Week 2 HW: DNA: Read, Write, and Edit


Class Assignment
Part 1: Benchling & In-silico Gel Art
Make a Benchling account. Import Lambda DNA. Simulate RE digestion using EcoRI, HindIII, BamHI, KpnI, EcoRV, SacI, and SalI. Create a Gel Art Pattern
The Benchling file is linked here: “https://benchling.com/s/seq-8fknpF6pJKrFg2o79vYc?m=slm-T9RMQvF1LswIKEhWWpSP
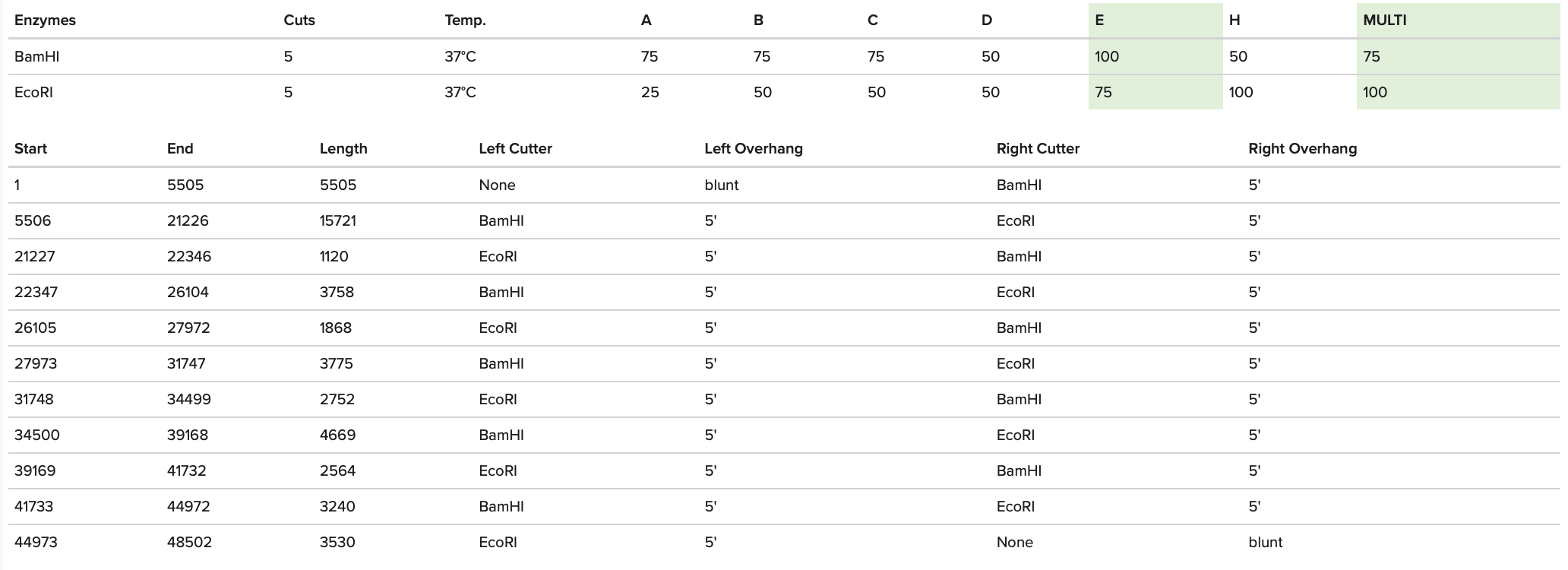
I imported the annotated Lambda DNA and ran a digest using the enzymes listed above.
