Homework
Please credit F.R. and cite the source if you use any content published on this website. If you find my work interesting, feel free to reach out to discuss collaborations or Art & Science consulting opportunities.
Please credit F.R. and cite the source if you use any content published on this website. If you find my work interesting, feel free to reach out to discuss collaborations or Art & Science consulting opportunities.


(c) Flo Razoux, AI generated
I. PROJECT DESCRIPTION
HOW TO GROW BIOLUMINESCENT MENSTRUAL BLOOD? A SYNTHETIC BIOART PROJECT
Bioluminescence is the production of visible light by living organisms. The light is produced through the oxidation of luciferin, which is catalyzed by an enzyme called luciferase. This phenomenon is believed to have evolved 540 million years ago in Earth’s ancient oceans [1]. Present in the majority of marine species but also in land-based ones such as fungi, bacteria and fireflies, bioluminescence serves biological purposes such as mating, hunting, and defense behaviors [2]. Bioluminescence has become an essential tool in biological engineering not just for sensing but also controlling biological processes [3-4].
While bioluminescence unanimously elicits attraction and curiosity, one cannot say the same for menstruation. The social stigma around it has slowly receded with increased visibility in the media over the past five years, but menstrual health remains under-researched. In particular, the precise biological impact of hormonal variation during perimenopause, which can last up to 10 years, remains unknown. When it comes to transgender menstrual health, the gap is even wider. Insights into how hormone therapy affects the menstrual cycle in trans men are sparse and usually extrapolated from research carried out on menopausal cis women [5–6].
Ironically, synthetic biology holds extraordinary potential to revolutionize trans health. For instance, by unlocking the expression of genes implicated in genital growth or creating organs de novo, synthetic biology methods could dramatically improve the quality of life of transgender patients undergoing gender-affirming surgeries, which are currently highly risky and often associated with poor outcomes.
The art installation invites the public to immerse themselves in a softly glowing, living artificial womb and observe pulsating “menstrual” blood being infused into it. By using synthetic biology to transform menstruation from a hidden process into a shared contemplative experience, I aim to raise awareness of the societal impact of scientific bias and the urgent need to invest in neglected research fields such as menstrual and trans health.
Shining light into the abyss of a womb is also a metaphorical invitation to regain the senses. Beyond the previously mentioned primary goal, I want to show that synthetic biology can be used in ways other than a product-centered perspective. In a world that is suffocating, it is meaningful to be reminded of life’s evolutionary timescales, as well as how the race for productivity and overconsumption affects Earth’s wonders such as embryonic development and bioluminescent life. The piece is a call to slow down, and rethink our vision of what the future of the synthetic biology revolution should look like.
Methodological strategies The menstruation-like fluid can either be derived from menses or created artificially [7]. Two strategies can be considered to enable the production of bioluminescence:
Bibliography [1] Danielle M. DeLeo et al. Evolution of bioluminescence in Anthozoa with emphasis on Octocorallia. Proc Biol Sci (2024) [2] Martini S. et al. Quantification of bioluminescence from the surface to the deep sea demonstrates its predominance as an ecological trait. Nature Scientific Reports (2017) [3] Widder E. et al. Review of Bioluminescence for Engineers and Scientists in Biophotonics. IEEE Journal of Selected Topics in Quantum Electronics (2013) [4] Love A. et al. Seeing (and using) the light: Recent developments in bioluminescence technology. Cell Chem Biol. (2020) [5] Perrone A. Effect of long-term testosterone administration on the endometrium of female-to-male (FtM) transsexuals. J Sex Med (2009) [6] Buck E. et al. Menstrual Suppression. Treasure Island (2025) [7] Tindal K. et al. The composition of menstrual fluid, its applications, and recent advances to understand the endometrial environment: a narrative review. F&S Reviews (2024) [8] France M. et al. Towards a deeper understanding of the vaginal microbiota. Nat Microbiol (2022)
II. GOUVERNANCE & POLICY GOALS (Synthetic Biology & Bioart in Berlin, Germany)
Overarching goal Ensure that the use of synthetic biology in artistic contexts is safe, non-maleficent, socially responsible, and inclusive, while complying with German and EU biosafety, biosecurity, and human rights frameworks.
GOAL 1: Ensure bio safety and prevent harm Sub-goal 1.1: Regulatory compliance and containment. Sub-goal 1.2: Prevention of misuse of the art work. Key institution: Institutional biosafety committees (Beauftragte für biologische Sicherheit)
GOAL 2: Promote equity and justice in biomedical narratives Sub-goal 2.1: Address epistemic bias in research priorities. Sub-goal 2.2: Protect the dignity and the autonomy of transgender patients. Key institution: German Ethics Council (Deutscher Ethikrat).
GOAL 3: Foster responsible innovation and public engagement Sub-goal 3.1: Transparency and public understanding of the art piece. Clearly communicate what aspects of the work are biological, synthetic, or metaphorical, supporting informed public engagement with synthetic biology. Sub-goal 3.2: Encourage reflective, non-product-centered innovation. Use the artwork to challenge efficiency- and market-driven narratives of biotechnology, aligning with broader German and EU discussions on sustainability and responsible research and innovation (RRI). Key institutions: European Commission (RRI framework), German Federal Ministry of Education and Research (BMBF).
III. GOUVERNANCE ACTIONS
Action 1: Mandatory Biosafety & Ethics Review for Art–Science Projects
Actor(s): Academic institutions, art schools, biosafety committees, federal regulators (BVL)
Purpose Current state: Biosafety review in Germany (GenTG) is robust for academic research, but art–science projects often fall into grey zones, especially when hosted outside of traditional labs. Proposed change: Require formal biosafety and ethics review for any art project involving synthetic biology or GMOs, regardless of whether it is framed as “research” or “art.”
Design Extend existing institutional biosafety committee (Beauftragte für biologische Sicherheit) oversight to art institutions collaborating with labs. Require project registration and approval before exhibition, similar to IRB-style review but adapted for bioart. Low administrative burden by using existing regulatory infrastructure under the Gentechnikgesetz.
