Week 2 HW: DNA Read, Write, & Edit


🎨 Benchling & In-silico Gel Art
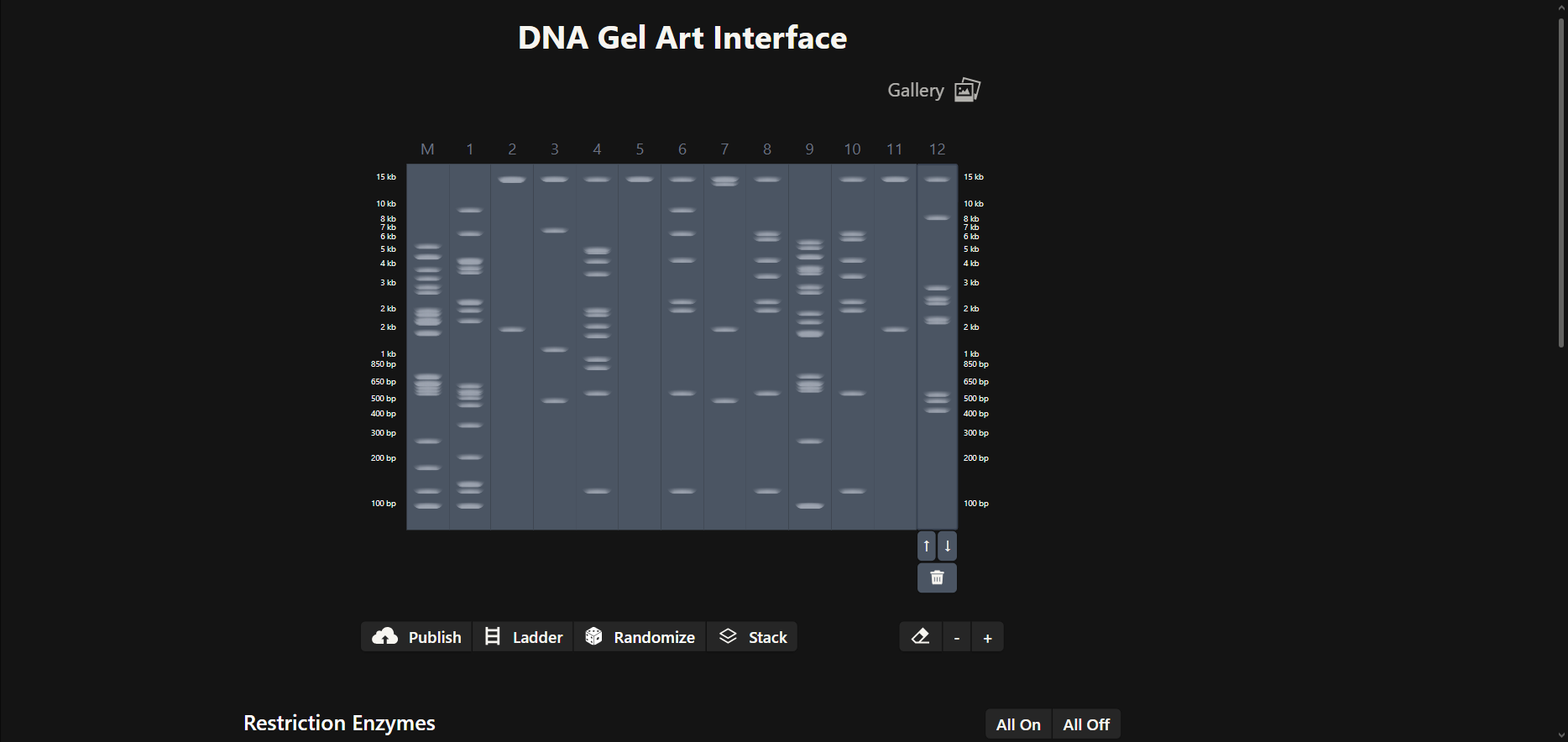
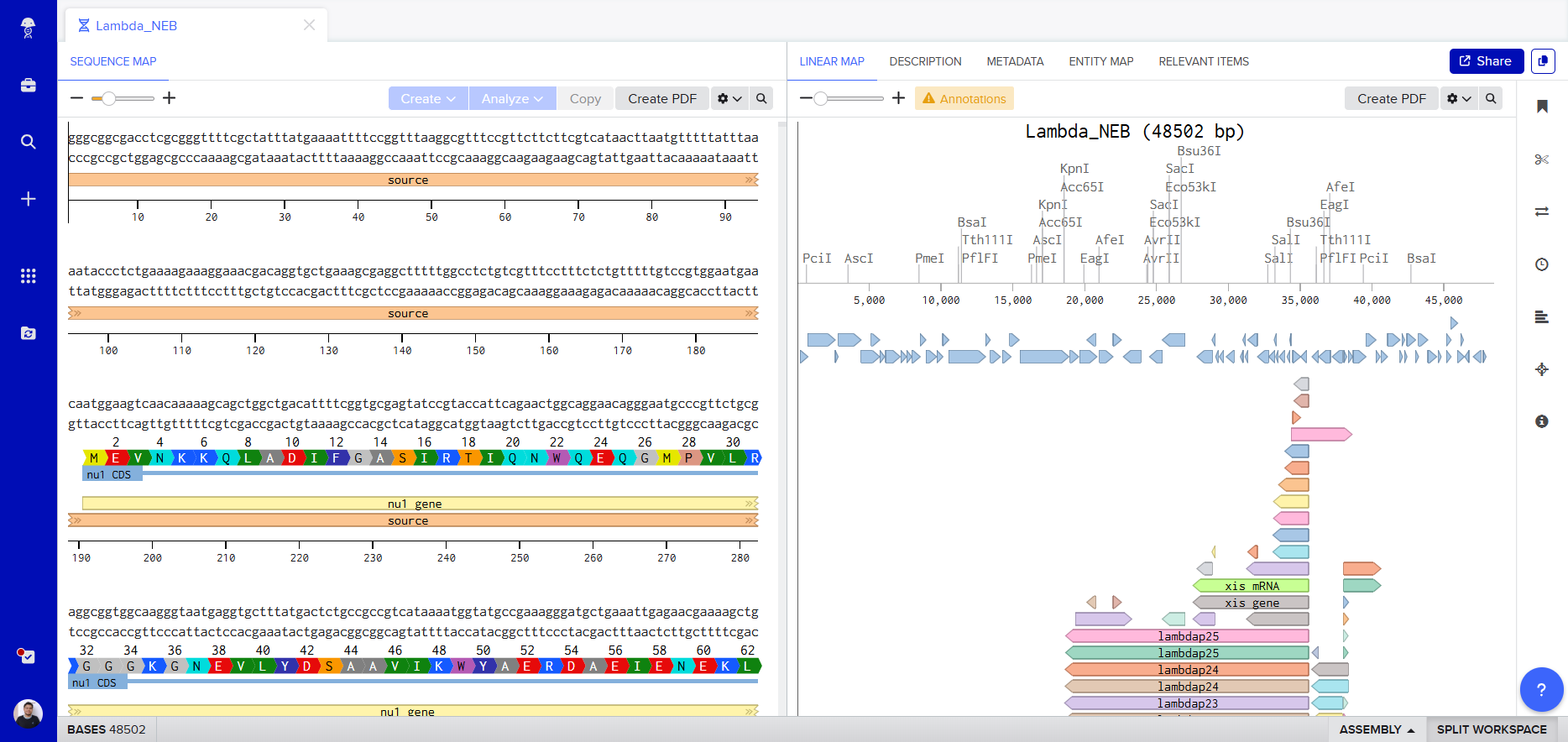
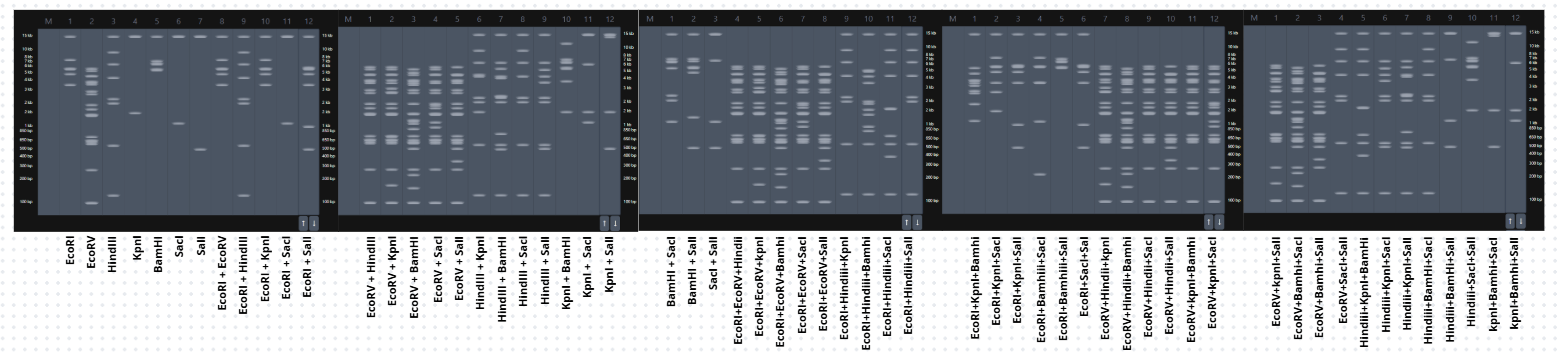
To begin my journey into understanding DNA, I made a “Gel Art” using restriction enzymes to create a beautiful image by cutting the Lambda sequence.
- Gel Art
- A technique that uses gel electrophoresis not for analytical purposes, but for artistic ones. Pigments or colored biomolecules are separated in an electric field to create abstract, controlled visual patterns and designs 1.
- Restriction enzymes
- They are “molecular scissors” (proteins) that cut DNA at specific sequences. They are fundamental tools in genetic engineering for isolating genes, recombining DNA, and analyzing samples 2.
- Well, to do that, I created a Benchling account, joined the HTGAA group, created a folder with my name, and imported my sequences there.
