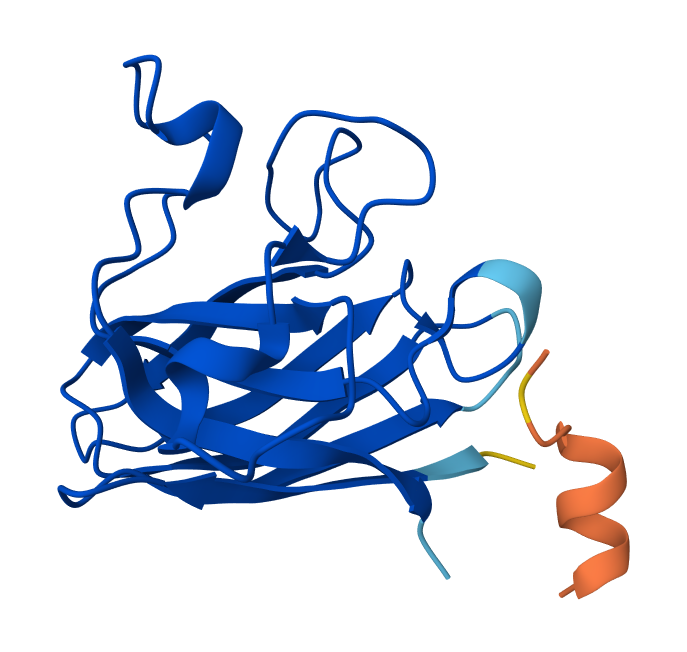

Optimized Peptide N°3:
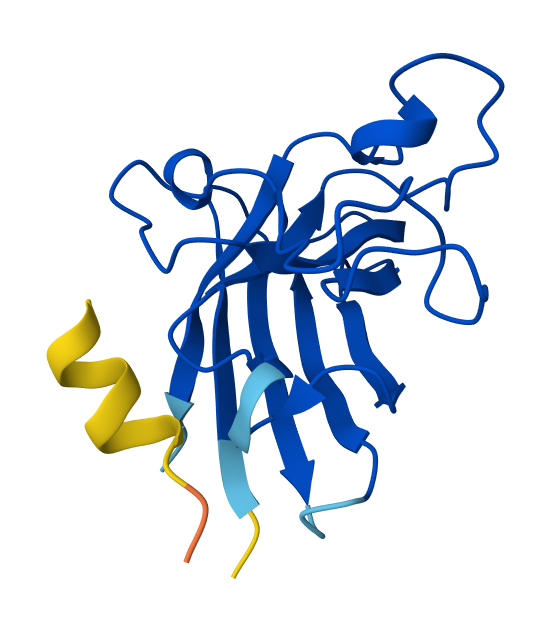

| Binder | AlphaFol3 Seed | ipTM score |
|---|---|---|
RVRTYKRTQKEM | 2124434605 | 0.34 |
KCYSLKLKKKYY | 276492257 | 0.32 |
YEYYKKKTCQKH | 1326061601 | 0.45 |
The third moPPIt-generated peptide is quite interesting since it achieved an ipTM score of 0.45 after evaluating it using AlphaFold3. This means that by “sculpting” a sequence specifically complemented to the Asp84-Asp102 motif, moPPIt created a high-affinity “molecular staple” that reinforces the SOD1 β-barrel core. This targeted motif approach balances affinity, solubility, and specificity and aims to prevent the structural collapse and toxic aggregation triggered by the A4V mutation, providing a precise lead candidate for clinical development.
Part B: BRD4 Drug Discovery Platform Tutorial (Gabriele)
(View Full Screen)
Skipping this one
Part C: Final Project: L-Protein Mutants
High level summary: The objective of this assignment is to improve the stability and auto-folding of the lysis protein of a MS2-phage. This mechanism is key to the understanding of how phages can potentially solve antibiotic-resistance.
This homework requires computation that might take you a while to run, so please get started early.
(View Full Screen)
L-Protein Engineering - Option 1: Mutagenesis
Step 1: Information Gathering