Assumptions Assumes that ethical risks in bioart are comparable to those in research. Assumes institutions are willing to take responsibility for hybrid practices.
Risks of Failure & “Success” Failure: Overregulation could discourage experimental art or push practices underground. Success risk: If normalized, review processes may become procedural and lose critical engagement, reducing ethics to box-ticking. Analogy: Drone registration systems that increased safety but initially slowed creative experimentation.
Action 2: Incentivizing Low-Risk, Contained Design Choices
Actor(s): Funding bodies (BMBF), foundations, academic labs, artists
Purpose Current state: Synthetic biology innovation is often optimized for scalability, performance, and commercial value. Proposed change: Create incentives (funding criteria, exhibition access) favoring contained, non-scalable, low-risk biological designs, especially in public-facing projects.
Design Funding calls explicitly reward projects that use Risk Group 1 organisms, non-reproductive systems, or synthetic analogues. Curatorial guidelines for public exhibitions prioritize containment and reversibility.
Assumptions Assumes artists and researchers respond meaningfully to incentive structures. Assumes “low-risk by design” can be assessed reliably.
Risks of Failure & “Success” Failure: Incentives may be ignored if prestige or novelty outweighs funding concerns. Success risk: Could unintentionally marginalize more radical or speculative research that challenges current risk models. Analogy: “Privacy-by-design” incentives in software development that improved norms but constrained some innovation paths.
Action 3: Transparency & Contextualization Requirements for Public Display
Actor(s): Exhibiting institutions, artists, regulators, public educators
Purpose Current state: Audiences often cannot distinguish between speculative, artistic, and clinical uses of biotechnology. Proposed change: Require clear public-facing contextualization for bioart using synthetic biology.
Design Mandatory disclosure explaining what is biological, synthetic, symbolic, or hypothetical. Clear statements that the work is non-therapeutic and non-clinical. Oversight by exhibiting institutions, not law enforcement.
Assumptions Assumes transparency increases public trust rather than fear. Assumes audiences engage with contextual information when provided.
Risks of Failure & “Success” Failure: Contextualization may be ignored or misunderstood. Success risk: Overexplanation could domesticate or neutralize critical artistic ambiguity. Analogy: Financial product disclosures that protect consumers but often overwhelm them.
IV. GOVERNANCE ACTIONS: SCORING
(from 1-3 with, 1 as the best, or n/a)
| Does the option: | Action 1 | Action 2 | Action 3 |
|---|---|---|---|
| Enhance Biosecurity | |||
| • By preventing incidents | 1 | 1 | 2 |
| • By helping respond | 1 | 1 | 2 |
| Foster Lab Safety | |||
| • By preventing incident | 1 | 1 | n/a |
| • By helping respond | 1 | 1 | n/a |
| Protect the environment | |||
| • By preventing incidents | 1 | n/a | 2 |
| • By helping respond | 1 | n/a | 2 |
| Other considerations | |||
| • Minimizing costs and burdens to stakeholders | 3 | 1 | 3 |
| • Feasibility? | 2 | 1 | 1 |
| • Not impede research | 2 | 2 | 1 |
| • Promote constructive applications | 2 | n/a | 1 |
V. PRIORITAZING STRATEGY FOR ACTION(S)
Action 01 should be prioritized because a foundational principle of academic biological research is the precautionary principle. Action 03 should also be prioritized because bioart can only be meaningful if it is conducted ethically and responsibly—not to create sensation, but to stimulate curiosity and deeper reflection.
VI. CONCLUSION: ETHICAL CONCERNS
Coming from an artistic perspective, I found it challenging to situate my project within the framework of the course. I was troubled by the fact that my proposal was not product-oriented: transforming the appearance of menstrual blood into light did not align with the “How To Grow” formulation.
As I am only beginning to engage with synthetic biology, it may seem presumptuous to question product-driven research. Yet, like many other fields, synthetic biology is shaped by the economic logics that have governed technological development since the Industrial Revolution. This raises the possibility of expanding its scope beyond productivity alone, toward applications that invite reflection, care, and alternative ways of relating to life.
AI support: ChatGPT. The tool was used to discuss the relevance of different final project ideas and to provide initial responses that served as a starting point for questions related to governance and policy, based on the prompts: project description and assignment questions.
HOMEWORK QUESTIONS FROM STEVEN JACOBSEN
After proofreading, DNA polymerase has an error rate of 1:106, meaning 1 error per 1 million base pairs. The human genome contains approx. 3 billions base pairs (3x109bp) in haploid cells and thus, 6 billions base pairs (6x109bp) in diploid cells. This means that thousands of errors occur during DNA replication, but the cell machinery has a post-replication mismatch repair (MMR) system that brings down DNA replication errors to only a few potential base pairs per division.
Human proteins are made of 20 amino acids (aa) whose code is stored in the DNA (A,C,G,T nucleotides coding). Ribosomes are macromolecules that synthesize proteins by translating messenger RNA (mRNA) into amino acid chains. This translation process is mediated by transfer RNA (tRNA) molecules that add a single amino acid corresponding to the mRNA code (A,C,G,U three-nucleotide codon/anticodon coding system). Because there are fewer amino acids than codon possibilities (4^3=64), multiple codons can encode for the same amino acid: a phenomenon called codon redundancy. Some codons are also associated to prompt the start and the end of the translation process. According to the genetic code, there are between two and four DNA code possibilities per amino acid. So in theory there are staggering possibilities to code for an average human protein (approx. 450-480 aa length).
But in practice, spatial configuration and kinetics can affect this process:
AI support: ChatGPT. Prompt: Please read this research article thoroughly and answer “In practice what are some of the reasons that all of these different codes don’t work to code for the protein of interest?”: https://www.science.org/doi/10.1126/science.1241459
HOMEWORK QUESTIONS FROM EMILY LEPROUST
Solid-phase phosphoramidite chemical synthesis is the industry-standard, automated method for creating custom DNA/RNA oligonucleotides.