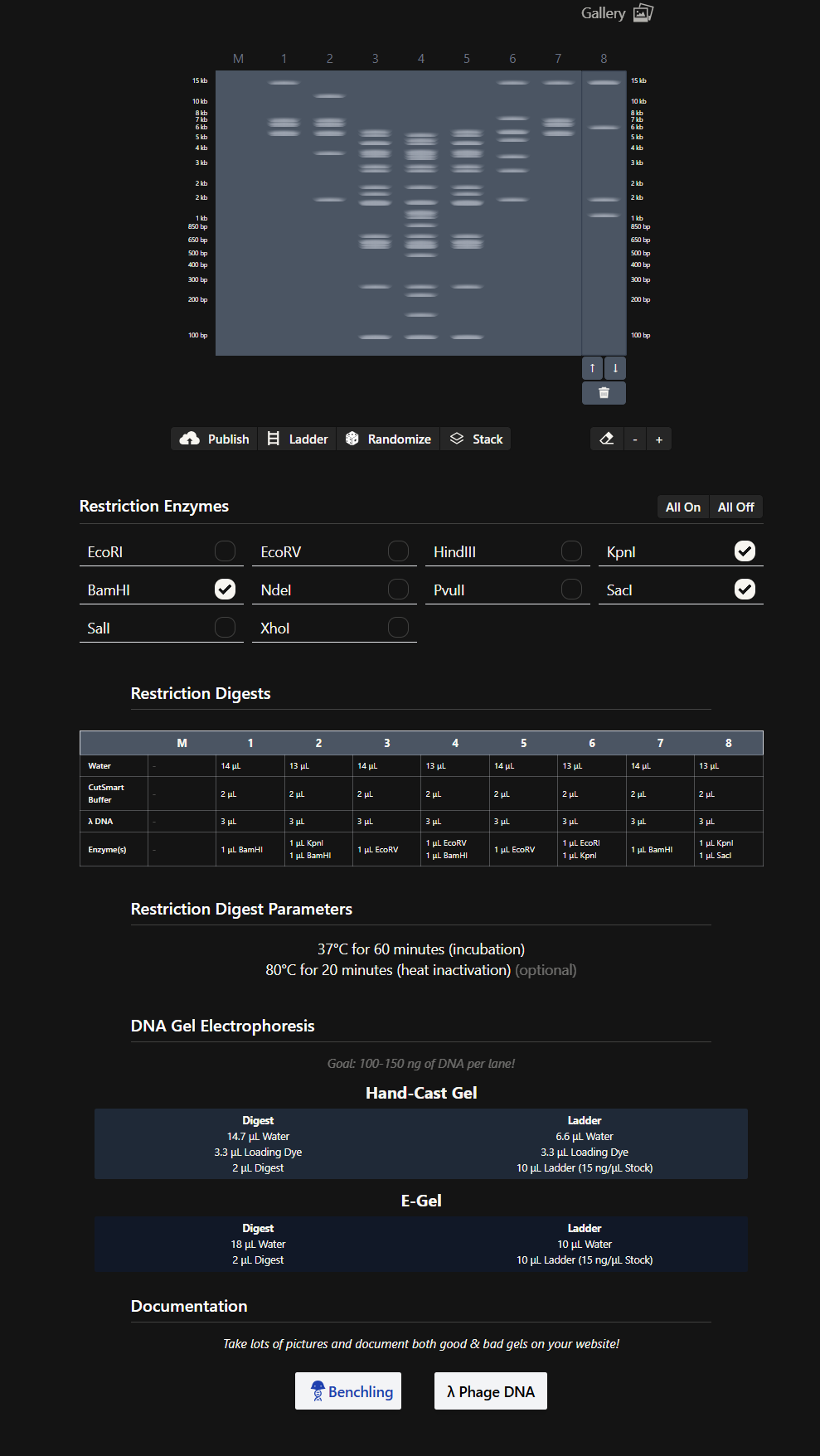
- Then, on my own, I started making combinations of the digests in the DNA Gel Art Interface on Ronan’s website.
In this part, I spent some time iterating to get an idea of what figure to create.
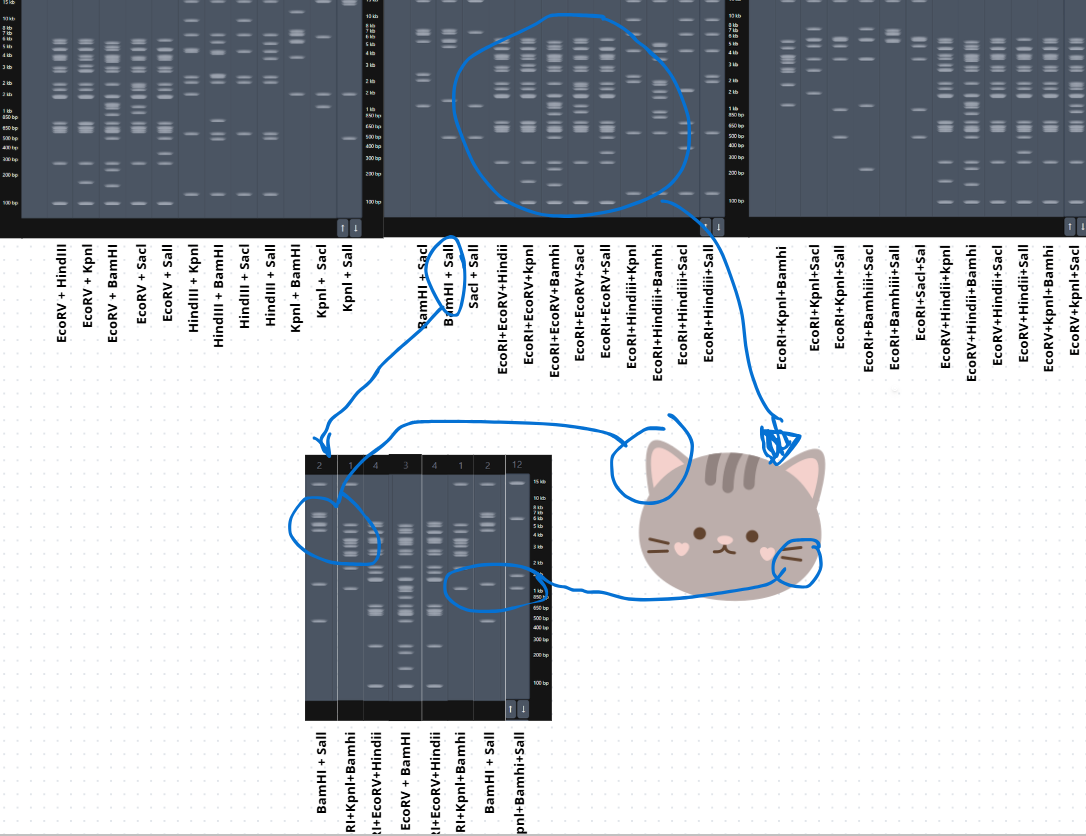
- After trying several combinations, I came up with this design for a cat’s face. 😸
- Then I performed the digestion in Benchling, and this was the result.
🧬 DNA Design Challenge
For this exercise, I chose the FIP-gts protein, an immunomodulatory protein identified in the Ganoderma tsugae fungus. I selected this protein after several searches on Google Scholar, Perplexity AI, and the Crow tool on the Edison platform.3 This protein caught my attention because it has therapeutic potential as an immunomodulatory, anticancer, and antiallergic agent. I feel that its antiallergic properties could be very useful, as many people I know suffer from problems caused by environmental and food allergies.
