Week 6 HW: Genetic Circuits Part I
Questions
1. What are some components in the Phusion High-Fidelity PCR Master Mix and what is their purpose?
- Phusion DNA Polymerase: Catalyzes the synthesis of new DNA strands. Has 3′→5′ exonuclease proofreading activity, which removes incorrectly added nucleotides. Phusion polymerase is a genetically engineered DNA polymerase fused to a DNA-binding domain. The fusion domain increases DNA binding, which improves processivity.
- Reaction buffer: Help to maintain a stable pH. Also provides optimal ionic strength for polymerase activity. Stabilizes enzyme structure at high temperatures.
- Magnesium Chloride (MgCl₂): Essential cofactor for DNA polymerases. Coordinates with the phosphate groups of incoming nucleotides. Helps stabilize primer–template interactions.
- dNTPs: Provide the substrates used to synthesize new DNA strands. Each nucleotide carries three phosphates, providing the energy needed for polymerization.
2. What are some factors that determine primer annealing temperature during PCR?
- Annealing temperature is primarily determined by the melting temperature (Tm) of the primers. Tm is influenced by primer length and GC content, as well as, sequence composition and distribution of bases. Salt concentration in the reaction and secondary structures (hairpins) can also impact the annealing temperature.
3. There are two methods from this class that create linear fragments of DNA: PCR, and restriction enzyme digests. Compare and contrast these two methods, both in terms of protocol as well as when one may be preferable to use over the other.
- PCR:
- Amplifies DNA. Uses a thermostable polymerase, primers, and thermal cycling to amplify a specific DNA sequence exponentially.
- Specificity: Determined by primer design and annealing temperature.
- Error Introduction: Can introduce polymerase errors (even with high-fidelity enzymes).
- Flexibility: Can amplify virtually any region with good primer design.
- Uses: Amplifying a specific gene or region from a genome, generate large amounts of DNA from tiny starting quantities, genotyping, mutagenesis, add tags, restriction sites, or homologous sequences for cloning, and diagnostic applications
- Restriction Enzyme Digests:
- Cuts DNA at specific recognition sequences. Uses sequence specific endonucleases to cut DNA at defined recognition sites.
- Specificity: Limited by recognition sites. The enzyme will cut everywhere its recognition site appears in the DNA.
- Error Introduction: Doesn’t synthesize new DNA, so no new errors are introduced.
- Flexibility: Constrained by where recognition sites occur.
- Uses: Cut DNA at known recognition sites, prepare compatible sticky or blunt ends for cloning, verifiy plasmid constructs by restriction mapping, linearize or fragment dna in predictable ways, and library construction.
Protocol Comparison
| Step | PCR | Restriction Enzyme Digest |
|---|---|---|
| Input DNA | Can work from nanogram or even picogram quantities | ng–µg range |
| Reagents | Polymerase, dNTPs, primers, buffer, MgCl₂ | Restriction enzyme(s), appropriate buffer |
| Thermal requirements | Thermocycler required (denature → anneal → extend, repeated) | Simple isothermal incubation, for 30-60 minutes |
| Time | 1–3 hours typically | 30-60 minutes |
| Visualization | Gel electrophoresis | Gel electrophoresis |
4. How can you ensure that the DNA sequences that you have digested and PCR-ed will be appropriate for Gibson cloning?
- The DNA pieces that are generated need to be designed so that adjacent fragments share matching end sequences, usually about 20-40 bp of overlap. During PCR, primers are designed that add the overlaps. Make sure the overlaps only match the intended neighboring fragment and not some other region in the assembly. The vector needs to be linearized so its ends match the insert overlaps. PCR and digested DNA should be purified prior to use. As long as overlaps are correct, Gibson Assembly can chew back the ends, let complementary regions anneal, fill in gaps, and seal the nicks.
5. How does the plasmid DNA enter the E. coli cells during transformation?
- Chemical Transformation (Heat Shock):
- Cells are first treated with cold CaCl₂. DNA and the bacterial membrane are both negatively charged because of their phosphate groups. That means they normally repel each other electrostatically. Calcium ions shield those negative charges and allow plasmid DNA to approach and stick to the outer membrane surface. Then comes the heat shock step, you briefly shift them to 42°C for 30–90 seconds, then return them to ice. This rapid temperature change is thought to drive uptake of DNA through temporary pores or disruptions in the membrane.