Direct synthesis of oligonucleotides (oligos) longer than 200 nucleotides (nt) is difficult primarily because of the cumulative, exponential decline in yield due to imperfect coupling efficiency and the accumulation of chemical errors. Cumulative Inefficiency: Standard oligo synthesis adds nucleotides one by one. Even if each step has a 99% success rate, the overall yield drops significantly as length increases. Longer sequences result in mostly truncated, incorrect, or incomplete products. Accumulation of Errors: With longer synthesis times, chemical side reactions increase, leading to a higher rate of sequence errors, such as deletions or misincorporations. Purification Challenges: As the length increases, it becomes difficult to separate the desired full-length, error-free product from the failed side products. Steric Hindrance: As the oligo grows, it can become tangled, making it harder for reagents to access the reactive end.
Making a 2000bp (base pair) gene via direct synthesis is currently not possible due to these limitations in efficiency, which result in a very low yield of the full-length, correct sequence. Exponentially Low Yield: Using standard 99% efficiency, a single-stranded DNA or RNA molecule that is 2000 bases in length would yield roughly effectively zero usable product. Error Rate vs. Length: The error rate is roughly one mistake per 200 bases, meaning a 2000bp strand would contain an average of 10 errors, making it highly unlikely to contain the correct sequence. Physical Limits of Support: The solid support material (e.g., controlled pore glass) becomes clogged by the growing DNA strands, preventing reagents from completing the synthesis.
AI support: ChatGPT, Gemini. Prompt: long oligonucleotide synthesis + “What’s the most commonly used method for oligo synthesis currently? Why is it difficult to make oligos longer than 200nt via direct synthesis? Why can’t you make a 2000bp gene via direct oligo synthesis?”
HOMEWORK QUESTIONS FROM GEORGE CHURCH
Lysine is one of the 10 essential amino acids found in all animals: Arginine, Histidine, Isoleucine, Leucine, Lysine, Methionine, Phenylalanine, Threonine, Tryptophan, and Valin. An amino acid is classified as essential in a species if the organism can’t produce it and therefore is required in the diet (or any other external supply) in order to survive.
In the movie Jurassic Park (1993), scientist Ray Arnold explains how the research team modified the genome of the dinosaurs to prevent them from surviving in the wild in case the dinosaurs would escape the park: “The lysine contingency is intended to prevent the spread of the animals in case they ever get off the island. Dr. Wu inserted a gene that makes a single faulty enzyme in protein metabolism. The animals can’t manufacture the amino acid lysine. Unless they’re continually supplied with lysine by us, they’ll slip into a coma and die.”
Lysine is classified as an essential amino acid in all known animals, including vertebrates. The movie portrayed the “Lysine Contingency” as an engineered weakness but it is likely that dinosaurs likely didn’t have an endogenous lysine biosynthesis pathway to remove in the first place. The auxotrophic strategy presented in the “Lysine Contingency” concept is also not valid. Indeed, lysine is widely present in nature, particularly animals but also in some plants. Carnivorous dinosaurs representing the main threat on the island, they would likely have no difficulty in finding their lysine supply in the wild. The idea of making an organism dependent on a non-natural amino acid would have been more plausible than preventing biosynthesis of a normal nutrient like lysine.
In the real world, synthetic biologists use more robust strategies to design genetic safeguards: Genetic kill switches: circuits that trigger death in certain environments. Synthetic amino acid dependencies: organisms engineered to depend on non-natural amino acids that aren’t in nature. Multiple overlapping dependencies: not just one but many safety constraints. Genetic firewalling: preventing horizontal gene transfer.
In conclusion, movies like Jurassic Park make synthetic biology look inherently dangerous, even though real scientists focus heavily on safety and careful regulation. The media shape how society feels about science, thus also have the responsibility to spark curiosity without creating unnecessary stigma around technologies that can also bring major benefits.
AI support: Gemini. Prompt: What could the scientist of the “Lysine Contingency” have proposed instead?


(c) Flo Razoux, AI generated
Part 1: Benchling & In-silico Gel Art
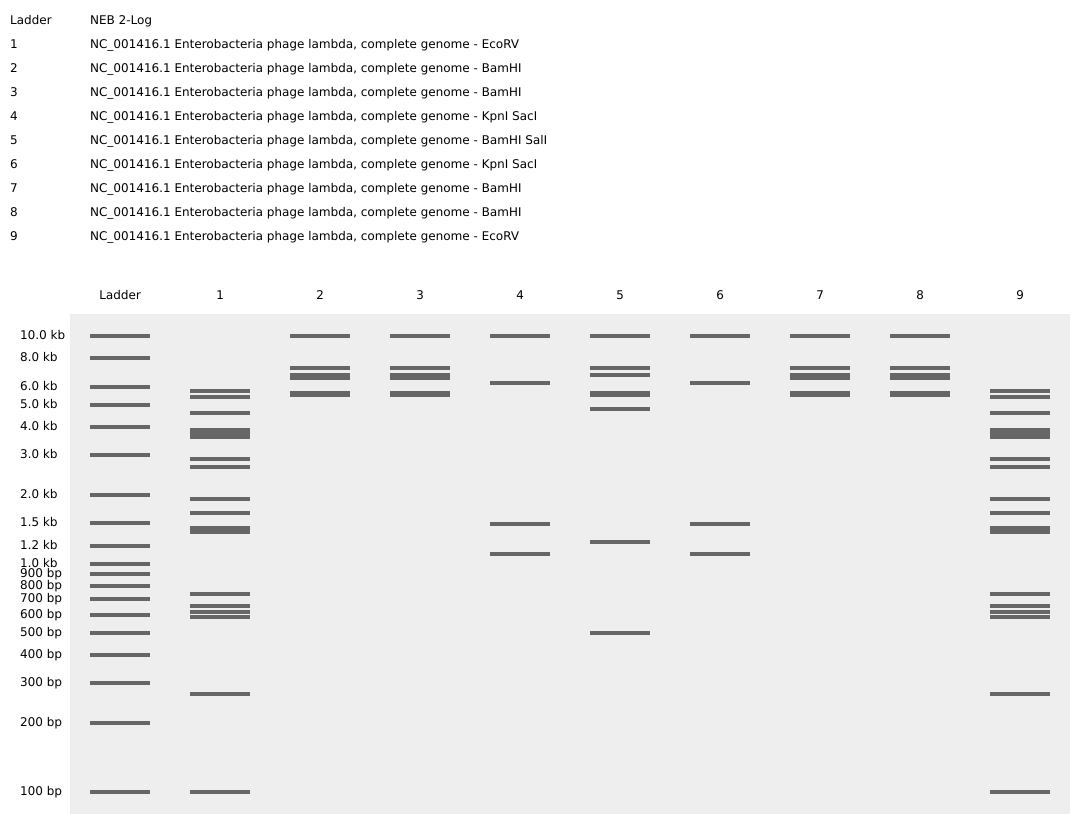

Part 2: Gel Art - Restriction Digests and Gel Electrophoresis
No access to lab
Part 3: DNA Design Challenge
3.1. LUCIFERASE
Bioluminescent art is based on the organic production of visible light by living organisms. This light is produced through the oxidation of luciferin, which is catalyzed by an enzyme called luciferase. For this week’s homework, we will focus on the protein coding for this enzyme, which was first identified in the firefly species (Photinus Pyrlis) [1].
