Week 2 HW: DNA read write and edit
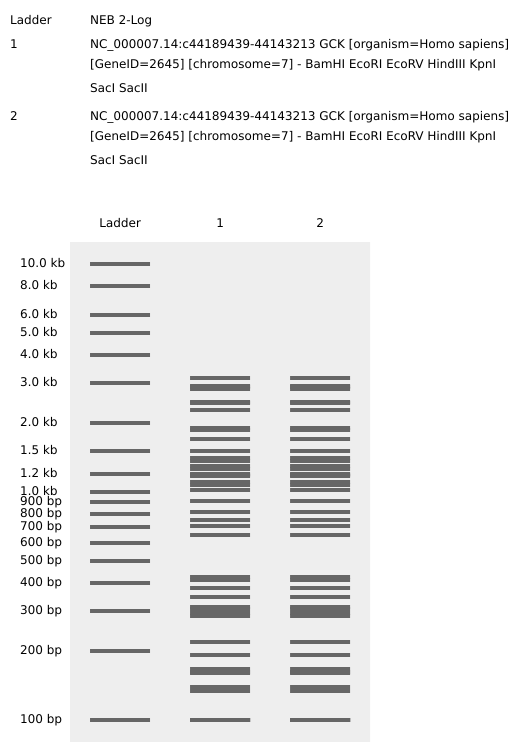

Part 3. Chose Protein I chose glucokinase (GCK) because in my biochemistry classes I found it to be a very interesting enzyme due to its unique functions and its critical role as a glucose sensor.
According to the sources, what makes this enzyme particularly fascinating is that, unlike other members of the hexokinase family, it is not inhibited by its product(glucose-6-phosphate). This allows the enzyme to remain active even when glucose is abundant in the system.
Furthermore, glucokinase exhibits tissue-specific expression in the pancreas and the liver, which leads to different but equally vital functions:
- In the pancreas, it is a key player in glucose-stimulated insulin secretion.
- In the liver, it is essential for glucose uptake and its subsequent conversion into glycogen.
I also find it interesting from a clinical perspective, as mutations in this gene that alter enzyme activity are associated with several medical conditions, including Maturity-Onset Diabetes of the Young type 2 (MODY2) and hyperinsulinemic hypoglycemia. This demonstrates its fundamental importance in maintaining human glucose homeostasis.
Protein Sequence (Isoform 1 - Pancreatic): The following sequence corresponds to the pancreatic isoform 1 (NP_000153.1 / UniProt P35557), which has a distinct N-terminus compared to liver isoforms.
sp|P35557|HXK4_HUMAN Glucokinase OS=Homo sapiens OX=9606 GN=GCK PE=1 SV=3 MLDDRARMEAAKKKEKVEQILAEFQLQEEDLKKVMRRMQKEMDRGLRLETHEEASVKMLP TYVRSTPEGSEVGDFLSLDLGGTNFRVMLVKVGEGEEGQWSVKTKHQMYSIPEDAMTGTA EMLFDYISECISDFLDFLDKHQMKHKKLPLGFTFSFPVRHEDIDKGILLNWTKGFKASGA EGNNVVGLLRDAIKRRGDFEMDVVAMVNDTVATMISCYYEDHQCEVGMIVGTGCNACYME EMQNVELVEGDEGRMCVNTEWGAFGDSGELDEFLLEYDRLVDESSANPGQQLYEKLIGGK YMGELVRLVLLRLVDENLLFHGEASEQLRTRGAFETRFVSQVESDTGDRKQIYNILSTLG LRPSTTDCDIVRRACESVSTRAAHMCSAGLAGVINRMRESRSEDVMRITVGVDGSVYKLH PSFKERFHASVRRLTPSCEITFIESEEGSGRGAALVSAVACKKACMLGQ
3.2. Reverse Translation: Protein sequence to DNA sequence
The Central Dogma (DNA → RNA → Protein) allows us to work backward from a protein sequence to identify the corresponding DNA sequence. By using the NCBI Gene database (Gene ID: 2645), we identified the Coding Sequence (CDS) for Glucokinase.