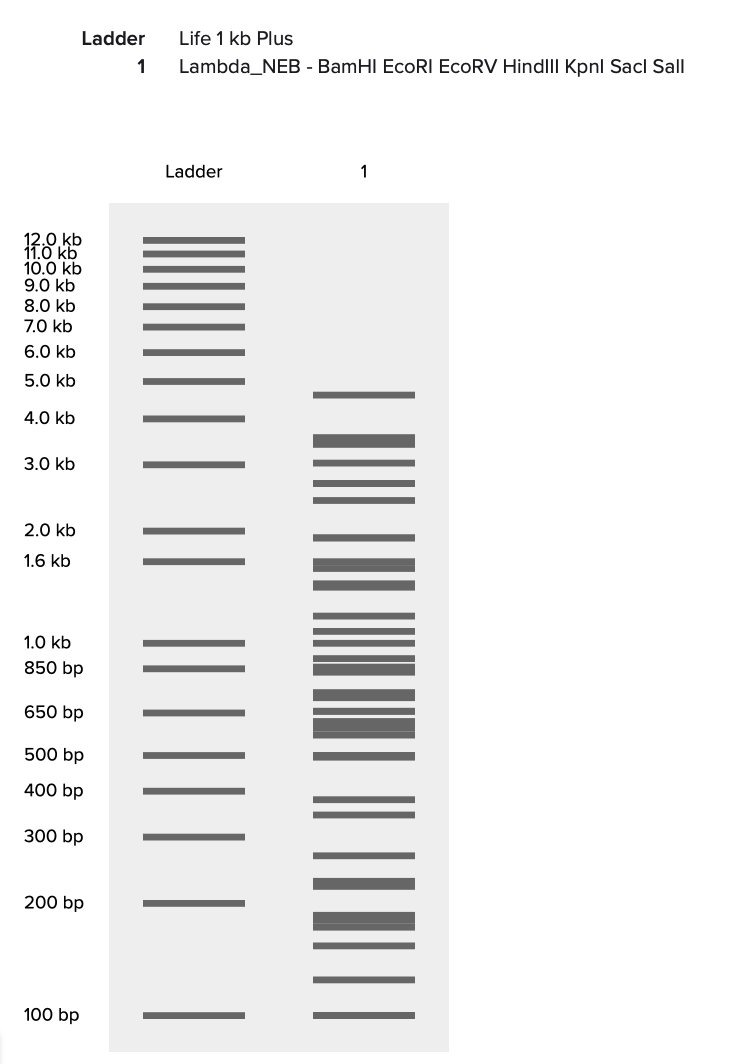
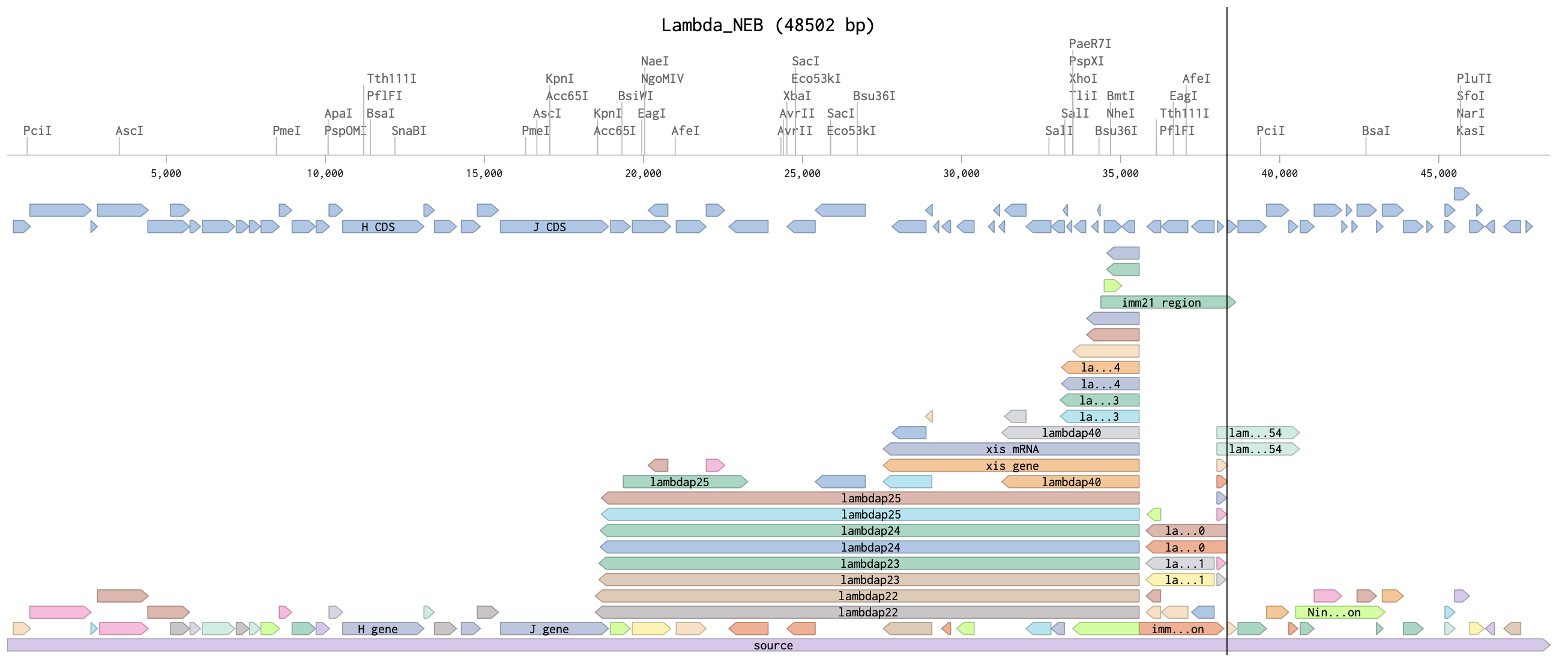



I then created a gel art pattern resembling a heart (to the best of my abilities!!!). To do so, I first simulated the gel run using DNA Gel Art Interface, before replicating the experiment on Benchling.
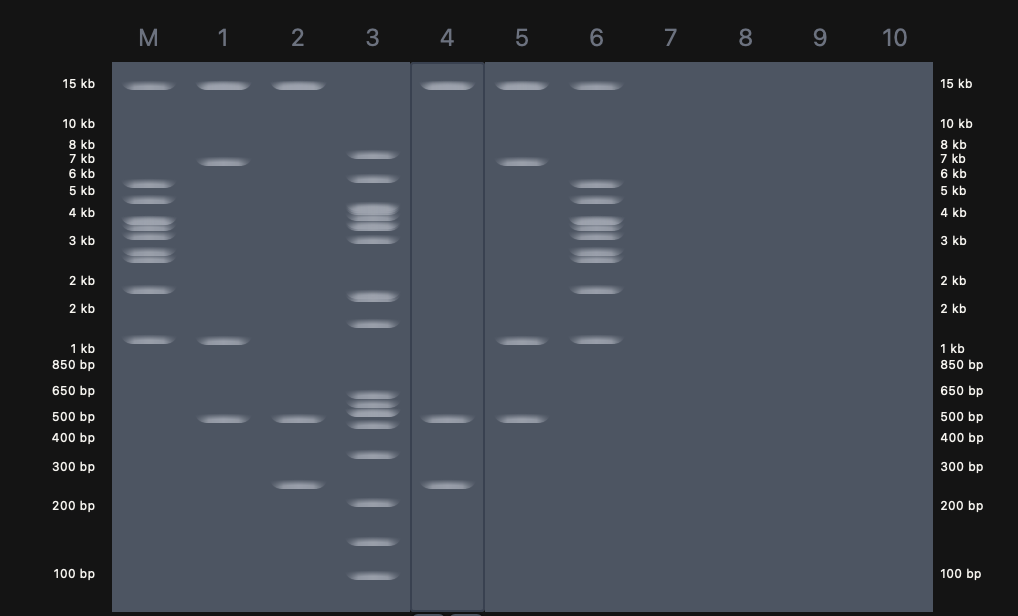

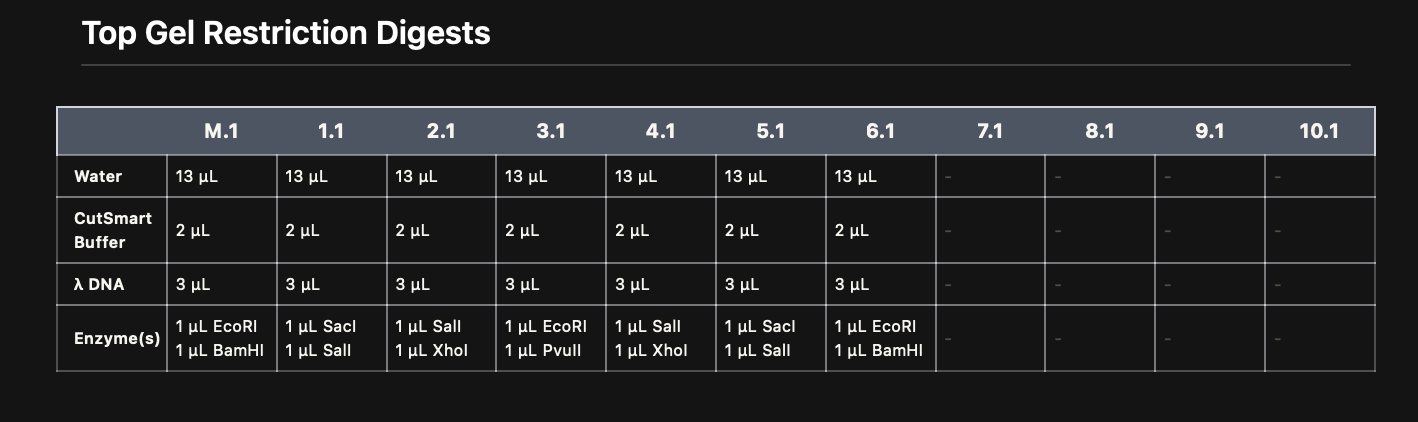

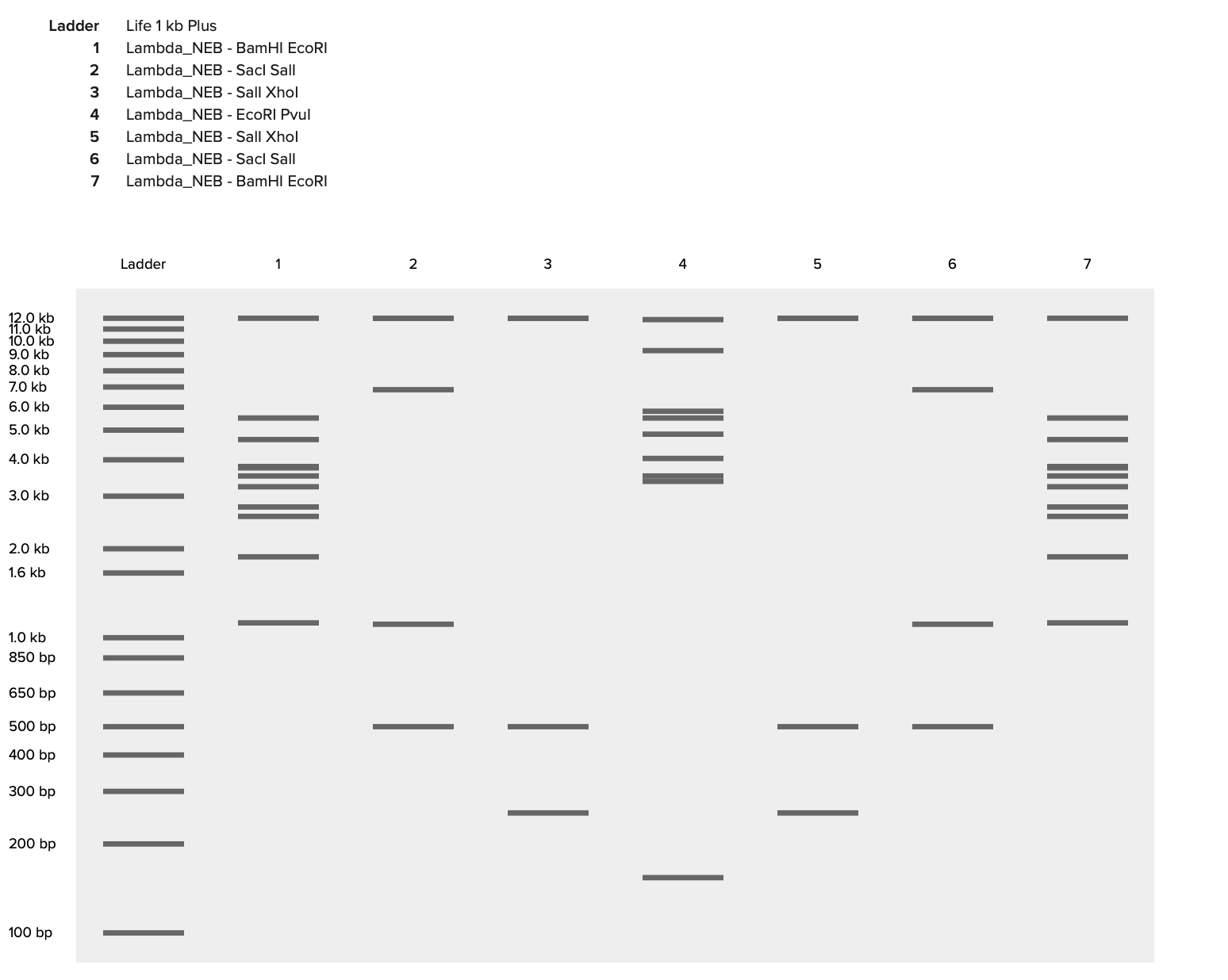


Part 3: DNA Design Challenge
In recitation, we discussed that you will pick a protein for your homework that you find interesting. Which protein have you chosen and why? Using one of the tools described in recitation (NCBI, UniProt, google), obtain the protein sequence for the protein you chose.