[1] De Wet J.R. et al. Firefly luciferase gene: structure and expression in mammalian cells. Mol Cell Biol (1987). https://pmc.ncbi.nlm.nih.gov/articles/PMC365129/
Sources
NCBI database search: https://www.ncbi.nlm.nih.gov/protein/BAF48396.1
Uniprot database search: https://www.uniprot.org/uniprotkb/P08659/entry#sequences
Protein Sequence
550 amino acids
1 medaknikkg papfypledg tageqlhkam kryalvpgti aftdahievn ityaeyfems
61 vrlaeamkry glntnhrivv csenslqffm pvlgalfigv avapandiyn erellnsmni
121 sqptvvfvsk kglqkilnvq kklpiiqkii imdsktdyqg fqsmytfvts hlppgfneyd
181 fvpesfdrdk tialimnssg stglpkgval phrtacvrfs hardpifgnq iipdtailsv
241 vpfhhgfgmf ttlgylicgf rvvlmyrfee elflrslqdy kiqsallvpt lfsffakstl
301 idkydlsnlh eiasggapls kevgeavakr fhlpgirqgy gltettsail itpegddkpg
361 avgkvvpffe akvvdldtgk tlgvnqrgel cvrgpmimsg yvnnpeatna lidkdgwlhs
421 gdlaywdede hffivgrlks likykgyqva paelesillq hpnifdagva glpdddagel
481 paavvvlehg ktmtekeivd yvasqvttak klrggvvfvd evpkgltgkr darkireili
541 kakkggkskl
3.2. Reverse Translate: Protein sequence to DNA sequence
Source
NCBI database search for P.pyralis (firefly) luciferase gene: https://www.ncbi.nlm.nih.gov/nuccore/M15077
Luciferase protein DNA sequence
1 ctgcagaaat aactaggtac taagcccgtt tgtgaaaagt ggccaaaccc ataaatttgg
61 caattacaat aaagaagcta aaattgtggt caaactcaca aacattttta ttatatacat
121 tttagtagct gatgcttata aaagcaatat ttaaatcgta aacaacaaat aaaataaaat
181 ttaaacgatg tgattaagag ccaaaggtcc tctagaaaaa ggtatttaag caacggaatt
241 cctttgtgtt acattcttga atgtcgctcg cagtgacatt agcattccgg tactgttggt
301 aaaatggaag acgccaaaaa cataaagaaa ggcccggcgc cattctatcc tctagaggat
361 ggaaccgctg gagagcaact gcataaggct atgaagagat acgccctggt tcctggaaca
421 attgcttttg tgagtatttc tgtctgattt ctttcgagtt aacgaaatgt tcttatgttt
481 ctttagacag atgcacatat cgaggtgaac atcacgtacg cggaatactt cgaaatgtcc
541 gttcggttgg cagaagctat gaaacgatat gggctgaata caaatcacag aatcgtcgta
601 tgcagtgaaa actctcttca attctttatg ccggtgttgg gcgcgttatt tatcggagtt
661 gcagttgcgc ccgcgaacga catttataat gaacgtaagc accctcgcca tcagaccaaa
721 gggaatgacg tatttaattt ttaaggtgaa ttgctcaaca gtatgaacat ttcgcagcct
781 accgtagtgt ttgtttccaa aaaggggttg caaaaaattt tgaacgtgca aaaaaaatta
841 ccaataatcc agaaaattat tatcatggat tctaaaacgg attaccaggg atttcagtcg
901 atgtacacgt tcgtcacatc tcatctacct cccggtttta atgaatacga ttttgtacca