Here are the L-protein and Dnaj sequences
Lysis Protein Sequence (UniProtKB ID: P03609)
METRFPQQSQQTPASTNRRRPFKHEDYPCRRQQRSSTLYVLIFLAIFLSKFTNQLLLSLLEAVIRTVTTLQQLLT
DnaJ sequence (UniProtKB ID: P03609)
MAKQDYYEILGVSKTAEEREIRKAYKRLAMKYHPDRNQGDKEAEAKFKEIKEAYEVLTDSQKRAAYDQYGHAAFEQGGMGGGGFGGGADFSDIFGDVFGDIFGGGRGRQRAARGADLRYNMELTLEEAVRGVTKEIRIPTLEECDVCHGSGAKPGTQPQTCPTCHGSGQVQMRQGFFAVQQTCPHCQGRGTLIKDPCNKCHGHGRVERSKTLSVKIPAGVDTGDRIRLAGEGEAGEHGAPAGDLYVQVQVKQHPIFEREGNNLYCEVPINFAMAALGGEIEVPTLDGRVKLKVPGETQTGKLFRMRGKGVKSVRGGAQGDLLCRVVVETPVGLNERQKQLLQELQESFGGPTGEHNSPRSKSFFDGVKKFFDDLTR
Note: Lysis protein contains a soluble N-terminal domain followed by a transmembrane protein (blue/last 35 residues). Transmembrane protein affects the lysis activity. The soluble domain (green) is the domain responsible for interaction with DnaJ. Soluble N-terminal domain:
METRFPQQSQQTPASTNRRRPFKHEDYPCRRQQRSSTLYVTM domain:LIFLAIFLSKFTNQLLLSLLEAVIRTVTTLQQLLT
Additionally, here’s a screenshot of the BLAST results for L-protein:


Lastly, these results were aligned using Clustal Omega, revealing a highly conserved “island” (HEDYPCRRQQRSST) at residues 24-38. These sites will be avoided during mutagenesis to preserve the critical interaction with DnaJ and overall biological function of the phage.
Clustal Job ID: clustalo-I20260311-043120-0780-2033785-p2m
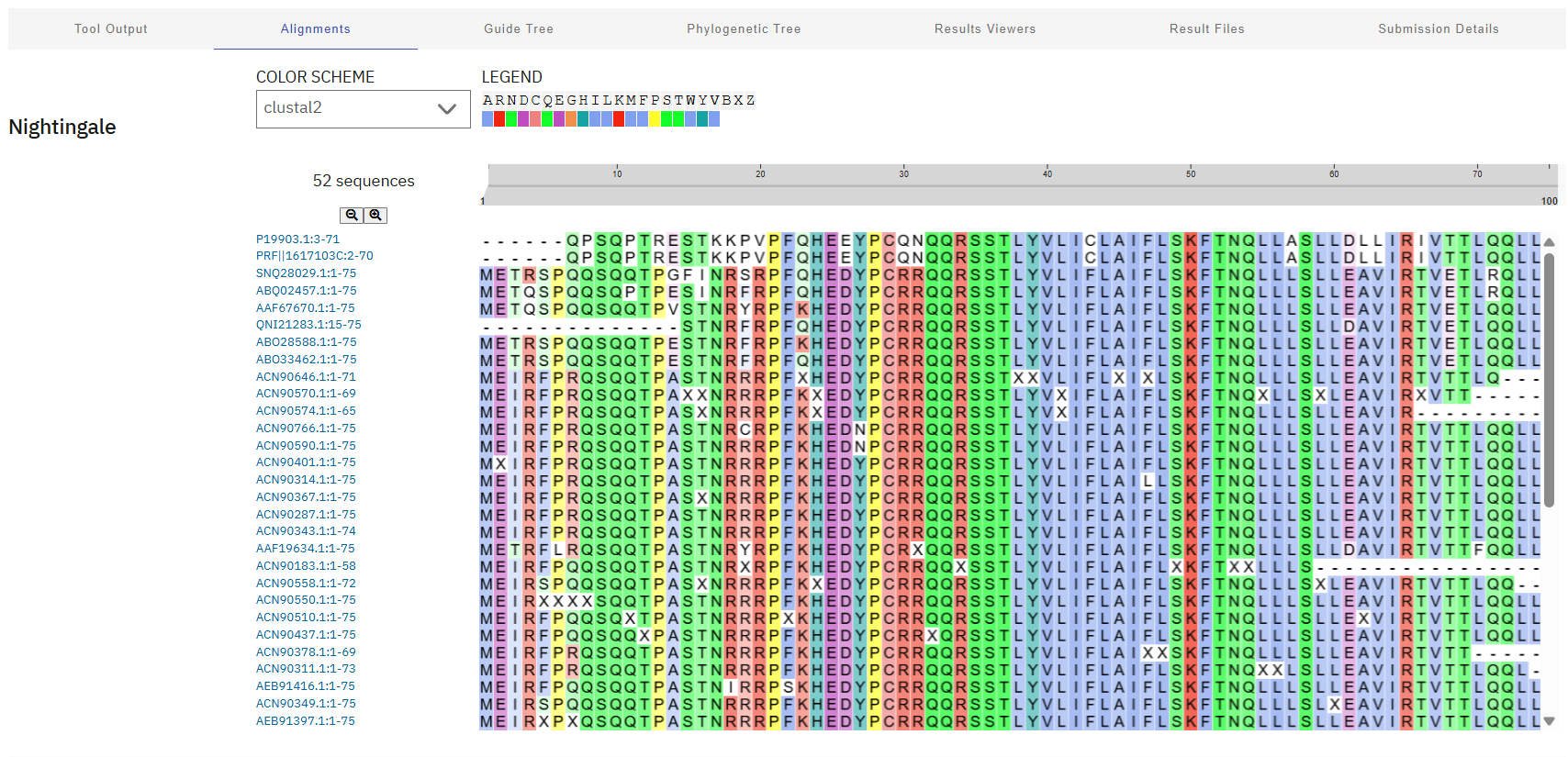

