Week 11 HW: Bioproduction & Cloud Labs
Part A: The 1,536 Pixel Artwork Canvas | Collective Artwork
I contributed a single pixel to the bioart project. It was one of the early additions—a red pixel placed in the bottom-left quadrant, about three rows down from the top of that section. At the time, the canvas was still mostly empty, and my contribution was eventually replaced as the artwork evolved into the final design, which included the word “Love.”
Even though my individual contribution was small, I think the project itself was a strong idea. It created a shared creative space where students across the HTGAA community could participate in building something collectively. This kind of setup is especially valuable since not everyone is physically located at MIT, Harvard, or a local node. It gave everyone a chance to contribute to a fun, interactive experiment while still being part of a larger collaborative effort.
One way this collaborative art experiment could be improved for next year is by adding a bit more structure or coordination without removing the creative freedom. For example, allowing students to preview or plan designs before placing pixels could reduce constant overwriting and lead to more intentional collaboration.
Part B: Cell-Free Protein Synthesis | Cell-Free Reagents
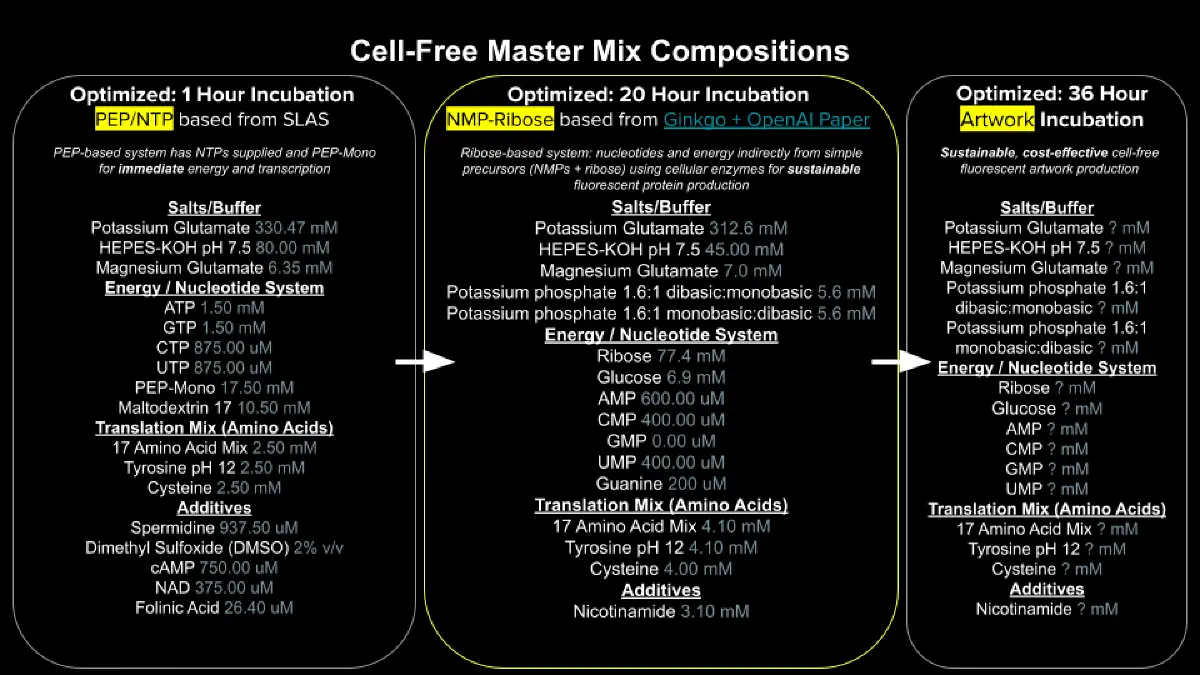

Cell-Free Protein Synthesis Reaction Components
E. coli Lysate
- BL21 (DE3) Star Lysate (includes T7 RNA Polymerase) - Provides the ribosomes, tRNAs, aminoacyl-tRNA synthetases, and metabolic enzymes needed for transcription and translation. The DE3 strain supplies T7 RNA polymerase for high-yield transcription from T7 promoters, and the “Star” (RNase E mutant) background slows mRNA degradation to extend protein output.