🟩 With the help of NCBI4, I searched for the sequence of the FIP-gts protein, but it did not appear as such. Reviewing other articles, I found the accession to the homologous protein, Ling zhi-85 (Lz-8, accession number AAA33350). Its amino acid sequence in FASTA format is:
>AAA33350.1 immunomodulatory protein [Ganoderma lucidum]
MSDTALIFRLAWDVKKLSFDYTPNWGRGNPNNFIDTVTFPKVLTDKAYTYRVAVSGRNLGVKPSYAVESDGSQKVNFLEYNSGYGIADTNTIQVFVVDPDTNNDFIIAQWN
Following the central dogma of molecular biology:
--- title: Central dogma of Molecular Biology --- graph LR; A[DNA] -->|Replication| A A -->|Transcription| B[RNA] B -->|Translation| C[Protein]
The dogma does not state that reverse translation can be done conventionally, even with the help of online bioinformatics tools, I performed reverse translation with the help of ProteinIQ and obtained the following sequence:
>AAA33350.1 immunomodulatory protein [Ganoderma lucidum]
ATGAGCGATACCGCCTTGATTTTCCGCTTGGCTTGGGATGTGAAGAAACTATCATTTGATTATACGCCAAACTGGGGGAGGGGTAACCCTAATAATTTTATCGACACTGTGACATTTCCAAAGGTACTCACTGATAAAGCTTACACTTATCGGGTCGCCGTTTCTGGACGTAACCTCGGCGTCAAGCCTAGCTACGCCGTGGAATCCGATGGTTCTCAAAAGGTGAACTTTCTTGAATATAACAGCGGATATGGCATCGCAGACACGAATACCATACAAGTGTTTGTAGTCGACCCGGATACCAACAATGATTTCATCATCGCACAATGGAAT
Out of curiosity, I compared it with its original genetic sequence:
>M58032.1 Ganoderma lucidum immunomodulatory protein (LZ-8) gene, complete cds
AAGTGGCATTATATCACCTTCGGATATCGCCATCCATAACCAACGTCTCAACCCCGAGATGGACGTCATCTGGAGGTAACGACCAAGGCGGTCTTCCGGCAACTGTGGTCTCGAAGACGTGAGGCGTTTACAAGGTTGGACATCTCGGGGCAATTCTGCCAAACTCGCAAGGAGAATGTACCGTCCTATCACCTGCAGTGGTCAGCAGGGGTTGCATCCAGGTCCACCTGCGGTGCAATTCGGTTCCCTGTGGCTTGCAGGGCCTCGCACGTCGTATGCACCCTGGTTTACATCATCGTGAAACGGCGCTCCGGTTGCCGTATAAAAGGACGGCAAAGGCGGCCAGTGGACTTCAGCACCTGCTCTTGTACCCACTTCTAGTAAGTCGCATACCACCTCTCCTGACAGCGACAGGTTCACTGACTAGTTCATAGTCCACAGCTCTTTGCCTTACAATCAAGGTTTGCCGACACCCTCTTTGAGCCCTCCCCCTTCAATAACCCCCTAACTTCTGCCCCGCAGCATCATGTCCGACACTGCCTTGATCTTCAGGCTCGCCTGGGACGTGAAGAAGCTCTCGTTCGACTACACCCCGAACTGGGGCCGCGGCAACCCCAACAACTTCATCGACACTGTCACCTTCCCGAAAGTCTTGACCGACAAGGCGTACACGTACCGCGTCGCCGTCTCCGGACGGAACCTCGGCGTGAAACCCTCGTACGCGGTCGAGAGCGACGGCTCGCAGAAGGTCAACTTCCTCGAGTACAACTCCGGGTATGGCATAGCGGACACGAACACGATCCAGGTGTTCGTTGTCGACCCCGACACCAACAACGACTTCATCATCGCCCAGTGGAACTAGGAGGAGGCAGTGACTGACCCCTGGCGGTCTAATCTGCGGGCCACTGTGGGGGAGGGGCATCGCCCATCAGCTTCCTTGTCTCATAATCATGCCCGTGTAACAATTGTAAATCGACCTTGTTGTACCATGCATCCAGCTTTTGTGGTGTGCCGTCTTATGTTTGGTTG
For this optimization, I used a model optimized for Komagataella phaffii (recognized as Pichia pastoris) because I believe that for the first tests it would be good to apply it to a closely compatible model, such as a yeast with high expression capacity that resembles the eukaryotic model of other fungi but with faster growth and productivity under controlled conditions.
🟦 The next part of the exercise involves optimizing the codon of the nucleotide sequence using an online tool. In this case, I chose the Codon Optimization Tool from Integrated DNA Technologies.