3.1. Choose your protein.
The protein I picked is luciferase, a bioluminescent protein that offers a sustainable, highly efficient alternative to high-intensity, artificial lighting any can aid in tackling light pollution.
The NCBI accession of luciferase from Armillaria gallica is A0A2H3E985.
3.2. Reverse Translate: Protein (amino acid) sequence to DNA (nucleotide) sequence.
Amino Acid Sequence: MSFIDSMKLDLVGHLFGIRNRGLAAACCALAVASTIAFPYIRRDYQTFLSGGPSYAPQNI RGYFIVCVLALFRQEQKGLAIYDRLPEKRRWLPDLPPRNGPRPITTSHIIQRQRNQAPDP KFALEELKATVIPRVQARHTDLTHLSLSKFEFHAEAIFLLPSVPIDDPKNVPSHDTVRRT KREIAHMHDYHDFTLHLALAAQDGKEVVSKGWGQRHPLAGPGVPGPPTEWTFIYAPRNEE ELAVVEMIIEASIGYMTNDPAGVVIA
Nucleotide Sequence: atgagctttattgatagcatgaaactggatctggtgggccatctgtttggcattcgcaac cgcggcctggcggcggcgtgctgcgcgctggcggtggcgagcaccattgcgtttccgtat attcgccgcgattatcagacctttctgagcggcggcccgagctatgcgccgcagaacatt cgcggctattttattgtgtgcgtgctggcgctgtttcgccaggaacagaaaggcctggcg atttatgatcgcctgccggaaaaacgccgctggctgccggatctgccgccgcgcaacggc ccgcgcccgattaccaccagccatattattcagcgccagcgcaaccaggcgccggatccg aaatttgcgctggaagaactgaaagcgaccgtgattccgcgcgtgcaggcgcgccatacc gatctgacccatctgagcctgagcaaatttgaatttcatgcggaagcgatttttctgctg ccgagcgtgccgattgatgatccgaaaaacgtgccgagccatgataccgtgcgccgcacc aaacgcgaaattgcgcatatgcatgattatcatgattttaccctgcatctggcgctggcg gcgcaggatggcaaagaagtggtgagcaaaggctggggccagcgccatccgctggcgggc ccgggcgtgccgggcccgccgaccgaatggacctttatttatgcgccgcgcaacgaagaa gaactggcggtggtggaaatgattattgaagcgagcattggctatatgaccaacgatccg gcgggcgtggtgattgcg
3.3. Codon optimization.
Codon optimization is a pre-processing step in recombinant DNA design which takes into account the chassis’s codon bias – the non-random, preferential use of specific synonymous codons over others to encode the same amino acid – in order to optimize the expressed protein’s translation, folding, and quantity. This works because, although the genetic code is degenerate, different organisms favor different codons. If a cognate tRNA is unavailable during protein synthesis, the amino acid’s translation may halt and impact the final product.