961 gagtcctttg atcgtgacaa aacaattgca ctgataatga attcctctgg atctactggg
1021 ttacctaagg gtgtggccct tccgcataga actgcctgcg tcagattctc gcatgccagg
1081 tatgtcgtat aacaagagat taagtaatgt tgctacacac attgtagaga tcctattttt
1141 ggcaatcaaa tcattccgga tactgcgatt ttaagtgttg ttccattcca tcacggtttt
1201 ggaatgttta ctacactcgg atatttgata tgtggatttc gagtcgtctt aatgtataga
1261 tttgaagaag agctgttttt acgatccctt caggattaca aaattcaaag tgcgttgcta
1321 gtaccaaccc tattttcatt cttcgccaaa agcactctga ttgacaaata cgatttatct
1381 aatttacacg aaattgcttc tgggggcgca cctctttcga aagaagtcgg ggaagcggtt
1441 gcaaaacggt gagttaagcg cattgctagt atttcaaggc tctaaaacgg cgcgtagctt
1501 ccatcttcca gggatacgac aaggatatgg gctcactgag actacatcag ctattctgat
1561 tacacccgag ggggatgata aaccgggcgc ggtcggtaaa gttgttccat tttttgaagc
1621 gaaggttgtg gatctggata ccgggaaaac gctgggcgtt aatcagagag gcgaattatg
1681 tgtcagagga cctatgatta tgtccggtta tgtaaacaat ccggaagcga ccaacgcctt
1741 gattgacaag gatggatggc tacattctgg agacatagct tactgggacg aagacgaaca
1801 cttcttcata gttgaccgct tgaagtcttt aattaaatac aaaggatatc aggtaatgaa
1861 gatttttaca tgcacacacg ctacaatacc tgtaggtggc ccccgctgaa ttggaatcga
1921 tattgttaca acaccccaac atcttcgacg cgggcgtggc aggtcttccc gacgatgacg
1981 ccggtgaact tcccgccgcc gttgttgttt tggagcacgg aaagacgatg acggaaaaag
2041 agatcgtgga ttacgtcgcc agtaaatgaa ttcgttttac gttactcgta ctacaattct
2101 tttcataggt caagtaacaa ccgcgaaaaa gttgcgcgga ggagttgtgt ttgtggacga
2161 agtaccgaaa ggtcttaccg gaaaactcga cgcaagaaaa atcagagaga tcctcataaa
2221 ggccaagaag ggcggaaagt ccaaattgta aaatgtaact gtattcagcg atgacgaaat
2281 tcttagctat tgtaatatta tatgcaaatt gatgaatggt aattttgtaa ttgtgggtca
2341 ctgtactatt ttaacgaata ataaaatcag gtataggtaa ctaaaaa
3.3. Codon optimization
According to the genetic code, there are fewer amino acids than codon possibilities (see chart below, image credit cdn.prod.website-files.com). Thus, in theory, multiple codons can encode for the same amino acid. But in practice, spatial configuration and kinetics factors affect the translation process. For instance, the use of some codons ressembling the STOP codons can interrupt prematurely the translation process. Thus, codon optimization is an important step when designing a nucleotide sequence.
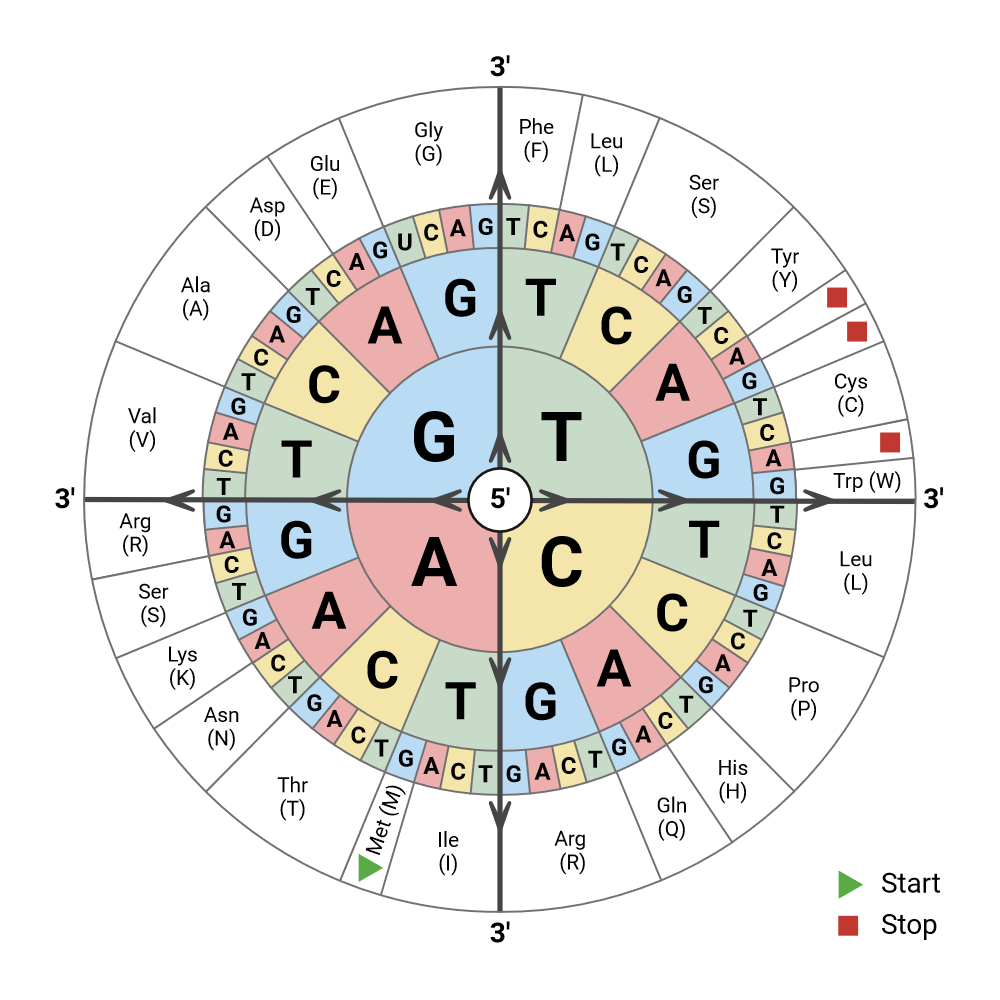

Luciferas protein DNA sequence with Codon-Optimization
TO BE COMPLETED ON MARCH 31st.