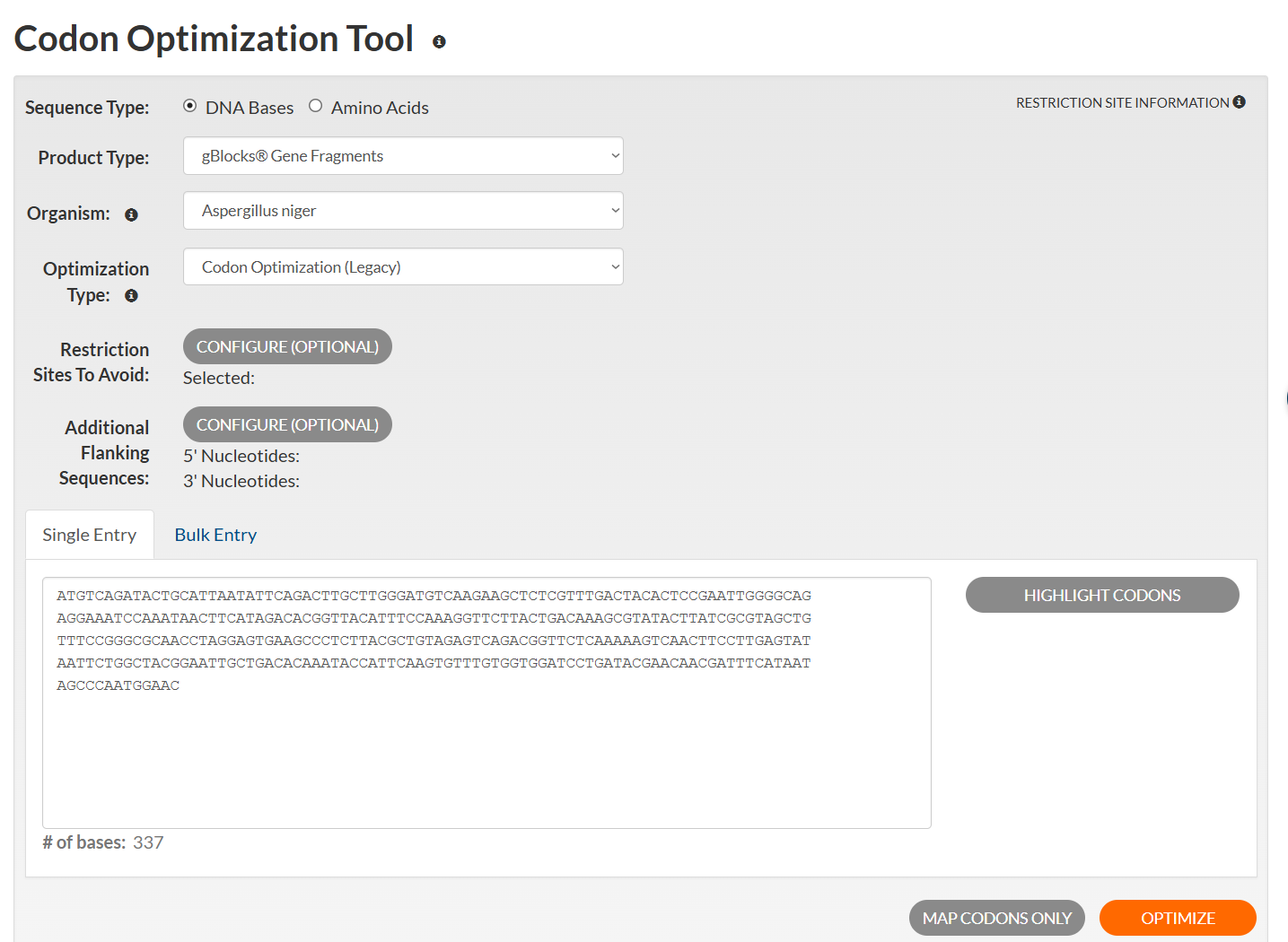

Optimized sequence
AAG TGG CAT TAC ATC ACC TTC GGA TAC AGA CAT CCA TAA CCT ACC AGT CAG CCA CGT GAT GGA AGA CAT CTG GAG GTG ACC ACC AAG GCT GTC TTT CGT CAG CTA TGG TCC AGG CGT CGT GAA GCC TTC ACC AGA CTT GAT ATT TCT GGA CAA TTT TGT CAA ACG AGG AAG GAA AAT GTG CCC TCA TAC CAT TTG CAG TGG TCC GCT GGT GTT GCT TCC AGA TCT ACT TGT GGC GCT ATT AGA TTT CCT GTG GCC TGT AGA GCT TCC CAT GTG GTT TGT ACT TTA GTT TAC ATC ATT GTT AAA AGG AGA AGT GGA TGT CGT ATT AAG GGA AGG CAG AGA AGA CCA GTC GAT TTT TCA ACG TGT TCA TGT ACC CAT TTT TAA TGA GTG GCA TAC CAC CTG AGT TAG CAA CGT CAA GTG CAT TAA TTG GTT CAT AGT CCA CAG CTA TTT GCC TTA CAA TCT AGA TTT GCT GAT ACA CTG TTT GAA CCA AGT CCA TTC AAT AAT CCC CTT ACC TCT GCT CCT CAA CAC CAT GTT AGA CAC TGT TTG GAT TTA CAA GCT CGT TTA GGA CGT GAG GAG GCA TTG GTG AGA TTG CAC CCA GAA CTA GGT CCT AGA CAG CCC CAA CAA CTT CAT AGA CAC TGC CAT TTA CCA GAA AGT TTA GAT AGA CAA GGT GTG CAT GTG CCA AGG AGA AGA TTG CGT ACT GAG CCA AGA AGA GAA ACT TTG GTT AGA GGT AGA GAG AGA AGA TTA GCA GAG GGT CAG TTG CCA AGG GTG CAA TTA CGT GTT TGG CAT TCT GGA CAT GAA CAT GAT CCA GGT GTA AGA TGT AGG CCA AGA CAT CAA CAA AGG TTG CAC CAT AGA CCA GTA GAA CTG GGT GGA GGC TCA GAT TAG CCT CTG GCA GTC TAG TCT GCT GGA CAC TGT GGA GGC GGA GCT TCA CCA ATT TCT TTT TTG GTG AGT TAA AGT TGT CCA TGT AAT AAT TGT AAG TCT ACC TTG CTT TAT CAC GCT TCC TCC TTT TGC GGA GTT CCT TCT TAT GTC TGG TTG
- To generate this protein, I believe that the most appropriate approach is to modify the Pichia pastoris yeast by inserting an expression vector using a strong promoter coupled to my optimized gene.
I believe this is the most feasible approach, considering this organism’s ability to accept the plasmid, integrate it into its transcriptional machinery, and generate multiple transcripts induced by a selective culture condition. The transcripts may undergo transcriptional modifications, be recognized by ribosomes, and be translated efficiently. I hope that the capabilities of P. pastoris will allow the necessary post-translational modifications to be made in order to fold the protein correctly, package it, and excrete it into the medium.
How does it work in nature/biological systems?