Step 2: Variant Selection Approach
My approach is very straightforward: I combined computational LLR scores with experimental lab data using a copy of the HTGAA Colab. I filtered for mutations that showed “active lysis” (value 1) in the experimental spreadsheet and high positive LLR scores in the notebook.
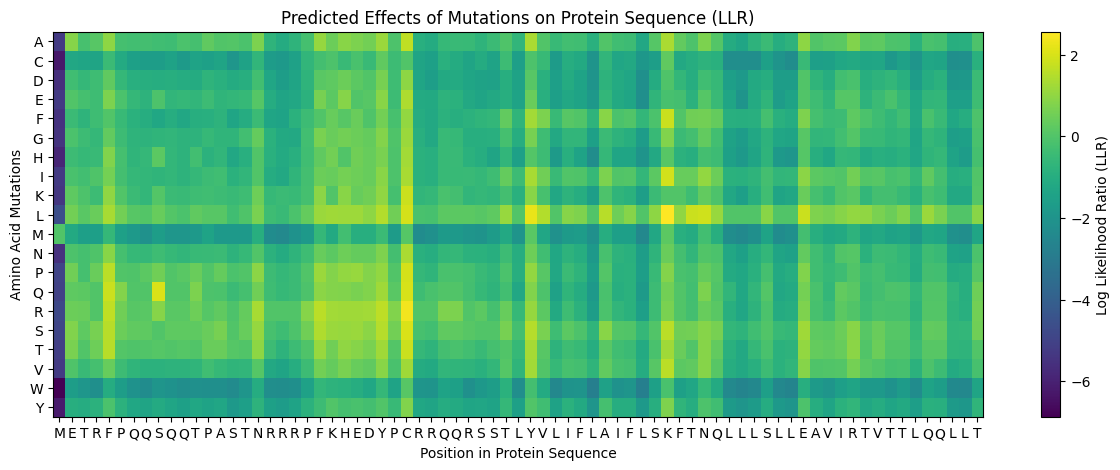