(c) Flo Razoux, AI generated
DOCUMENTATION
(1) Searching for “uterus image vector” on the web (2) Loading best image (simple graphic design and good color contrast) in Automation Art Interface (3) Adjusting Zoom (0.9), Brightness (70%), Contrast (110%) Pixelation (-10%) and Saturation (300%) (4) Refining design manually (incl. testing different colorations) (5) Exporting coordinates: mscarlet_i_points = [(-7.7, 20.9),(-5.5, 20.9),(-3.3, 20.9),(-1.1, 20.9),(1.1, 20.9),(3.3, 20.9),(5.5, 20.9),(7.7, 20.9),(-14.3, 18.7),(-12.1, 18.7),>(-9.9, 18.7),(-7.7, 18.7),(7.7, 18.7),(9.9, 18.7),(12.1, 18.7),(14.3, 18.7),(-29.7, 16.5),(-27.5, 16.5),(-25.3, 16.5),(-23.1, 16.5),(-20.9, 16.5),(-18.7, 16.5),(-16.5, 16.5),(-14.3, 16.5),(-12.1, 16.5),(-5.5, 16.5),(-3.3, 16.5),(-1.1, 16.5),(1.1, 16.5),(3.3, 16.5),(5.5, 16.5),(12.1, 16.5),(14.3, 16.5),(16.5, 16.5),(18.7, 16.5),(20.9, 16.5),(23.1, 16.5),(25.3, 16.5),(27.5, 16.5),(29.7, 16.5),(-31.9, 14.3),(-29.7, 14.3),(-27.5, 14.3),(-25.3, 14.3),(-23.1, 14.3),(-20.9, 14.3),(-18.7, 14.3),(-16.5, 14.3),(-9.9, 14.3),(-7.7, 14.3),(-5.5, 14.3),(-3.3, 14.3),(-1.1, 14.3),(1.1, 14.3),(3.3, 14.3),(5.5, 14.3),(7.7, 14.3),(9.9, 14.3),(16.5, 14.3),(18.7, 14.3),(20.9, 14.3),(23.1, 14.3),(25.3, 14.3),(27.5, 14.3),(29.7, 14.3),(31.9, 14.3),(-34.1, 12.1),(-31.9, 12.1),(-29.7, 12.1),(-16.5, 12.1),(-14.3, 12.1),(-12.1, 12.1),(-9.9, 12.1),(-7.7, 12.1),(-5.5, 12.1),(-3.3, 12.1),(-1.1, 12.1),(1.1, 12.1),(3.3, 12.1),(5.5, 12.1),(7.7, 12.1),(9.9, 12.1),(12.1, 12.1),(16.5, 12.1),(29.7, 12.1),(31.9, 12.1),(34.1, 12.1),(-34.1, 9.9),(-29.7, 9.9),(-14.3, 9.9),(-12.1, 9.9),(-9.9, 9.9),(-7.7, 9.9),(-5.5, 9.9),(-3.3, 9.9),(-1.1, 9.9),(1.1, 9.9),(3.3, 9.9),(5.5, 9.9),(7.7, 9.9),(9.9, 9.9),(14.3, 9.9),(29.7, 9.9),(34.1, 9.9),(-34.1, 7.7),(-29.7, 7.7),(-27.5, 7.7),(-25.3, 7.7),(-14.3, 7.7),(-12.1, 7.7),(-9.9, 7.7),(-7.7, 7.7),(-5.5, 7.7),(-3.3, 7.7),(-1.1, 7.7),(1.1, 7.7),(3.3, 7.7),(5.5, 7.7),(7.7, 7.7),(12.1, 7.7),(14.3, 7.7),(25.3, 7.7),(27.5, 7.7),(34.1, 7.7),(-34.1, 5.5),(-23.1, 5.5),(-12.1, 5.5),(-9.9, 5.5),(-7.7, 5.5),(-5.5, 5.5),(-3.3, 5.5),(-1.1, 5.5),(1.1, 5.5),(3.3, 5.5),(5.5, 5.5),(7.7, 5.5),(12.1, 5.5),(23.1, 5.5),(34.1, 5.5),(-34.1, 3.3),(-23.1, 3.3),(-20.9, 3.3),(-12.1, 3.3),(-9.9, 3.3),(-7.7, 3.3),(-5.5, 3.3),(-3.3, 3.3),(-1.1, 3.3),(1.1, 3.3),(3.3, 3.3),(5.5, 3.3),(12.1, 3.3),(20.9, 3.3),(23.1, 3.3),(34.1, 3.3),(-31.9, 1.1),(-18.7, 1.1),(-12.1, 1.1),(-9.9, 1.1),(-7.7, 1.1),(-5.5, 1.1),(-3.3, 1.1),(-1.1, 1.1),(1.1, 1.1),(3.3, 1.1),(5.5, 1.1),(9.9, 1.1),(12.1, 1.1),(18.7, 1.1),(31.9, 