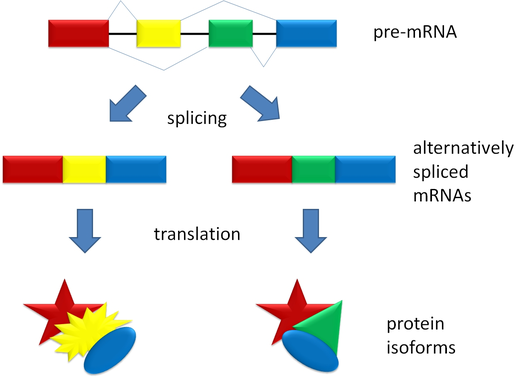
During post-transcriptional processes, there is a stage known as Alternative Splicing, which is a process that occurs after the removal of introns from the messenger RNA sequence, where the translatable fragments (exons) can be rearranged or omitted to form the final transcript. In this process, the order of the sequences can give rise to different forms of the same peptide, known as isoforms. These isoforms can have different folding, activity, and efficiency depending on the arrangement, which can respond to different circumstances depending on the conditions of the cell.
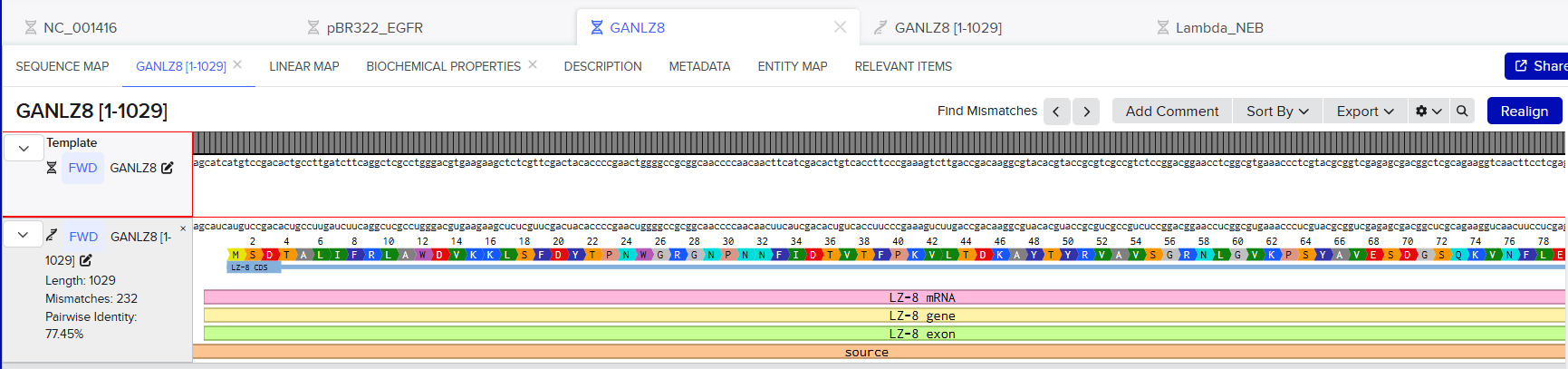

- I performed this alignment using Benchling’s alignment tool. First, I generated a copy of the RNA-transcribed sequence, and then I aligned it, including the amino acid and protein markers.
🪢 Twist DNA Synthesis Order
In this part, I built my vector with the following sequences:
Promoter BBa_J23106:
TTTACGGCTAGCTCAGTCCTAGGTATAGTGCTAGCRBS BBa_B0034:
CATTAAAGAGGAGAAAGGTACCStart Codon:
ATGCoding Sequence M58032.1 Ganoderma lucidum immunomodulatory protein (LZ-8) optimized:
AAGTGGCATTACATCACCTTCGGATACAGACATCCATAACCTACCAGTCAGCCACGTGATGGAAGACATCTGGAGGTGACCACCAAGGCTGTCTTTCGTCAGCTATGGTCCAGGCGTCGTGAAGCCTTCACCAGACTTGATATTTCTGGACAATTTTGTCAAACGAGGAAGGAAAATGTGCCCTCATACCATTTGCAGTGGTCCGCTGGTGTTGCTTCCAGATCTACTTGTGGCGCTATTAGATTTCCTGTGGCCTGTAGAGCTTCCCATGTGGTTTGTACTTTAGTTTACATCATTGTTAAAAGGAGAAGTGGATGTCGTATTAAGGGAAGGCAGAGAAGACCAGTCGATTTTTCAACGTGTTCATGTACCCATTTTTAATGAGTGGCATACCACCTGAGTTAGCAACGTCAAGTGCATTAATTGGTTCATAGTCCACAGCTATTTGCCTTACAATCTAGATTTGCTGATACACTGTTTGAACCAAGTCCATTCAATAATCCCCTTACCTCTGCTCCTCAACACCATGTTAGACACTGTTTGGATTTACAAGCTCGTTTAGGACGTGAGGAGGCATTGGTGAGATTGCACCCAGAACTAGGTCCTAGACAGCCCCAACAACTTCATAGACACTGCCATTTACCAGAAAGTTTAGATAGACAAGGTGTGCATGTGCCAAGGAGAAGATTGCGTACTGAGCCAAGAAGAGAAACTTTGGTTAGAGGTAGAGAGAGAAGATTAGCAGAGGGTCAGTTGCCAAGGGTGCAATTACGTGTTTGGCATTCTGGACATGAACATGATCCAGGTGTAAGATGTAGGCCAAGACATCAACAAAGGTTGCACCATAGACCAGTAGAACTGGGTGGAGGCTCAGATTAGCCTCTGGCAGTCTAGTCTGCTGGACACTGTGGAGGCGGAGCTTCACCAATTTCTTTTTTGGTGAGTTAAAGTTGTCCATGTAATAATTGTAAGTCTACCTTGCTTTATCACGCTTCCTCCTTTTGCGGAGTTCCTTCTTATGTCTGGTTG7x His Tag:
CATCACCATCACCATCATCACStop Codon:
TAATerminator BBa_B0015:
CCAGGCATCAAATAAAACGAAAGGCTCAGTCGAAAGACTGGGCCTTTCGTTTTATCTGTTGTTTGTCGGTGAACGCTCTCTACTAGAGTCACACTGGCTCACCTTCGGGTGGGCCTTTCTGCGTTTATA
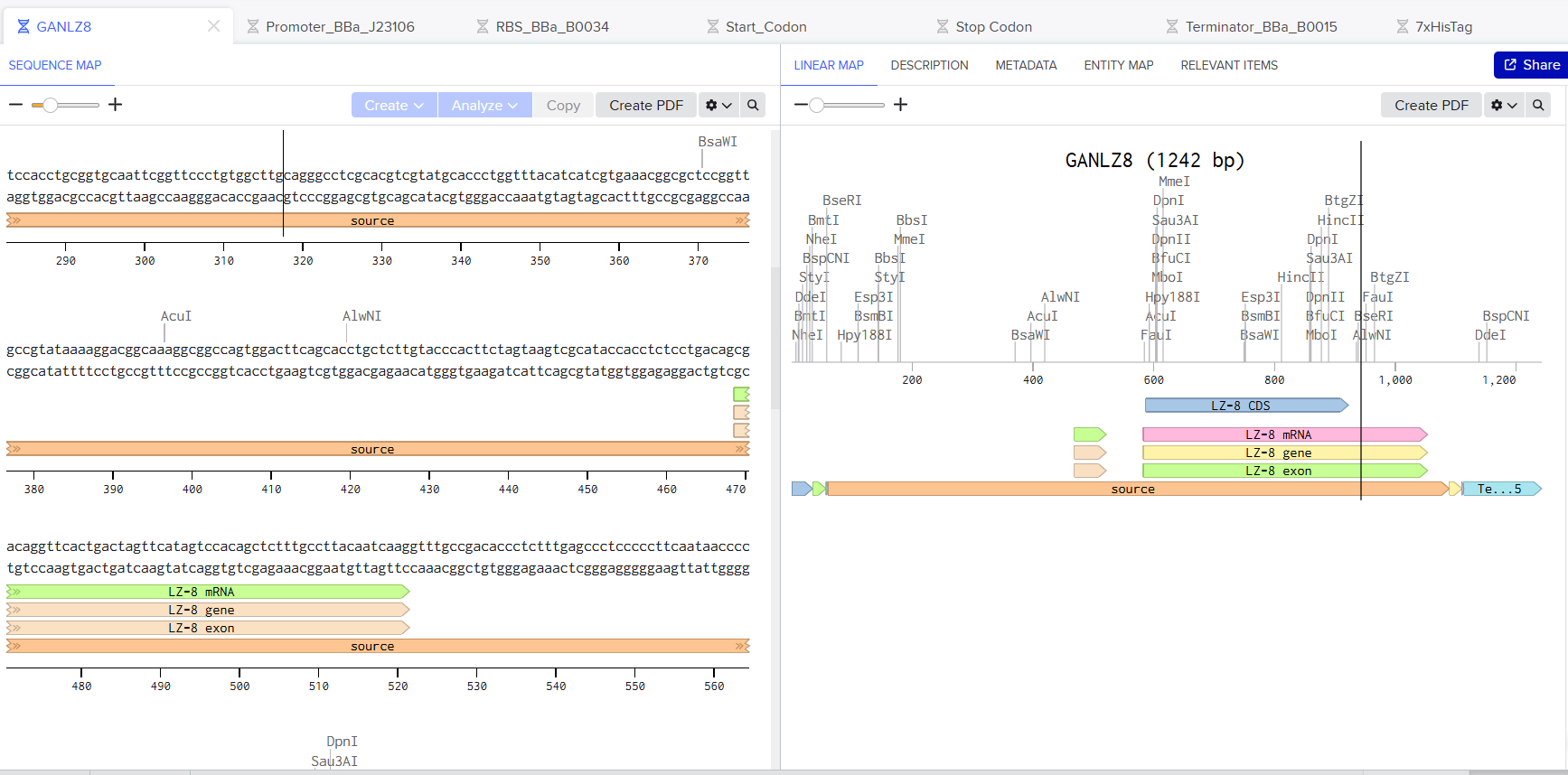