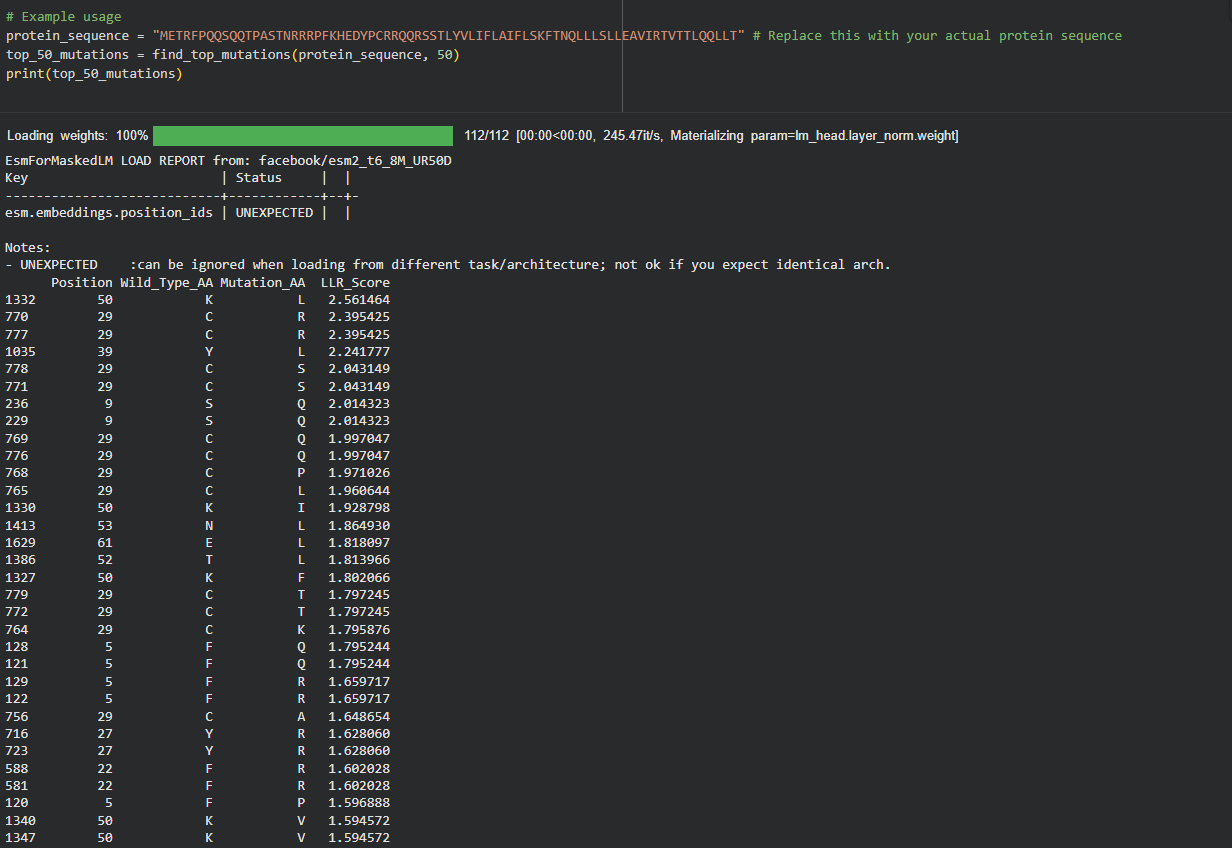

Step 3: Filtering and Ranking
I used AlphaFold 3 to model the 8-chain assembly. This step was used to rank candidates that showed both positive computational scores and confirmed experimental activity, ensuring they don’t disrupt the pore’s symmetry.
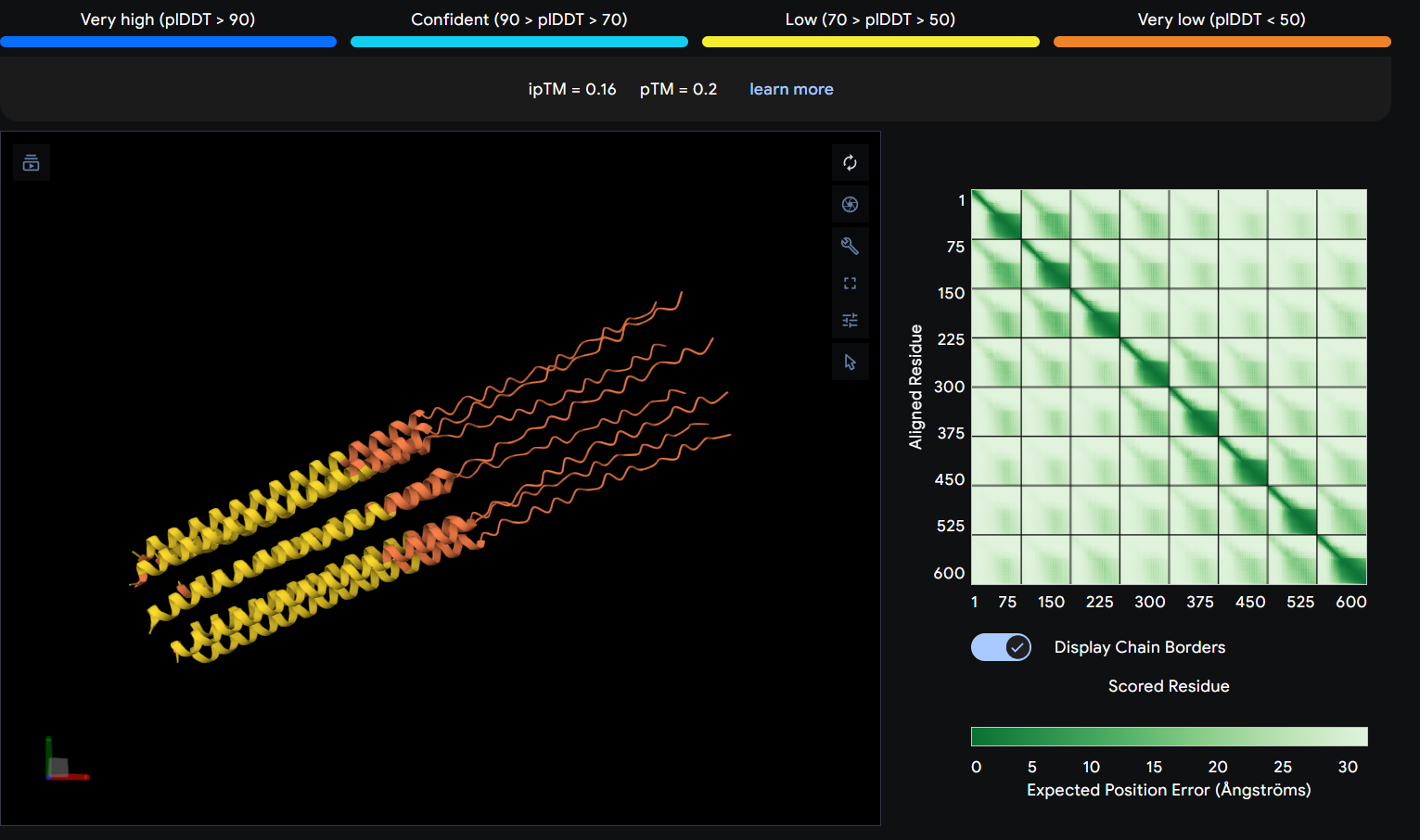

Step 4: Final Mutated Sequences
These 5 mutations were selected because they are experimentally proven to maintain lysis (score 1) and show improved or stable computational scores.
| Region | Mutation | LLR Score (ESM-2) | Experimental Lysis | Rationale |
|---|---|---|---|---|
| Soluble | S9Q | 2.01 | Active (1) | High computational confidence; replaces Serine with Glutamine to stabilize the N-terminal loop. |
| Soluble | C29R | 2.39 | Active (1) | One of the top scores; removing this Cysteine likely prevents incorrect disulfide bonding. |
| TM Domain | Y39L | 2.24 | Active (1) | High confidence score in the TM interface; optimizes hydrophobicity for membrane entry. |
| TM Domain | A45L | 1.53 | Active (1) | Consistent with experimental data; improves the hydrophobic core of the lytic pore. |
| TM Domain | N53L | 1.86 | Active (1) | Replaces a polar Asparagine with Leucine, significantly improving helix-helix packing in the multimer. |
S9Q mutation 8-chain assembly:

C29R mutation 8-chain assembly:

Y39L mutation 8-chain assembly:

A45L mutation 8-chain assembly:


N53L mutation 8-chain assembly:

While AF3 structures were used to visualize the multimeric orientation, the ipTM scores remained low (~0.17) across all mutations. This is expected given the small, intrinsically disordered nature of the L-protein and the high flexibility required for its lytic function, which challenges standard multimeric confidence metrics.