1.1),(-31.9, -1.1),(-18.7, -1.1),(-9.9, -1.1),(-7.7, -1.1),(-5.5, -1.1),(-3.3, -1.1),(-1.1, -1.1),(1.1, -1.1),(3.3, -1.1),(9.9, -1.1),(18.7, -1.1),(31.9, -1.1),(-29.7, -3.3),(-27.5, -3.3),(-20.9, -3.3),(-9.9, -3.3),(-7.7, -3.3),(-5.5, -3.3),(-3.3, -3.3),(-1.1, -3.3),(1.1, -3.3),(3.3, -3.3),(7.7, -3.3),(9.9, -3.3),(20.9, -3.3),(27.5, -3.3),(29.7, -3.3),(-25.3, -5.5),(-23.1, -5.5),(-7.7, -5.5),(-5.5, -5.5),(-3.3, -5.5),(-1.1, -5.5),(1.1, -5.5),(3.3, -5.5),(7.7, -5.5),(23.1, -5.5),(25.3, -5.5),(-7.7, -7.7),(-5.5, -7.7),(-3.3, -7.7),(-1.1, -7.7),(1.1, -7.7),(7.7, -7.7),(-7.7, -9.9),(-3.3, -9.9),(-1.1, -9.9),(1.1, -9.9),(7.7, -9.9),(-7.7, -12.1),(-3.3, -12.1),(-1.1, -12.1),(1.1, -12.1),(3.3, -12.1),(7.7, -12.1),(-7.7, -14.3),(-5.5, -14.3),(5.5, -14.3),(7.7, -14.3),(-5.5, -16.5),(5.5, -16.5),(-5.5, -18.7),(-3.3, -18.7),(3.3, -18.7),(5.5, -18.7),(-5.5, -20.9),(-3.3, -20.9),(3.3, -20.9),(5.5, -20.9),(-5.5, -23.1),(-3.3, -23.1),(3.3, -23.1),(5.5, -23.1),(-5.5, -25.3),(-3.3, -25.3),(3.3, -25.3),(5.5, -25.3),(-5.5, -27.5),(-3.3, -27.5),(3.3, -27.5),(5.5, -27.5),(-5.5, -29.7),(-3.3, -29.7),(3.3, -29.7),(5.5, -29.7)] mko2_points = [(14.3, 12.1),(12.1, 9.9),(-31.9, 7.7),(9.9, 7.7),(31.9, 7.7),(-29.7, 5.5),(9.9, 5.5),(29.7, 5.5),(-29.7, 3.3),(-27.5, 3.3),(-25.3, 3.3),(7.7, 3.3),(9.9, 3.3),(25.3, 3.3),(27.5, 3.3),(29.7, 3.3),(-29.7, 1.1),(-27.5, 1.1),(-25.3, 1.1),(-23.1, 1.1),(-20.9, 1.1),(7.7, 1.1),(20.9, 1.1),(23.1, 1.1),(25.3, 1.1),(27.5, 1.1),(29.7, 1.1),(-27.5, -1.1),(-25.3, -1.1),(-23.1, -1.1),(5.5, -1.1),(7.7, -1.1),(23.1, -1.1),(25.3, -1.1),(27.5, -1.1),(5.5, -3.3),(5.5, -5.5),(3.3, -7.7),(5.5, -7.7),(3.3, -9.9)] mrfp1_points = [(-5.5, 18.7),(-3.3, 18.7),(-1.1, 18.7),(1.1, 18.7),(3.3, 18.7),(5.5, 18.7),(-9.9, 16.5),(-7.7, 16.5),(7.7, 16.5),(9.9, 16.5),(-14.3, 14.3),(-12.1, 14.3),(12.1, 14.3),(14.3, 14.3),(-31.9, 9.9),(31.9, 9.9),(29.7, 7.7),(-31.9, 5.5),(-27.5, 5.5),(-25.3, 5.5),(25.3, 5.5),(27.5, 5.5),(31.9, 5.5),(-31.9, 3.3),(31.9, 3.3),(-29.7, -1.1),(-20.9, -1.1),(20.9, -1.1),(29.7, -1.1),(-25.3, -3.3),(-23.1, -3.3),(23.1, -3.3),(25.3, -3.3),(-5.5, -9.9),(5.5, -9.9),(-5.5, -12.1),(5.5, -12.1),(-3.3, -14.3),(-1.1, -14.3),(1.1, -14.3),(3.3, -14.3),(-3.3, -16.5),(-1.1, -16.5),(1.1, -16.5),(3.3, -16.5),(-1.1, -18.7),(1.1, -18.7),(-1.1, -20.9),(1.1, -20.9),(-1.1, -23.1),(1.1, -23.1),(-1.1, -25.3),(1.1, -25.3),(-1.1, -27.5),(1.1, -27.5)]
(6) Laptop/internet crashed before submission for publication in the gallery 😢
DOCUMENTATION ChatGPT was used to (1) understand code lines from the CoLab examples, (2) understand and fix errors when trying to write my own code from the CoLab examples, (3) and later when trying to adapt the code downloaded from Ronan Donovan’s AAI. Step (2) and (3) were not successful despite many attempts. At last, documentation of other students were checked and the code of the design Golden Lyre by Katherine Kolin was used as a base. The code was easily adapted and the first test images were generated in Colab quickly after.
Finally, coloring tests were made to get as close as possible of the fluorescent images published in the gallery
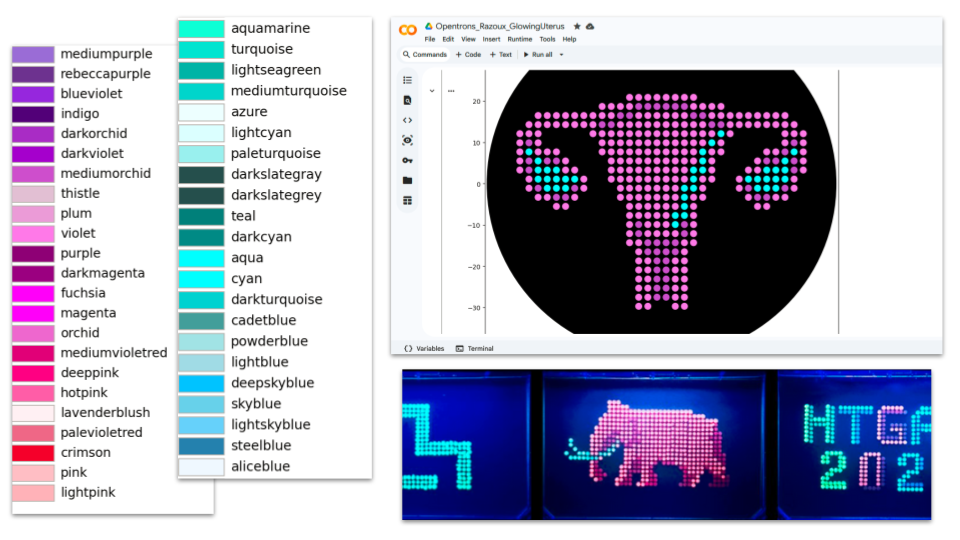
To be updated on March 10th
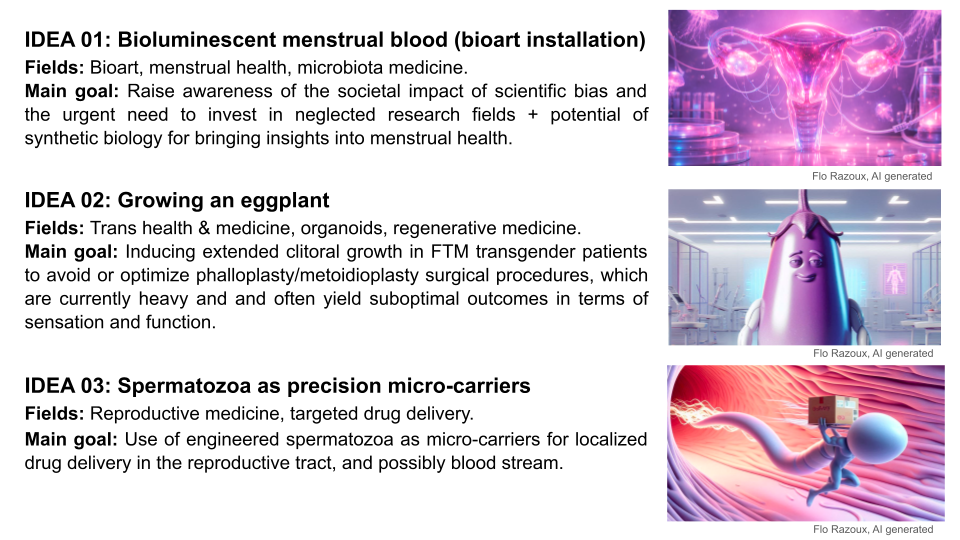



FINAL PROJECT IDEA 01 | BIOLUMINESCENT MENSTRUAL BLOOD (BIOART INSTALLATION)
Fields: Bioart, menstrual health, microbiota medicine.