In this part of the activity, I should be working on the Twist Bioscience platform. Unfortunately, after creating my account, the platform did not allow me to create my vector 😿 due to restrictions in my geographical area, which prevents me from accessing the tools as shown in the following image.

📝 DNA Read/Write/Edit
👓 Read
- What DNA would you want to sequence (e.g., read) and why?From my project to this exercise, I have focused on obtaining information on fungal genes with potential therapeutic functionality as a way to generate drugs or solutions based on secondary metabolites that stimulate the human defense system and inhibit the growth of pathogenic organisms since there are many fungi that I believe conceal their incredible machinery for combating other microorganisms or even controlling conditions such as cancerous tumors or viral infections.
- What technology or technologies would you use to perform sequencing on your DNA and why?Currently, two generations of technology are available: Illumina sequencing (2nd generation) can be used to obtain complete reference genomes or perform gene expression studies (RNA-seq) with good accuracy, high throughput, and low cost. There are also third-generation technologies such as Pacific Biosciences (PacBio) and MinION from Oxford Nanopore Technologies (ONT); both produce readings of tens of thousands of bases, ideal for resolving complex genomic structures and obtaining high-quality assemblies that reach the level of a complete chromosome . I can combine these technologies to ensure high accuracy for in-depth studies of fungal diversity in complex samples (metabarcoding).
- What is your input? How do you prepare your input (e.g. fragmentation, adapter ligation, PCR)?To sequence the DNA of my fungi, high molecular weight DNA is mechanically fragmented to obtain the desired size: for Illumina, short fragments of 200-500 bp are sought using ultrasound; while for PacBio or ONT, long fragments (>10 kb) are required using a device such as Covaris g-TUBEs, which allows for extensive reads that resolve complex regions of the fungal genome. Next, the ends of the fragments are repaired and an adenine tail is added to facilitate the ligation of specific adapters to each platform. In addition, if the amount of DNA is limited, PCR amplification is performed to generate a sufficient library for sequencing.