- Electroporation:
- Typically considered more efficient than chemical transformation. Cells are placed in a cuvette with DNA and subjected to a brief, intense electrical pulse. This electric field temporarily creates nanoscale pores in the membrane. DNA present in the solution is driven through these pores by the electric field itself. After the pulse ends, the membrane reseals and the DNA remains inside.
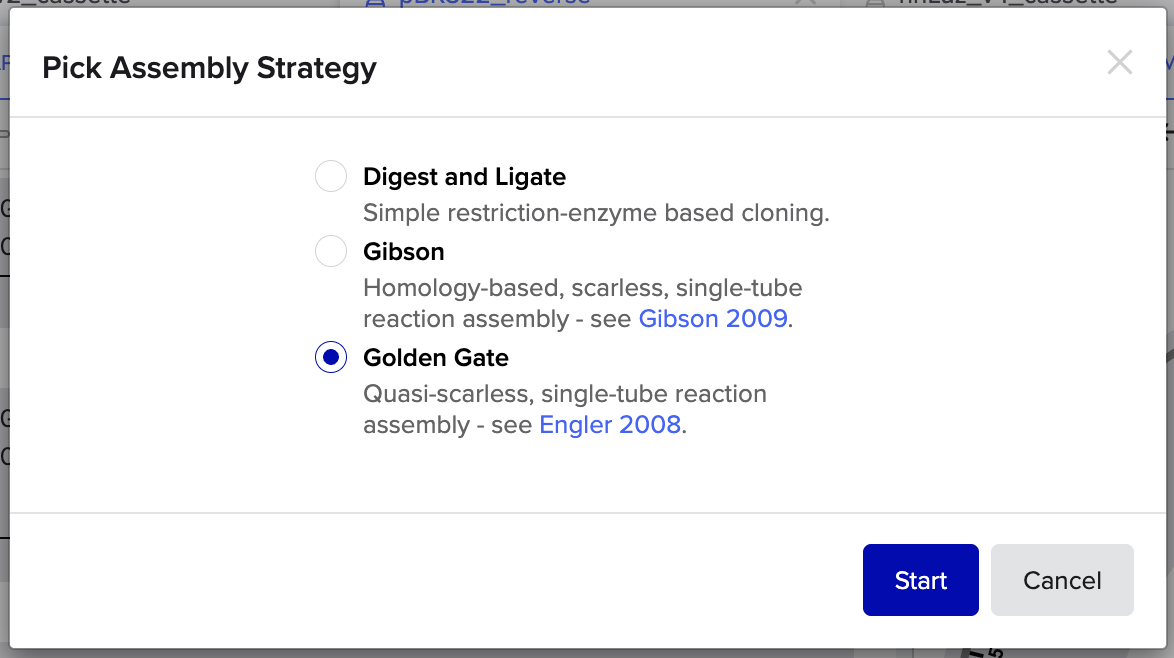
6. Describe another assembly method in detail (such as Golden Gate Assembly)
- 1. Explain the other method in 5 - 7 sentences plus diagrams (either handmade or online).
- Golden Gate Assembly is a DNA assembly method that uses Type IIS restriction enzymes to cut DNA outside of their recognition sites, this allows for the design of custom overhangs that determine how fragments join together. Enzymes such as BsaI recognize a specific sequence but cleave a few bases away from it, producing programmable sticky ends. Because the overhang sequences can be chosen, multiple DNA fragments can be assembled in a precise order within a single reaction. The reaction mixture contains the restriction enzyme and a DNA ligase, and the protocol cycles between temperatures that allow cutting and ligation to occur repeatedly. When fragments join correctly, the recognition sites are removed from the final construct, preventing the enzyme from cutting the assembled product again. This design allows many fragments (often 5–10 or more) to be assembled simultaneously in a one pot reaction. Golden Gate Assembly is especially useful for modular cloning systems where standardized parts such as promoters, coding sequences, and terminators need to be assembled rapidly.
