Individual Final Project
Here are my three individual final project ideas!
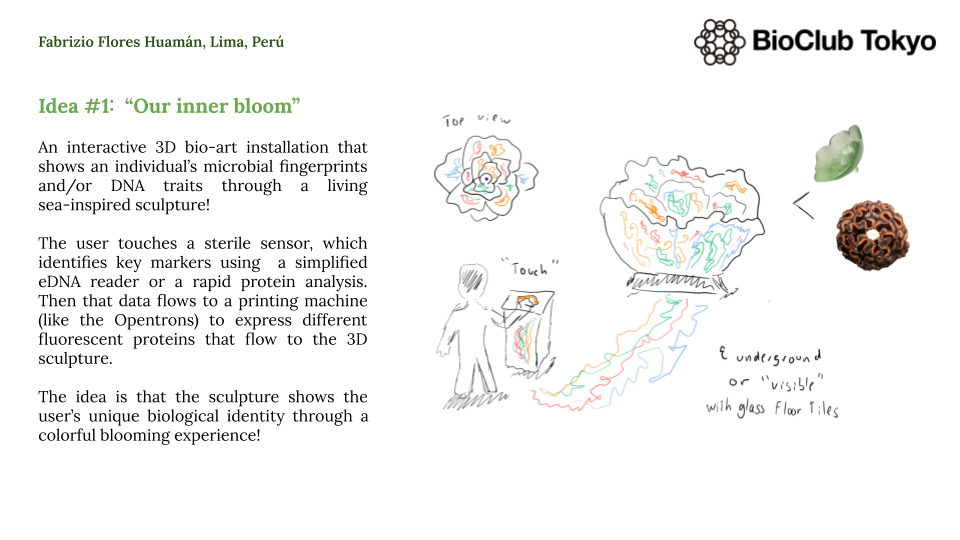

An interactive 3D bio-art sculpture where human touch meets living bioluminiscence bloom.
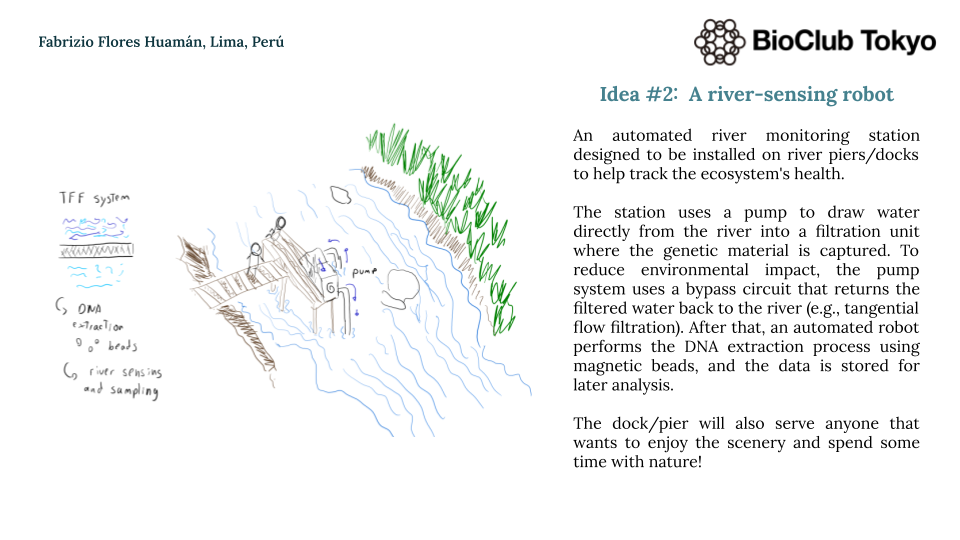


A river-sensing automated robot system that helps both advance research and act as a tourist spot for the community to reunite and spend time together.