Salts/Buffer
- Potassium Glutamate - Maintains ionic strength and mimics intracellular conditions to stabilize proteins and ribosomes.
- HEPES-KOH pH 7.5 - Buffers the reaction to maintain a stable pH optimal for enzymatic activity.
- Magnesium Glutamate - Essential cofactor for ribosomes and enzymes; critical for translation and nucleotide interactions.
- Potassium Phosphate Monobasic / Dibasic - Provides additional buffering capacity and helps maintain phosphate balance for metabolic reactions.
Energy/Nucleotide System
- Ribose - Serves as a precursor for nucleotide synthesis in NMP-based energy systems.
- Glucose - Fuels metabolic pathways in the lysate to regenerate energy (ATP).
- AMP, CMP, UMP - Nucleoside monophosphates that the lysate’s endogenous nucleotide kinases phosphorylate up to NDPs and NTPs, supplying the ATP/CTP/UTP needed for transcription and translation.
- GMP - Listed at 0 µM in the middle master mix composition. Would normally be the GTP precursor, but this formulation supplies guanine instead.
- Guanine - Nucleobase used to support GMP/GTP synthesis for transcription and translation.
Translation Mix (Amino Acids)
- 17 Amino Acid Mix - The bulk substrate for protein synthesis; supplies all amino acids except the three broken out separately for solubility/stability reasons.
- Tyrosine - Tyrosine is poorly soluble at neutral pH, so it’s added from a high-pH stock to keep it dissolved before it’s diluted into the reaction.
- Cysteine - Added separately because it is prone to oxidation and needs controlled availability.
Additives
- Nicotinamide - Supports redox balance by maintaining NAD+/NADH pools, improving metabolic efficiency and protein yield.
Backfill
- Nuclease Free Water - Brings the reaction to the correct final volume without introducing nucleases that could degrade RNA or DNA.
The one hour PEP/NTP mix supplies energy and nucleotides directly as NTPs (ATP, GTP, CTP, UTP) with PEP-Mono and maltodextrin as the energy source, giving fast but short lived output because PEP is consumed quickly and phosphate buildup inhibits the reaction. The 20 hour NMP and Ribose mix instead feeds in cheap upstream precursors, ribose and glucose for energy regeneration, NMPs (AMP, CMP, UMP) and guanine that the lysate’s own kinases phosphorylate up to NTPs. So the system continuously regenerates its own ATP and NTPs rather than burning through a finite pool. The NMP and Ribose formulation also drops pricey additives like cAMP, NAD, folinic acid, spermidine, and DMSO in favor of just nicotinamide, making it dramatically cheaper and more sustainable at the cost of slower kinetics.
Bonus Question: Transcription can still occur because the cell-free lysate contains enzymes from the nucleotide salvage pathway that convert free guanine into GMP, and then further phosphorylate it into GDP and GTP. The produced GTP is the actual substrate used by RNA polymerase during transcription, so supplying guanine provides a precursor that the system can metabolically convert as needed.
Part C: Planning the Global Experiment | Cell-Free Master Mix Design
Fluorescent Proteins
- sfGFP - chromophore maturation requires molecular oxygen for the final oxidation step (as with all avGFP-derived FPs), so in a sealed 36-hour cell-free reaction oxygen gets depleted and newly translated sfGFP accumulates as non-fluorescent apo-protein. Even with its fast folding, the readout plateaus once dissolved O₂ runs out.