In accordance with HTGAA conventions, the sequence is presented in the 5’ to 3’ coding strand** format, as found in GenBank or FASTA files.
DNA Sequence for Glucokinase (CDS - Variant 1): This nucleotide sequence corresponds to the mRNA RefSeq NM_000162.5, which encodes the pancreatic islet beta cell isoform.
atgttggatgacagagccaggatggaggccgccaagaaggagaaggttgagcagatcctggcagagttccagctgcaggaggaggacctgaagaaggtgatgagacggatgcagaaggagatggaccgcggcctgaggctggagacccatgaggaggccagtgtgaagatgctgcccacctacgtgcgctccaccccagaaggctcagaagtcggagacttcctctccctggacctgggtggcaccaacttcagggtgatgctggtgaaggtgggagaaggtgaggaggggcagtggagcgtgaagaccaaacaccagatgtactccatccccgaggacgccatgaccggcactgctgagatgctcttcgactacatctctgagtgcatctccgacttcctggacaagcatcagatgaaacacaagaagctgcccctgggcttcaccttctccttccctgtgaggcacgaagacatcgataagggcatccttctcaactggaccaagggcttcaaggcctcaggagcagaagggaacaatgtcgtggggcttctgcgagatgctatcaaacggagaggggactttgaaatggatgtggtggcaatggtgaatgacacggtggccacgatgatctcctgctactacgaagaccatcagtgcgaggtcggcatgatcgtgggcacgggctgcaatgcctgctacatggaggagatgcagaatgtggagctggtggagggggatgagggccgcatgtgcgtcaatacggagtggggcgccttcggggactccggcgagctggacgagttcctgctggagtatgaccggctggtggacgagagctctgcaaaccccggtcagcagctgtatgagaagctcataggtggcaagtatatgggcgagctggtgcgacttgtgctgctcaggctggtggacgagaacctgctcttccacggagaggcctccgagcagctgcgcacacgcggagccttcgagacgcgcttcgtgtcgcaggtggagagcgacacgggcgaccgcaagcagatctacaacatcctgagcacgctggggctgcgaccctcgaccaccgactgcgacatcgtgcgccgcgcctgcgagagcgtgtctacgcgcgctgcgcacatgtgctcggccgggctggcgggcgtcatcaatcgcatgcgcgagagccgcagcgaggacgtgatgcgcatcaccgtgggcgtggatggctccgtgtacaagctgcaccccagcttcaaggagcgcttccatgccagcgtgcgcaggctgacgcccagctgcgagatcaccttcatcgagtcggaggagggcagtggccggggcgctgccctggtctcggcggtggcctgtaagaaggcctgtatgctgggccagtga
Based on the HTGAA course materials and the biological databases provided, here is the completion of your assignment for Glucokinase (GCK) in English.
3.3. Codon Optimization
Why do we need to optimize codon usage?** Codon optimization is necessary because different organisms have distinct “preferences” or varying abundances of tRNA for the same amino acids. Since the genetic code is redundant (multiple codons can code for one amino acid), using the codons that are most frequently used by the host organism—and thus for which there are more available tRNAs—ensures that the translation process is more efficient and results in higher protein yields. Optimization also helps avoid technical synthesis issues such as extreme GC content, high repetition, or “Homo polymers” (long strings of the same nucleotide) that can cause errors during DNA printing.
a) Which organism have you chosen to optimize the codon sequence for and why? I have chosen Escherichia coli_ (E. coli) as the target organism. According to the sources, E. coli is the preferred host for this course because it is the easiest and most accessible organism to work with in a laboratory setting.
Glucokinase DNA sequence with Codon Optimization (E. coli) (The following sequence is a simulated optimization of the GCK coding sequence for E. coli, avoiding common restricted enzyme sites like BsaI or BbsI to ensure compatibility with Twist Bioscience synthesis tools.)