Since A. gallica is not typically used industrially, E. coli will be used. Thus, the nucleotide sequence will be codon optimized to E. coli.
Optimized Nucleotide Sequence:
ATG AGC TTT ATT GAT AGC ATG AAA CTG GAC CTG GTC GGC CAC CTG TTT GGT ATC CGC AAC CGC GGC CTG GCG GCG GCC TGC TGT GCG CTG GCG GTG GCC AGC ACC ATT GCC TTC CCG TAT ATC CGT CGT GAT TAT CAG ACC TTC CTC TCC GGT GGG CCG AGC TAT GCT CCG CAG AAC ATC CGC GGT TAT TTT ATC GTC TGT GTG CTG GCG CTG TTC CGC CAG GAG CAG AAA GGC CTG GCG ATT TAT GAC CGC CTG CCG GAG AAA CGC CGC TGG CTG CCG GAC CTG CCG CCG CGC AAC GGT CCG CGT CCG ATT ACC ACC AGC CAC ATC ATT CAG CGC CAG CGT AAC CAG GCA CCG GAT CCG AAA TTT GCG CTG GAA GAG CTG AAA GCG ACC GTG ATT CCG CGC GTG CAG GCG CGC CAC ACC GAC CTG ACC CAC CTC TCG CTG AGC AAG TTT GAG TTC CAT GCA GAA GCG ATC TTC CTG CTG CCG TCT GTG CCG ATC GAT GAT CCG AAA AAC GTG CCC AGC CAC GAC ACC GTG CGC CGC ACC AAA CGT GAG ATC GCC CAC ATG CAC GAC TAC CAC GAC TTC ACC CTG CAC CTG GCG CTG GCG GCA CAG GAT GGC AAA GAG GTG GTG AGC AAA GGC TGG GGG CAG CGC CAT CCG CTG GCG GGC CCG GGT GTG CCG GGC CCG CCG ACC GAA TGG ACC TTT ATT TAT GCG CCG CGC AAC GAA GAA GAG CTG GCG GTG GTT GAA ATG ATC ATT GAA GCC TCA ATC GGT TAC ATG ACC AAC GAC CCG GCA GGC GTG GTG ATT GCC
Part 4: Prepare a Twist DNA Synthesis Order
4.2. Build Your DNA Insert Sequence
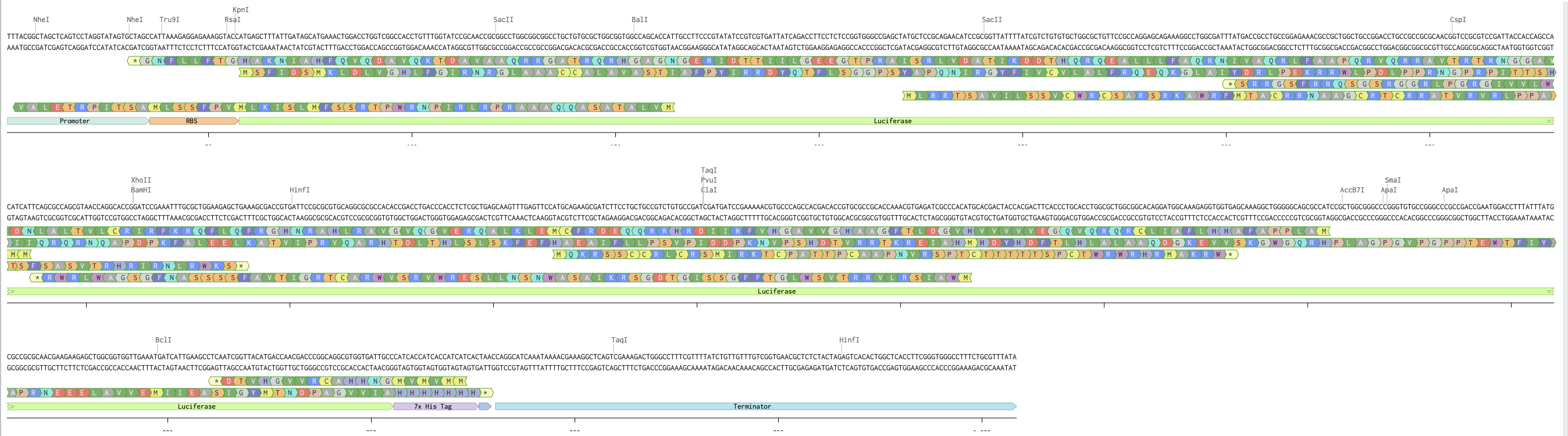
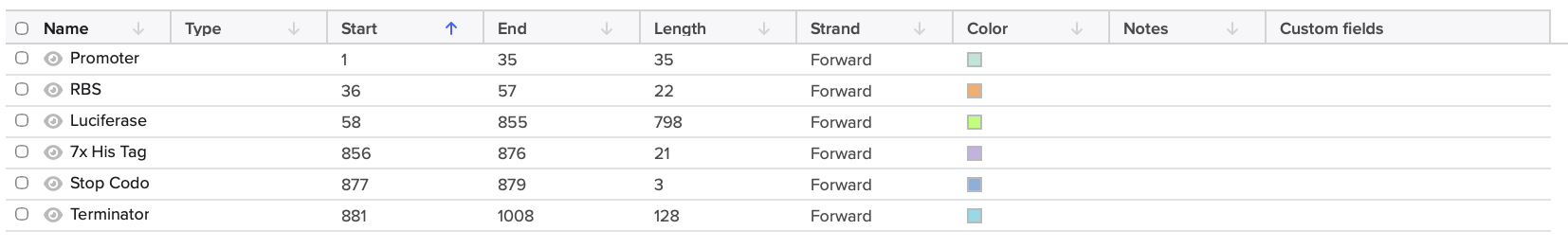
Using the listed promtoer, RBS, 7x His-tag, and terminator, I assembled and annotated the gene expression casette below.





Part 5: DNA Read, Write, Edit
5.1 DNA Read