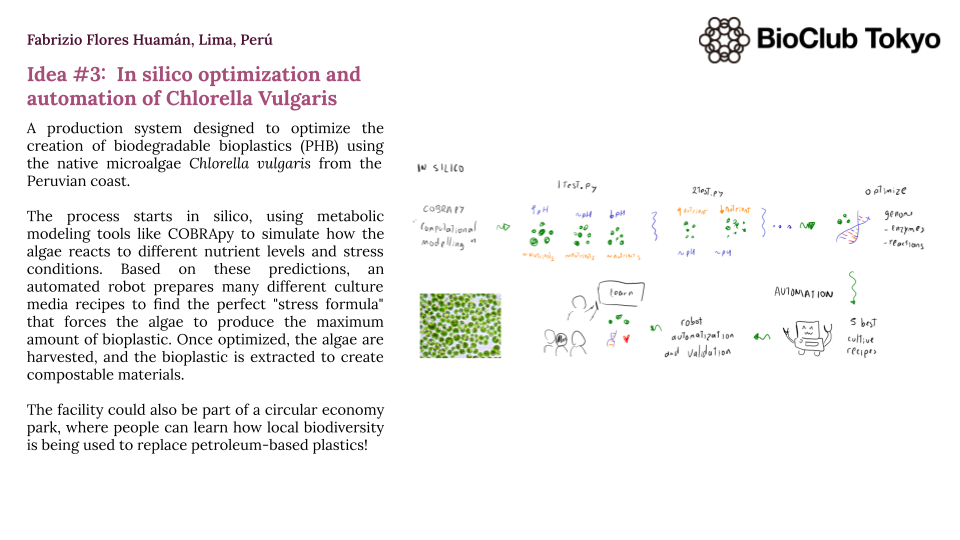


Chlorella vulgaris in silico optimization and automation for the optimal accumulation of polyhydroxybutyrate (PHB).
After some thought and consideration, I decided to go with my third idea for my final individual project, here you can find a more detailed view of it:


SECTION 1: ABSTRACT
Provide a concise, self-contained summary of your project (minimum 150 words). The abstract should allow a reader to understand the purpose, approach, and expected outcomes of the work without referring to other sections.
Your abstract should briefly address the following elements:
Significance: What problem or question does the project address, and why is it important?
The accumulation of petroleum-derived plastics has led to irreversible ecological damage, particularly in marine ecosystems like the ones in the coast of Peru. While Polyhydroxybutyrate (PHB) offers a 100% biodegradable and biocompatible alternative, its industrial scaling is hindered by low yields and the high cost of optimizing metabolic pathways. Microalgae like Chlorella vulgaris are ideal “chassis” because they fix CO2, but current optimization methods rely on slow, manual trial-and-error, creating a significant gap between laboratory research and industrial relevance.
Broad Objective: What is the overall goal of the project?
The overall goal of the project is to develop a high-throughput, closed-loop pipeline that integrates computational metabolic modeling with robotic automation. The goal is to rapidly identify and implement optimal “metabolic stress” environments that pivot Chlorella vulgaris from standard vegetative growth to specialized bioplastic accumulation, making the production of sustainable materials more predictable and effective.
Hypothesis: What prediction or principle is the project testing or demonstrating?
I hypothesize that Flux Balance Analysis (FBA) can accurately identify the metabolic “tipping points” where nutrient limitation (e.g., nitrogen or phosphorus) forces a shift in carbon flux. By using these models to predict the exact concentration of nutrients that inhibit biomass production without killing the cell, we can design “stress recipes” that maximize the partition of carbon into the PHB biosynthetic pathway (such as phaA, phaB, phaC).
Specific Aims: What key steps or milestones will be completed to achieve the objective?
Some of the steps I attempt to cover in the project are as follows:
Step 1: In Silico Modeling Reconstruct and simulate a genome-scale metabolic model of C. vulgaris using COBRApy to identify optimal flux distributions for PHB precursors (Acetyl-CoA). This involves mapping the metabolic shifts during the transition from biomass growth to polymer accumulation.
Step 2: Automated Protocol Design Develop and simulate Python-based automation protocols for the Opentrons OT-2 to execute parallelized micro-cultivations. The goal is to program the logic for complex “nutrient stress recipes” that can be deployed in a Biofoundry setting to validate model-driven growth and yield predictions.
