Week 1 HW: Principles & Practices

🦠Brighter Autonomous Bioluminescence🦠
I would love to improve the intensity of the glow that is emitted from autonomous bioluminescent organisms whether natural or synthetic.
There are several different organisms that produce bioluminescence through various forms of luciferases (the enzyme that catalyzes the light emitting reaction) and luciferins (the substrate). However, most of them require the addition of the substrate to the growing medium to induce bioluminescence, typically coelenterazine or D-Luciferin. This to me just does not seem like the most convenient way to do this, so I am more interested in autonomous bioluminescent systems, such as Lux (bacterial luciferase) and Luz (fungal luciferase). These systems are the only two bioluminescent systems that have been fully elucidated. This means that they are fully genetically encoded, cells express luciferase and the enzymes necessary for substrate synthesis. This enables continuous supply of substrate without having to worry about adding the substrate to the growing medium or tissues to produce a glow.
The first bioluminescent organism I ever cultivated was the fungus Panellus stipticus. The culture was given to me by a mycologist I was working with at the time. In order to get P. stipticus to glow I was directed to subculture onto bread crumb agar (simply agar and bread crumbs from the grocery store). Once cultured on the bread crumb agar the P. stipticus cultures did glow! However, this is where it should be noted that the glow that is produced from bioluminescence is typically not very bright. I will include a picture that I had taken of my first plate, the glow seems brighter than what it actually appears in person due to a longer exposure time on my iPhone camera settings.

While in undergrad we transformed cells to produce GFP in an Advanced Microbiology Lab course.
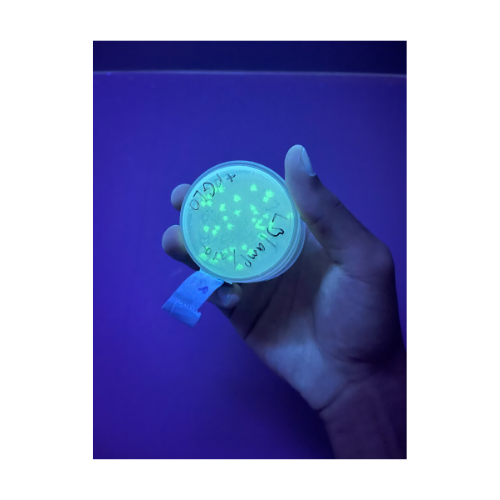


You can see the difference in how bright the glow is even with the long exposure time on the photo for the bioluminescent fungus. However, it should be noted in order to get organisms to produce fluorescence with GFP light is used. Whereas the glow from the bioluminescence may not be as bright there is no need for any other light sources to see the glow. So, I would prefer to use bioluminescence as there is no need to use light.
Then most recently, about a week ago, I transformed E. coli, on my own, with pVIB to produce bioluminescent cells. Again, I found myself with the same feeling that I had all those years ago when I first cultured P. stipticus, this is amazing but not exactly what I had imagined.
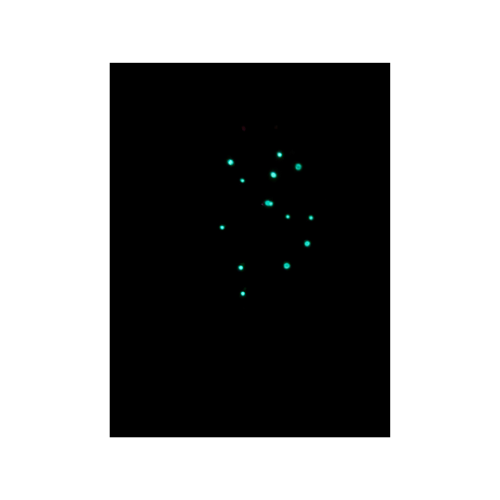