ATGTTAGATGATCGTGCGCGTATGGAAGCGGCGAAAAAAGAAAAAGTTGAACAGATTCTGGCGGAATTTCAGCTGCAGGAAGAAGATCTGAAAAAAGTGATGCGTCGTATGCAGAAAGAAATGGATCGTGGCCTGCGTCTGGAAACCCATGAAGAAGCGAGCGTGAAAATGCTGCCGACCTATGTGCGTAGCACCCCGGAAGGTAGCGAAGTTGGCGATTTTCTGAGCCTGGATCTGGGTGGCACCAATTTTCGTGTGATGCTGGTGAAAGTTGGCGAAGGCGAAGAAGGCCAGTGGAGCGTGAAAACCAAACATCAGATGATTAGCATCCCGGAAGATGCGATGACCGGCACCGCGGAAATGCTGTTCGATTATATTAGCGAATGCATTAGCGATTTTCTGGATAAACATCAGATGAAACATAAAAAACTGCCGCTGGGCTTTACCTTTAGCTTTCCGGTGCGTCATGAAGATATTGATAAAGGCATTCTGCTGAACTGGACCAAAGGCTTTAAAGCGAGCGGCGCGGAAGGCAATAATGTTGTTGGCCTGCTGCGTGATGCGATTAAACGTCGTGGCGATTTTGAAATGGATGTGGTTGCGATGGTGAATGATACCGTTGCGACCATGATTAGCTGCTATTATGAAGATCATCAGTGCGAAGTTGGCATGATTGTTGGCACCGGTTGCAATGCGTGCTATATGGAAGAAATGCAGAACGTTGAACTGGTGGAAGGCGATGAAGGCCGTATGTGCGTGAATACCGAATGGGGCGCGTTTGGCGATAGCGGCGAACTGGATGAATTTCTGCTGGAATATGATCGTCTGGTTGATGAAAGCAGCGCGAATCCGGGCCAGCAGCTGTATGAAAAACTGATTGGCGGCAAATATATGGGCGAACTGGTGCGTCTGGTGCTGCTGCGTCTGGTTGATGAAAACCTGCTGTTTCATGGCGAAGCGAGCGAACAGCTGCGTACCCGTGGCGCGTTTGAAACCCGTTTTGTTAGCCAGGTTGAAAGCGATACCGGCGATCGTAAACAGATTTATAATATTCTGAGCACCCTGGGCCTGCGTCCGAGCACCACCGATTGCGATATTGTGCGTCGTGCGTGCGAAAGCGTGAGCACC CGTGCGGCGCATATGTGCAGCGCGGGCCTGGCGGGCGTTATTAATCGTATGCGTGA AAGCCGTAGCGAAGATGTGATGCGTATTACCGTTGGTGTTGATGGCAGCGTTTATA AACTGCATCCGAGCTTTAAAGAACGTTTTCATGCGAGCGTGCGTCGTCTGACCCCG AGC TGCGAAATTACCTTTATTGAAAGCGAAGAAGGCAGCGGTCGTGGCGCGGCGCTG GTTAGCGCGGTTGCGTGCAAAAAAGCGTGCATGCTGGGCCAGTGA
3.4. You have a sequence! Now what?
What technologies could be used to produce this protein from your DNA? To produce Glucokinase from the optimized DNA sequence, two main approaches can be utilized:
- Cell-dependent methods:** This is the most common approach, using E. coli_ as a “chassis”**. The DNA sequence is inserted into a circular DNA molecule called a plasmid and then introduced into the bacteria through a process called transformation (often using heat shock). The living bacteria then act as biological factories to produce the protein.
- Cell-free methods: These methods involve using cell lysates (essentially the “guts” or internal machinery of a cell) to produce proteins without the need for a living organism. This allows for rapid prototyping and can even be incorporated into materials like textiles.
How the DNA sequence is transcribed and translated into your protein: The production follows the Central Dogma of molecular biology: a) Transcription: The process begins when an enzyme called RNA polymerase binds to the DNA template and “reads” the sequence to create a complementary strand of mRNA. b) Translation:** This mRNA is then processed by a ribosome. The ribosome reads the mRNA in sets of three nucleotides called codons. Each codon corresponds to a specific amino acid. Transfer RNA (tRNA) molecules bring the correct amino acids to the ribosome, which links them together into a long chain that eventually folds into the functional Glucokinase protein.