Step 3: Visionary Scaling of the proccess Develop a theoretical framework for an AI-guided, modular photobioreactor system. This conceptual design focuses on the integration of real-time sensing data with metabolic models to enable decentralized bioplastic manufacturing in coastal communities, empowering them to utilize local biodiversity.
Methods: What experimental or technical approaches will be used?
The project follows a Design-Build-Test-Learn (DBTL) cycle, prioritizing computational engineering and standardized automation protocols:
Metabolic Modeling (Design): I will utilize COBRApy (Python) to perform Flux Balance Analysis (FBA) on a genome-scale metabolic model of Chlorella vulgaris. This involves defining a mathematical objective function for PHB production and applying environmental constraints (nitrogen, phosphorus, and carbon levels) to predict the metabolic states that maximize bioplastic precursors like acetyl-CoA.
Automation Protocol Development (Build): The predicted nutrient “recipes” will be translated into Python scripts for the Opentrons OT-2 robot. This ensures that the logic for combinatorial screening is high-precision and reproducible. By programming the liquid handling workflows, the project demonstrates how to scale the preparation of complex media that would be prone to human error if done manually.
Micro-scale Validation (Test - If resource availability permits): If laboratory access and resources allow, the programmed protocols will be executed in 24-well or 96-well plates. Growth dynamics would be monitored via optical density (OD) measurements at 680 nm and 750 nm to track chlorophyll health and cell density, providing the real-world data needed to refine the initial FBA models.
Genetic Construct Design (Future Build): Using Benchling, I will design a synthetic operon containing the phaA, phaB, and phaC genes. These sequences will be codon-optimized for Chlorella and placed under the control of nitrate-inducible promoters (e.g., NIT1). This genetic “switch” ensures that bioplastic synthesis only triggers when the system detects or creates a nitrogen-depleted environment, as predicted by the model.
System Architecture (Vision): For the visionary phase, the focus shifts to a theoretical system architecture for a modular photobioreactor. Instead of physical manufacturing, this involves defining the integration of low-cost sensors (pH, temperature, and light) with a closed-loop AI model to maintain the optimal metabolic conditions identified during the computational phase.
SECTION 2: PROJECT AIMS
Define three aims for your final project (minimum one sentence per aim).
Aim 1: Experimental Aim (this project):
“The first aim of my final project is to [achievable experimental goal] by utilizing [protocols, tools, or strategies].” This aim should describe the core experimental objective you will attempt during this class. List or link any relevant methods or resources you plan to use (e.g., experimental protocols, automation workflows, DNA or protein designs, protein design tools, or Twist orders). You will provide a detailed step-by-step experimental plan for Aim 1 in the Experimental Design section of this assignment.
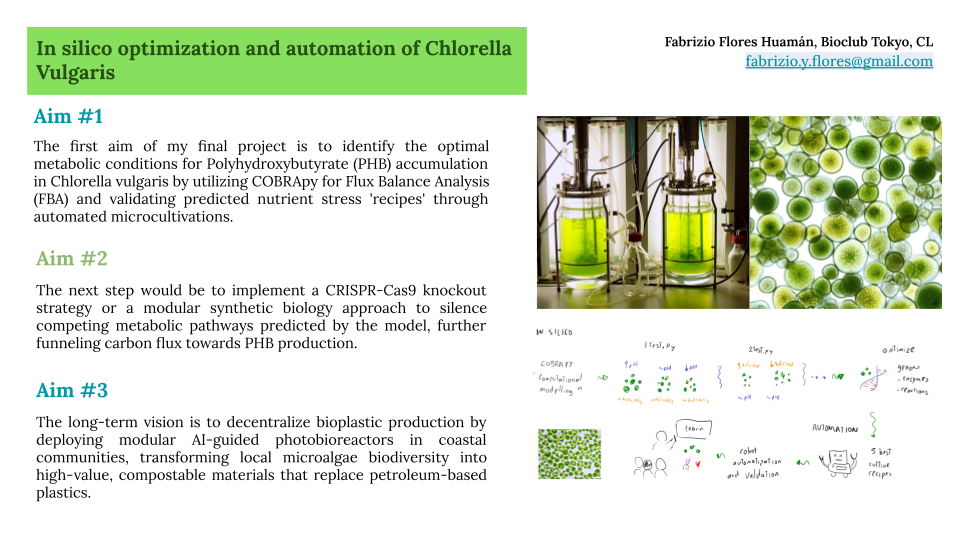
“The first aim of my final project is to identify the optimal metabolic conditions for PHB accumulation in Chlorella vulgaris by utilizing COBRApy for Flux Balance Analysis (FBA) and developing automated Python-based protocols for the Opentrons OT-2 to simulate high-throughput nutrient stress ‘recipes.’”