So now through this class I would like to take the opportunity to see if it would be possible to improve brightness/intensity of the glow produced from bioluminescent organisms!
Governance Goals
Goal 1: Ensure Safety & Prevent Harm
Biocontainment Standards: Making sure that containment strategies are in place that require genetic safeguards (e.g., inhibiting reproduction, auxotrophy or kill switches). This would prevent survival outside of intended environments. Safeguards should be validated and verified before deployment or release.
Operational Biosafety Protocols: Establish biosafety training and certification for anyone working with engineered bioluminescent organisms, including DIY biologists. This would mirror existing requirements in higher-biosafety labs and reduce risks posed by accidental exposure or poor technique.
Goal 2: Equity & Access
Equitable Access to Research Tools: Support public funding or bioluminescence kits for educational institutions and community labs with built-in safety guidelines, so that whoever is interested can participate responsibly in this area of science.
Ethics Education: Integrate ethics and governance training into the curriculum for any program funding or advising engineered organism projects, helping ensure researchers understand broader societal impact and their responsibilities.
Governance Actions
1) Community Driven Codes of Conduct for DIY and Institutional Labs
A community code of conduct would address the current gap between informal safety norms and clear expectations for work with engineered organisms. A coalition of academic groups, community labs, and DIYbio chapters could create a public code of conduct for responsible biodesign describing standards for organism handling, transparent documentation, and peer review, with voluntary adoption by spaces that host workshops or shared laboratories. This approach assumes that practitioners care about reputation and will align behavior with visible community values, though voluntary norms may struggle to reach groups that want independence over guidance. The main risk of failure is adoption without real behavioral change, while success could normalize safer habits and provide newcomers with a clear ethical baseline that reduces harm from inexperience.
2) Community Lab Micro-Grant Network
A micro-grant network would expand access to research by providing small, flexible funding to community labs, schools, and independent creators who lack institutional backing. Foundations, universities, or regional science hubs could distribute grants of $500–$2,000 paired with basic safety mentorship and simple reporting requirements, allowing projects in education, art, and environmental sensing to flourish. This approach assumes that modest resources can meaningfully broaden participation. The risk is that funds could be captured by already privileged groups or used without adequate guidance, but success would mean a more diverse ecosystem of synthetic biologist and ensure that the future of synthetic biology reflects many voices.
3) Peer Audit & Recognition Program for Safety Practices
A voluntary peer audit network would replace external enforcement with collaborative review, drawing on models from open source software and engineering design critique. Community organizations, DIYbio groups, iGEM teams, and university clubs could review one another’s safety documentation, containment strategies, and workflows, with participating projects receiving a public “Safety Verified” recognition. The model assumes that practitioners will invest time in mutual review and that social trust can motivate improvement, which may not hold equally across all groups. Audits could fail if reduced to paperwork exercises, yet success would foster a culture where safety is shared, allowing many eyes to strengthen emerging technologies.
Scoring of Governance Actions Against Policy Goals
Scale: 1 = best alignment, 2 = moderate, 3 = weak/indirect, N/A = not applicable
| Goals & Sub-Goals | 1) Codes of Conduct | 2) Community Micro-Grants | 3) Peer Audit & Recognition |
|---|---|---|---|
| Goal 1: Ensure Safety & Prevent Harm | |||
| Support biocontainment standards & safety-by-design | 1 | 3 | 2 |
| Encourage operational biosafety protocols | 1 | 2 | 1 |
| Reduce accidental exposure or misuse | 1 | 2 | 1 |
| Goal 2: Equity & Access | |||
| Equitable access to research tools | 2 | 1 | 2 |
| Integrate ethics & governance education | 1 | 2 | 1 |
| Broaden participation beyond elite labs | 2 | 1 | 2 |
| Other Considerations | |||
| Minimize costs and burdens | 1 | 1 | 2 |
| Feasibility | 2 | 1 | 2 |
| Not impede research | 1 | 1 | 1 |
| Promote constructive applications | 1 | 1 | 1 |
Drawing on the scoring table, I would prioritize a combined approach centered on Community Driven Codes of Conduct, the Community Lab Micro-Grant Network, and Peer Audit & Recognition. The Codes of Conduct scored highest for fostering safety and preventing harm by establishing shared expectations around biocontainment and biosafety without restricting research. The Micro-Grant Network performed best for equity and access, directly lowering barriers for schools, community labs, and independent creators while including mentorship to support responsible practice. Peer Audit & Recognition complements both by reinforcing operational safety through collaborative review and practical ethics learning, helping translate norms into everyday lab behavior.
This combination reflects key trade-offs and it relies on voluntary incentives rather than formal enforcement, assuming that reputation, community trust, and access to resources could meaningfully shape the community. The approach may struggle if some actors do not value these social levers, and micro-grants could still be captured by already privileged groups. However, for audiences such as MIT leadership, community lab networks, and nonprofit funders, this strategy offers a feasible path that protects safety, expands participation, and promotes constructive uses of bioluminescence while keeping burdens on researchers low and innovation open.
Week 2 Prep Questions
Homework Questions from Professor Jacobson:
Nature’s machinery for copying DNA is called polymerase. What is the error rate of polymerase? How does this compare to the length of the human genome. How does biology deal with that discrepancy?
- The error rate of polymerase is 1:10^6. The length of the human genome is 3.2 Gbp. Biology deals with this discrepancy by incorporating proofreading capabilities within the DNA polymerase. Certain DNA polymerases contain 3’ to 5’ exonuclease activity allowing them to remove any incorrect DNA bases. There are also other mismatch repair systems such as the MutS system that detect mismatched DNA bases and fixes them.
How many different ways are there to code (DNA nucleotide code) for an average human protein? In practice what are some of the reasons that all of these different codes don’t work to code for the protein of interest?
- An average human protein can be encoded by an enormous number of different DNA sequences because most amino acids are specified by multiple codons. Even though DNA sequences could encode the same amino-acid chain, they are not functionally the same because for example, different codons are translated at different speeds due to tRNA abundance.
Homework Questions from Dr. LeProust:
- What’s the most commonly used method for oligo synthesis currently?
- Solid-phase syntheis using the phosphoramadite method.
- Why is it difficult to make oligos longer than 200nt via direct synthesis?
- Each step in the phosphoramadite method has an efficiency of about 99%, which means that even at an oligo length of just 200nt most products will be truncated or contain deletions and therefore not be usable.
- Why can’t you make a 2000bp gene via direct oligo synthesis?
- Once oligos start to reach this length there are issues with the strand starting to fold back on itself as well as the error issue mentioned in the answer to the previous question.
Homework Question from George Church:
What are the 10 essential amino acids in all animals and how does this affect your view of the “Lysine Contingency”?
- Histidine, isoleucine, leucine, lysine, methionine, phenylalanine, threonine, tryptophan, valine, and arganine. Lysine is an essential amino acid which means that it cannot be synthesized by human or other animal cells. Therefore, it has to be obtained from the diet. This means the “Lysine Contingency” that was referenced within the Jurassic Park series makes no sense, as animals already do not synthesize lysine. So this edit would not do anything at all lol.