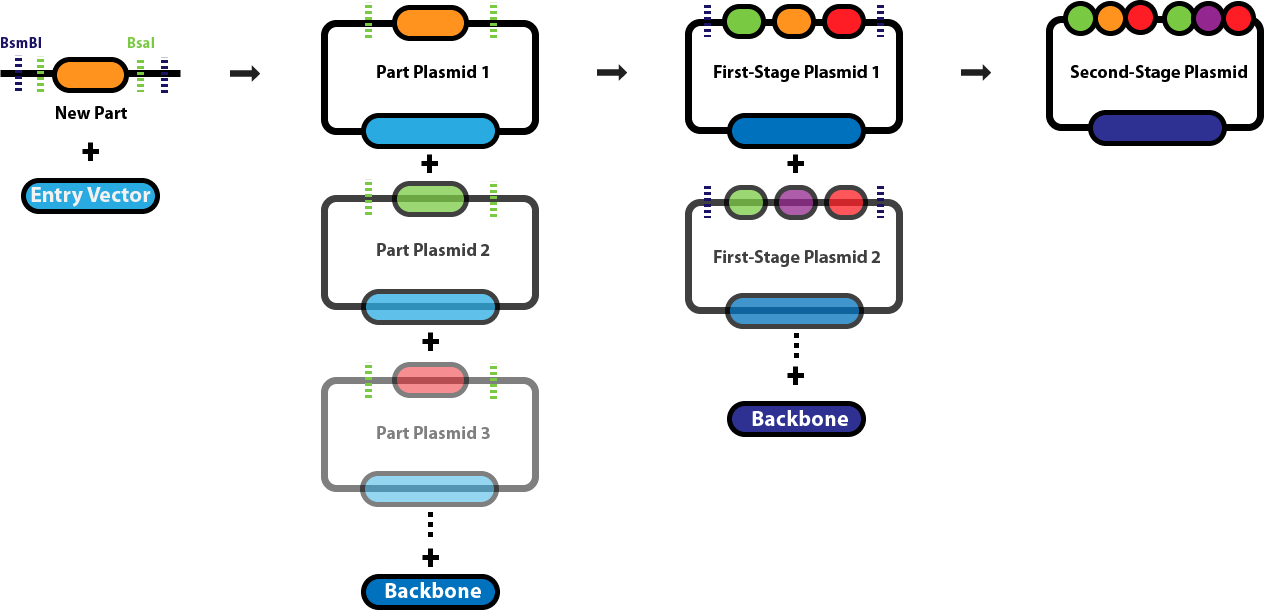

- Diagram explanation:
- A new DNA part is first inserted into an entry vector using Type IIS restriction enzymes such as BsaI, which cuts outside its recognition site to create specific overhangs.
- This cloning step generates individual part plasmids, each containing a single genetic component (such as a promoter, coding sequence, or terminator) flanked by designed overhangs.
- Multiple part plasmids are then mixed in a single reaction with BsaI and DNA ligase, allowing the enzyme to cut and generate complementary sticky ends.
- The matching overhangs guide the DNA fragments to ligate together in a specific predetermined order, forming larger constructs called first-stage plasmids.
- Because the restriction sites are removed during assembly, correctly assembled fragments are no longer cut by the enzyme, increasing assembly efficiency.
- Finally, several first-stage plasmids can be combined with a backbone plasmid to form a second-stage plasmid containing many genetic parts arranged sequentially.
- Golden Gate Assembly is a DNA assembly method that uses Type IIS restriction enzymes to cut DNA outside of their recognition sites, this allows for the design of custom overhangs that determine how fragments join together. Enzymes such as BsaI recognize a specific sequence but cleave a few bases away from it, producing programmable sticky ends. Because the overhang sequences can be chosen, multiple DNA fragments can be assembled in a precise order within a single reaction. The reaction mixture contains the restriction enzyme and a DNA ligase, and the protocol cycles between temperatures that allow cutting and ligation to occur repeatedly. When fragments join correctly, the recognition sites are removed from the final construct, preventing the enzyme from cutting the assembled product again. This design allows many fragments (often 5–10 or more) to be assembled simultaneously in a one pot reaction. Golden Gate Assembly is especially useful for modular cloning systems where standardized parts such as promoters, coding sequences, and terminators need to be assembled rapidly.