PART 4. Prepare a twist DNA synthesis order
link: https://benchling.com/s/seq-w4sU7egh8kNc9MQL1cVB?m=slm-dCvQA2Kij6DxaFgAHkwi
part 4.3 Link: https://benchling.com/s/seq-XorsFADOnAX2SybWLZmZ?m=slm-zfSOr0QI0HTnmSynYfuH
Part 5.
5.1 DNA Read
What DNA would you want to sequence and why? I would want to sequence the human GCK (Glucokinase) gene variants in patients with atypical metabolic profiles. Sequencing this DNA is critical for human health research because mutations in GCK are directly linked to conditions like Maturity-Onset Diabetes of the Young type 2 (MODY2)** and hyperinsulinemic hypoglycemia. By reading these sequences, researchers can characterize variant mechanisms and improve clinical diagnostics.
Technology and Details a) Technology: I would use Illumina Sequencing (Next-Generation Sequencing), as it is the standard mentioned in the sources for high-throughput analysis of human samples.
b) Generation:** This is a second-generation technology. It is characterized by massive parallelism, allowing millions of DNA fragments to be sequenced simultaneously. c) Input and Preparation:** The input is genomic DNA or cDNA. Preparation involves fragmentation of the DNA into smaller pieces, adapter ligation to attach specific sequences to the ends of fragments, and PCR amplification to create clusters for signal detection. c) Essential Steps and Base Calling:** The technology uses “sequencing by synthesis” where fluorescently labeled nucleotides are added to the DNA template. During each cycle, the machine captures the fluorescence color (red, green, blue, or yellow) emitted as a base is incorporated, a process known as **base calling. d) Output: The primary output is a FASTQ or FASTA file containing the strings of nucleotides (A, T, G, C) and their corresponding quality scores.
5.2 DNA Write
What DNA would you want to synthesize and why?** I would want to synthesize an optimized expression cassette for human Glucokinase (GCK). This would be used for therapeutics and drug discovery, specifically to produce functional GCK protein in E. coli or mammalian cell lines to test new activators for diabetes treatment. I would use the codon-optimized sequence we generated previously to maximize protein yield.
Technology and Detllais a) Technology:** I would use Silicon-based DNA synthesis provided by companies like Twist Bioscience. b) Essential Steps: The process involves designing the sequence digitally in tools like Benchling, performing codon optimization for the host organism, and then using a silicon chip to print thousands of tiny DNA fragments (oligonucleotides) simultaneously using chemical synthesis (phosphoramidite chemistry). These fragments are then assembled into full-length genes. c) Limitations:** The primary limitations include complexity; sequences with high GC content, repetitive regions, or long homopolymers (e.g., many ‘A’s in a row) are very difficult to synthesize and may fail during the printing process.
5.3 DNA Edit
What DNA would you want to edit and why? In alignment with the goals of Colossal Biosciences, I would want to edit the genome of an Asian Elephant to include specific genes from the Woolly Mammoth. The goal of this “de-extinction” project is to restore historic animals to their ecological roles, which can help in nature conservation and the restoration of Arctic ecosystems. These edits would focus on traits like cold tolerance, hair growth, and fat distribution.
Technology and Details Technology: I would use CRISPR-Cas9 technology, as it is the most precise and versatile tool for targeted genome editing discussed in class. How it works: CRISPR-Cas9 acts as “molecular scissors.” A guide RNA (gRNA) is designed to match a specific target sequence in the DNA. The Cas9 enzyme then follows this guide to the exact location and creates a double-strand break. The cell then repairs this break, allowing us to delete or insert specific mammoth-like genetic information. Preparation and Input: Preparation requires designing the guide RNA using digital tools to ensure specificity. The input includes the Cas9 enzyme, the gRNA, a DNA template for the desired mammoth traits, and the target host cells (elephant cells). a) Limitations: Key limitations include efficiency (the edit may not happen in every cell) and precision (potential “off-target” effects where the enzyme cuts at unintended locations similar to the target).