- What are the essential steps of your chosen sequencing technology, how does it decode the bases of your DNA sample (base calling)?The essential steps of PacBio begin with the preparation of an SMRTbell library, where hairpin adapters are ligated to DNA fragments to create a circular template. This is immobilized in a Zero-Mode Waveguide (ZMW) with a DNA polymerase. To decode the bases, the polymerase incorporates nucleotides carrying a fluorophore attached to the phosphate group. Upon incorporation, the fluorophore is cleaved and detected in real time, identifying the specific base.
- What is the output of your chosen sequencing technology?PacBio produces long reads (10-25 kb) with high accuracy. The result is sequence data in HiFi (High Fidelity) format with >99.9% accuracy, ideal for assembling complete genomes and resolving complex regions.
⌨️ Write
- What DNA would you want to synthesize (e.g., write) and why?I would like to synthesize expression vectors containing sequences that encode the production of antimicrobial and antiviral compounds or peptides (such as the immunomodulatory protein LZ-8).
- What technology or technologies would you use to perform this DNA synthesis and why?As reviewed during the course, I would use high-throughput chemical synthesis on chip platforms such as the model offered by Twist Bioscience (provided they allow me access to their platform) due to its scalability and low cost. In the case of long fragments or complex sequences, enzymatic synthesis with TdT is ideal, as I have read that it operates under mild conditions and generates high-fidelity DNA without toxic solvents.
- What are the essential steps of your chosen sequencing methods?Chemical synthesis on a chip involves the synthesis of oligonucleotides on a silicon platform with thousands of nanocavities, using phosphoramidite chemistry in miniaturized reaction volumes (100 nL). while enzymatic synthesis with TdT involves iterative cycles where a directed evolution-modified terminal deoxynucleotidyl transferase (TdT) adds nucleotides blocked with 3’-phosphate groups, followed by gentle enzymatic unblocking.
- What are the limitations of your sequencing method (if any) in terms of speed, accuracy, scalability?The limitations of the chemical method are its dependence on toxic organic solvents and a drop in yield for fragments >1,800 bp, although it offers high scalability (9,600 genes per chip) and reduced cost (~10 cents/base). Performing the same analysis as the enzymatic method reveals that there is residual sequence bias and lower commercial maturity, although this is offset by greater speed (90 sec/cycle), coupling efficiency >99%, an environmentally friendly aqueous process, and ultra-long reads.
🔀 Edit
- What DNA would you want to edit and why?It might be possible to edit genes associated with fungal pathogen effectors to see how they affect virulence and pathogenicity.
- What technology or technologies would you use to perform these DNA edits and why?I’m considering implementing the CRISPR/Cas9 system, as it is the most efficient and versatile model for genetic editing in filamentous fungi. Recent studies show that CRISPR achieves recombination frequencies five times higher than traditional methods such as transformation mediated by other microorganisms such as Agrobacterium. Its superiority lies in its ability to generate specific cuts guided by a guide RNA, allowing everything from simple gene disruptions to multiple edits and gene regulation, significantly accelerating the functional exploration of genes in these genetically complex organisms.
- How does your technology of choice edit DNA? What are the essential steps?According to what I have read, CRISPR/Cas9 edits by generating double-strand breaks (DSBs) in DNA at a precise location determined by the guide RNA (sgRNA). The Cas9 endonuclease, guided by the sgRNA, recognizes a PAM sequence (5′-NGG-3′) and cuts both strands, producing blunt ends. Subsequently, the cellular repair machinery acts: either through NHEJ (non-homologous end joining), which introduces insertions/deletions that inactivate the gene, or through homologous repair (HR), which incorporates a donor DNA template to introduce precise modifications such as insertions or substitutions.
- What preparation do you need to do (e.g. design steps) and what is the input (e.g. DNA template, enzymes, plasmids, primers, guides, cells) for the editing?Employing this technology requires a bioinformatic design of the guide RNA (sgRNA) complementary to the genomic target, as well as the construction of the vector with Cas9 and sgRNA, or preparation of the Cas9-sgRNA ribonucleoprotein complex (RNP). In the case of HR editing, a DNA donor template with homology arms (500-2000 bp flanking the region to be modified) and a selectable marker such as hygromycin are required. In addition, the Cas9 enzyme, ARNg, restriction enzymes, high-fidelity polymerases, and transformation reagents such as PEG and protoplast buffers are required.
- What are the limitations of your editing methods (if any) in terms of efficiency or precision?The main limitation is multinucleation and heterokaryosis in filamentous fungi, considering that cells contain multiple nuclei, and unedited nuclei persist during regeneration, making it difficult to obtain stable homokaryotic strains. Off-target effects also persist, where NgRNA recognizes unwanted similar sequences, although Cpf1 (Cas12a) shows a lower incidence than Cas9.
5 Technogenesis: Aesthetic Dimensions of Art and Biotechnology ↩︎
Restriction enzymes and their use in molecular biology: An overview ↩︎
Fungal immunomodulatory proteins (FIPs) from Ganoderma lucidum: Unveiling their immunomodulatory mechanisms and potential for cancer therapy ↩︎
High-Level Expression, Purification and Production of the Fungal Immunomodulatory Protein-Gts in Baculovirus-Infected Insect Larva ↩︎
Next-Generation Sequencing (NGS): Platforms and Applications ↩︎
Chapter Six - Enzymatic synthesis and modification of high molecular weight DNA using terminal deoxynucleotidyl transferase ↩︎
Scaling DNA synthesis with a microchip-based massively parallel synthesis system. ↩︎
CRISPR-Cas9 genome editing approaches in filamentous fungi and oomycetes ↩︎
Recent Advances in Genome Editing Tools in Medical Mycology Research ↩︎