- mRFP1 - a somewhat slowly maturing monomer whose DsRed family chromophore requires two oxidation steps (not one), making it both oxygen hungry and slow; it’s also dimmer and less photostable than its parent DsRed. In cell-free reactions the slow, oxygen-dependent red chromophore formation means a lot of translated protein sits as the non-fluorescent green intermediate instead of reaching the mature red state.
- mKO2 - strongly oxygen-dependent during chromophore maturation, with a pO₂·50 of 0.9%, meaning fluorescence drops by half at oxygen tensions that are easily reached inside a closed reaction tube. This makes it the most hypoxia sensitive FP on the list and a major liability in a long, sealed cell-free incubation where O₂ gets consumed by the lysate’s residual respiratory enzymes.
- mTurquoise2 - The main cell-free weakness for mTurquoise2 is that its chromophore is built from a Tyr-Gly backbone cyclization plus oxidation, and like other avGFP derivatives that oxidation step consumes O₂ and produces H₂O₂ as a byproduct; over 36 hours in a closed reaction, both oxygen depletion and H₂O₂ accumulation can cap the mature fluorophore pool. Its excitation also sits in the near-UV/violet range where lysate components (flavins, NADH) contribute background that hurts signal-to-noise.
- mScarlet_I - The T74I mutation gives mScarlet-I markedly faster maturation but at the cost of a reduced quantum yield (0.54) and shorter fluorescence lifetime (3.1 ns), and like all DsRed lineage RFPs its chromophore requires two sequential oxidation steps to reach the red state. Incomplete maturation (“dead-end” green intermediates) and oxygen limitation over a 36-hour reaction both suppress the final red signal.
- Electra2 - a blue FP derived from mRuby3 (Entacmaea quadricolor lineage); its blue emission at ~456 nm overlaps heavily with the autofluorescence of lysate cofactors like NADH (~460 nm) and flavins, which tanks signal-to-noise in cell-free systems. It also inherits the eqFP611/DsRed family two step oxidative maturation, so oxygen depletion over 36 hours hurts yield just as it does for the RFPs.
Hypothesis
Hypothesis: Boost cysteine to accelerate Electra2 chromophore maturation and final fluorescence yield over 36 hours, as well as, boost guanine to relieve GTP bottleneck.
Protein: Electra2
Reagent adjustment: Cysteine 4.0 mM → 8.0 mM & Guanine 0.156 mM → 0.4 mM
Rationale and expected effect: Electra2 is derived from mRuby3, which descends from eqFP611, a DsRed/Anthozoa lineage chromophore. These chromophores form from an X-Tyr-Gly tripeptide and require oxidative cyclization to mature. Critically, the parent mRuby3 scaffold contains internal cysteines that participate in chromophore environment and folding, and free cysteine in solution is needed both as a building block (Electra2’s sequence still contains Cys residues) and as a reducing agent that prevents misoxidation of those internal thiols during the long folding/maturation window. In a 36-hour reaction, the free cysteine pool gets depleted and oxidized (cystine, mixed disulfides), which can stall folding of cysteine containing FPs and leave a larger fraction stuck as immature/misfolded protein. Doubling the cysteine pool extends the reducing capacity and amino acid availability across the full 36-hour window, which should increase the fraction of Electra2 that reaches the mature, blue fluorescent state. The expected effect is higher final Electra2 fluorescence intensity at 456 nm at the 36-hour endpoint, with the largest gains in the second half of the incubation (12–36 hr) when the baseline cysteine pool would otherwise be exhausted.
Secondly, for the boost guanine to relieve GTP bottleneck, translation elongation consumes 2 GTP per amino acid added, and Electra2 is a 236-residue protein, that’s ~470 GTP per molecule made. In a 36-hour reaction, GTP regeneration via the salvage pathway (guanine → GMP → GDP → GTP) becomes rate-limiting. The expected effect here is more guanine means more sustained GTP, means more Electra2 translated. Particularly relevant since GMP is set to 0 in this mix, so guanine is the entire GTP feedstock.