Aim 2: Development Aim:
Describe the next step that would follow a successful Aim 1, extending the work beyond the scope of this course. This aim should represent a realistic progression of the project, such as executing additional experiments, solving a technical limitation, or developing the system or technology further.
“The next step following the computational validation would be to experimentally implement the designed genetic constructs using a modular synthetic biology approach, overexpressing the PHB biosynthetic pathway under the control of the nitrate-inducible NIT1 promoter to evaluate real-time polymer accumulation.”
Aim 3: Visionary Aim:
Describe the long-term vision for the project. Explain how the broader concept could have an impact if fully realized. Examples include: Challenging an existing paradigm or clinical practice. Addressing a major barrier in a field. Enabling a new experimental capability or research approach.
“The long-term vision is to establish a decentralized framework for bioplastic production through AI-guided, closed-loop modular photobioreactors, enabling coastal communities to transform local microalgae biodiversity into sustainable, high-value compostable materials that replace petroleum-based plastics.”
SECTION 3: BACKGROUND
Background and Literature Context Provide background research that explains the current state of knowledge and identifies the gap in knowledge or capability that your project addresses.
Briefly summarize two peer-reviewed research citations relevant to your research (minimum four sentences).
Explain how your project is novel or innovative. (Minimum 3 sentences.) Examples of topics to discuss: New applications or uses of existing biological tools or concepts. Development of new approaches, methodologies, or technologies. Ways the project challenges existing paradigms or assumptions. How the work expands the boundaries of synthetic biology.
Explain why your project matters and what impact it could have. (Minimum 5 sentences.) Examples of topics to discuss: The problem addressed: What pressing real-world problem does your project attempt to solve? Importance of the problem: Why is this problem significant, or what critical barrier to progress in the field does it represent? Broader societal contribution: How could the outcomes of your project benefit society beyond the immediate research context? Advancement of knowledge or capability: How might the project improve scientific understanding, technical capability, or clinical practice within one or more fields? Field-level change: If your aims are achieved, how could the concepts, methods, technologies, treatments, services, or preventative approaches used in this field of research change?
Describe the ethical implications associated with your project and identify relevant ethical principles (e.g., non-maleficence, beneficence, justice, or responsibility). (Minimum 2 paragraphs.) First paragraph: Include what ethical implications are involved in your project. Try to suggest ethical the principle(s) you may apply (e.g. non-maleficence, justice)? Second paragraph: Describe the measures that should be taken to ensure that your project is ethical (both in how the research is conducted and in its broader implications for society). You may wish to answer the following questions: What action(s) do you propose? What are potential unintended consequences of your proposed actions? What could you have been wrong (e.g., incorrect assumptions and uncertainties)? What are alternatives to your proposed actions? Note: in an NIH proposal, an ethics statement is used to describe the relevance of this research to public health
SECTION 4: EXPERIMENTAL DESIGN, TECHNIQUES, TOOLS, AND TECHNOLOGY
Use Claude AI skills to refine your HTGAA final project experimental design here
Create a detailed experimental plan for your final project. Include a timeline for each part of your experimental plan (i.e., how long you would expect each step in your final project to take). (min. 15 lines/sentences—a numbered list is acceptable) Include specific methods/tools/technologies/biological concepts for each part of the final project and analysis This section will be used to determine whether the experiments are well designed, feasible, and likely to succeed in testing your hypothesis Often this section is broken into discrete tasks/sub-aims For each experiment and/or analysis, include a description of your expected results If possible, include figure(s) that visually shows a broad workflow of your project or a specific aspect of your experimental plan Reminder: All HTGAA projects must include some DNA design! Make sure this form is submitted.