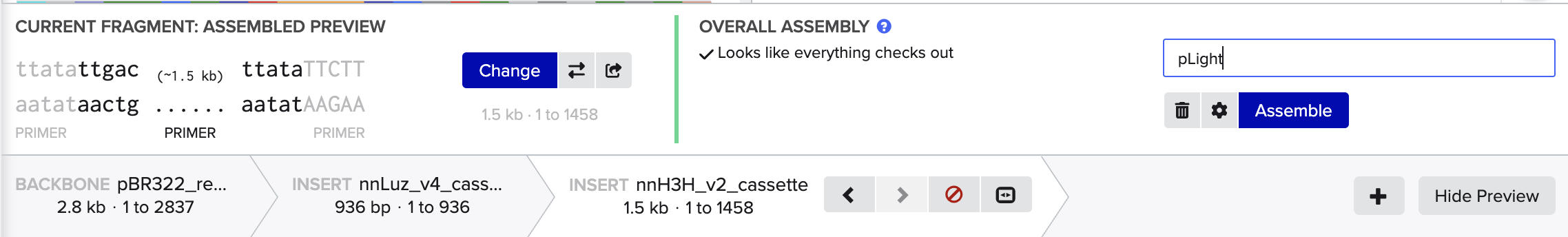
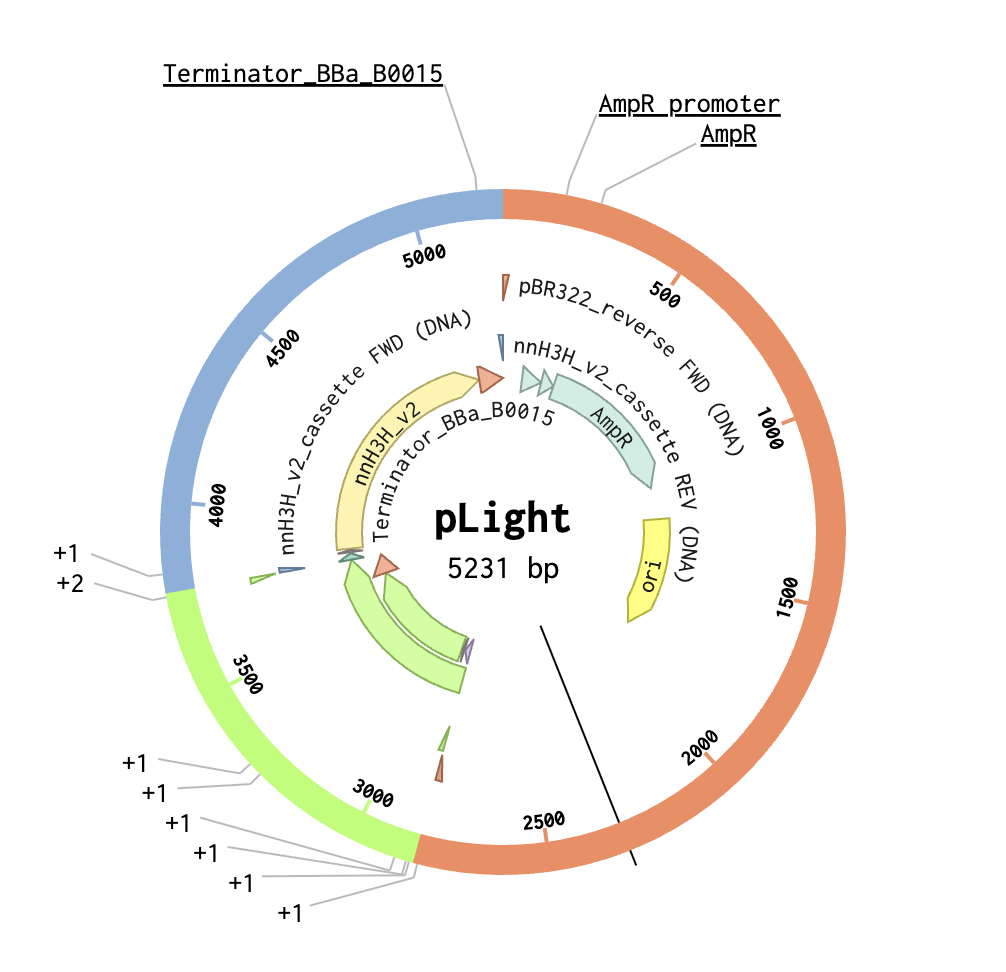
- 2. Model this assembly method with Benchling or Asimov Kernel!