(i) I would want to sequence the microbiome (MB) associated with a specific niche, such as the gut, focusing on both species abundance and functionalities. Bacteria co-evolved with a lot of species and thus play crucial roles in their health. In humans, this involves digestion, immunity, and even gut-brain health.
(ii) I would use Illumina sequencing for shotgun metagenomics. This is second-generation sequencing, where input DNA is extracted from stool, fragmented, adapter-ligated, PCR-amplified, and sequenced. Bases are decoded via fluorescence (Illumina), producing FASTQ files for downstream taxonomic and functional analysis.
5.2 DNA Write
(i) I would synthesize a genetic circuit in gut bacteria that senses inflammation and responds by producing a therapeutic molecule, such as an anti-inflammatory peptide or short-chain fatty acid. This enables localized, on-demand treatment of gut disorders while minimizing systemic side effects. The circuit would include a sensor promoter, regulatory logic, and an effector gene.
(ii) I would use phosphoramidite-based DNA synthesis to generate individual circuit components, followed by Gibson Assembly.
Essential steps include gene and oligo synthesis, genetic parts assembly, cloning, and validation (in vitro and in vivo). Limitations include sequence length constraints, synthesis errors, and batch-to-batch variations.
5.3 DNA Edit
(i) I would edit the genomes of gut-resident microbes to improve circuit stability, safety, and therapeutic performance. This could include knocking out competing pathways, tuning promoter strength, or enhancing stress tolerance to ensure reliable function in the gut environment.
(ii) I would utilize various forms of CRISPR technolgoies (e.g. Cas9, CRISPRa/CRISPRi) to introduce edits of different magnitdues as needed.