We discussed and practiced various techniques related to synthetic biology throughout the semester. Place a check next to the techniques relevant to your project.
Pipetting Pipetting Lab Safety Bioethical Considerations (must check this box)
DNA Gel Art DNA Sequencing DNA Editing DNA Construct Design Restriction Enzyme Digestion Gel Electrophoresis DNA Purification From Gel Databases (e.g., GenBank, NCBI, Ensembl, and UCSC Genome Browser)
Lab Automation Creating Code for Laboratory Automation Using Liquid Handling Robots (e.g., Opentrons) Designing a Twist Order Creating a plan to use the Autonomous lab at Ginkgo Bioworks
Protein Design Protein Design Use of Boltz or PepMLM Use of Asimov Kernel Use of Benchling Models and Notebooks Databases Bioproduction Bioproduction Chassis Selection (e.g., DH5alpha) Registry of Standard Biological Parts Plasmid Preparation Bacterial Culturing Quality Control/Analysis Bacterial Processing (e.g., Centrifugation, Lysis, DNA Purification)
Cell-Free Systems Cell Free Reactions Freeze-Dried Cell Free Systems miniPCR Tools Protein Purification
Gibson Assembly Primer Design or Selection PCR Reactions Gibson Assembly Other Cloning Methods (e.g., Restriction Enzyme Digestion or Gateway Cloning)
CRISPR CRISPR/Cas9 Designing Prime Editing gRNA
Expand upon two techniques you checked in the previous question by describing how you would utilize those techniques in your final project. (min. 4 sentences)
Identify any How To Grow (Almost) Anything Industry Council companies which are associated with your final project (optional)
Addgene Asimov (Kernel) ATCC Basecamp Research BioFabricate Biome Consortia Bolt Boltz.bio Cultivarium DeepCure Epibone Ginkgo Bioworks Helix Nano Millipore Sigma Mycoworks New England Biolabs Nuclera Opentrons SecureDNA Takeda Pharmaceuticals Thermo Fisher Scientific Transfyr.ai Twist Biosciences Upside Foods Waters Corporation
SECTION 5: Results & Quantitative Expectations
You are required to validate at least one aspect of your final project aims. This is to ensure that you are able to successfully apply a relevant synthetic biology technique to your project. Include figures if you have them—accuracy is critical in figures, tables, and graphs Here is a non-exhaustive list of acceptable validations: Designing DNA relevant to your final project Performing a PCR reaction using primers relevant to your final project Performing a Gibson assembly relevant to your final project Creating and performing a cell-free assay related to your final project Creating and running code to validate an aspect of your final project Developing a model or completing a computational analysis relevant to your project Designing DNA construct(s) that can express at least one gene of interest, ordering it (via Twist), and testing of the expression of the construct(s) (potentially using an Opentrons robot)
What aspect of your final project did you choose to validate? (min. 2 sentences)
Write down a detailed protocol of how you validated this aspect of your final project. (Numbered list or paragraph is fine)
What synthetic biology techniques did you utilize in validating this aspect of your final project? You can refer to the list of techniques in question 8. (min. 4 sentences)
You must present data as part of your final project and include some analysis of that data. The data may be collected experimentally in the lab or generated as simulated data (e.g., using the Asimov Kernel or another simulation method). (min. 2 sentences)
Did you encounter any unexpected challenge(s) when performing your validation? If so, describe the challenge(s) and strategies to overcome it. If not, discuss potential problems, difficulties, limitations, and/or alternative strategies to overcome challenges in your final project. (min. 4 sentences).
SECTION 6: ADDITIONAL INFORMATION
List all references cited in this assignment (bullet-point list)
Create a supply list and budget for your project (bullet-point list)
- What supplies, equipment, and budget is needed for your project to work?