Main goal: Raise awareness of the societal impact of scientific bias and the urgent need to invest in neglected research fields + potential of synthetic biology for bringing insights into menstrual health.
Methods: Option 01 Change the genome of bioluminescent marine microorganisms to help them adapt to a liquid culture made of menstrual blood serum and elicit a photonic response under specific stimuli (biosensor approach) Option 02 Insert luciferase/luciferin genes into vaginal microbiota to monitor early “cellular ecosystem” biomarkers (biosensor approach) Option 03 Create a protein ressembling hemoglobin with luciferase properties that produce light when binding to dioxygen (design approach)
Documentation: https://pages.htgaa.org/2026a/flo-razoux/homework/week-01-hw-principles-and-practices/hw-governance/index.html
FINAL PROJECT IDEA 02 | GROWING AN EGGPLANT
Fields: Trans health & medicine, organoids, regenerative medicine.
Main goal: Inducing extended clitoral growth in FTM transgender patients to avoid or optimize phalloplasty/metoidioplasty surgical procedures (hybrid methods), which are currently heavy and often yield suboptimal outcomes in terms of sensation and function.
Methods: Gene-circuit strategy applied locally (corpora cavernosa) to reopen or bypass embryonic gene programs that locked the penile or/and clitoral architecture. Reactivation of the expression of the genes involved in tissue growth such as androgens receptors or SRY downstream targets.
Documentation: What is Bottom Growth? A trans perspective: https://www.tiktok.com/@bugandalex/video/7287946425164434720 ChatGPT, prompt: “What are the biological mechanisms underlying bottom-growth, and why does growth stops at the some point?” https://chatgpt.com/s/t_699c7b5178e48191aa41ad42cddf2e9b ChatGPT, prompt: “According to previous discussion, how could synthetic biology be implemented to “unlock” the bottom-growth ceiling?” https://chatgpt.com/s/t_699c7c245f1081919576d9e7d082bc98
FINAL PROJECT IDEA 03 | ENGINEERED SPERMATOZOA AS PRECISION MICRO-CARRIERS
Fields: Reproductive medicine, targeted drug delivery.
Main goal: Use of engineered spermatozoa as micro-carriers for localized drug delivery in the reproductive tract (and in a second step, possibly via blood stream).
Methods: This would depend on the application: targeted uterine/ovarian cancer, anti-proliferative therapy in endometriosis, or contraceptive innovation.
Documentation: ChatGPT, prompt: “Write a list of applications on how sperm cells could be used to deliver drugs locally, with a focus on reproductive health” https://chatgpt.com/s/t_699c7e1d63f88191a129fdab9c3b69a4
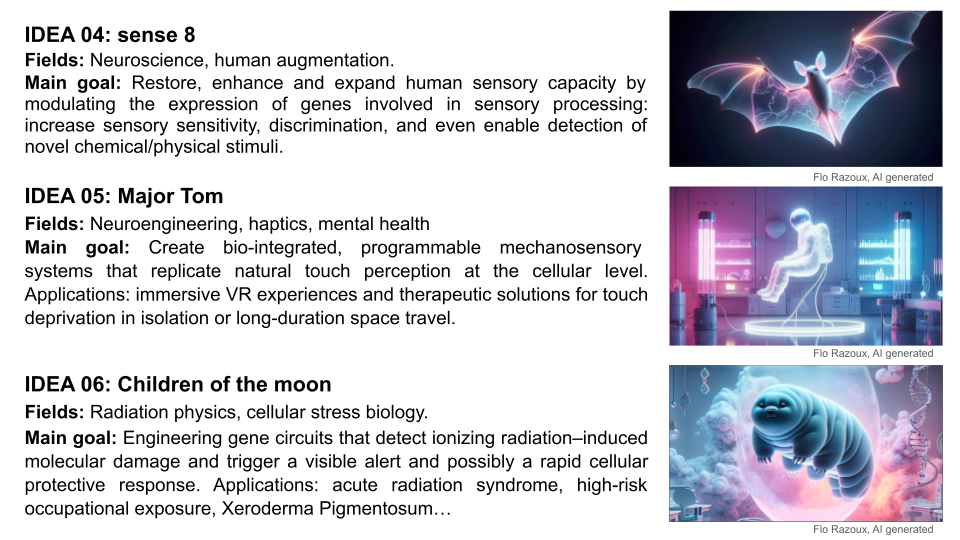
FINAL PROJECT IDEA 04 | SENSE 8
Fields: Neuroscience, human augmentation.
Main goal: Restore, enhance and expand human sensory capacity by modulating the expression of genes involved in sensory processing: increase sensory sensitivity, discrimination, and even enable detection of novel chemical/physical stimuli.
Documentation: ChatGPT, prompt: “How would you pitch a project on using synthetic biology to enhance senses (especially the olfaction). Please suggest different directions to explore” https://chatgpt.com/s/t_699e3902307c8191bb9abf5aa7add2d7
FINAL PROJECT IDEA 05 | MAJOR TOM
Fields: Neuroengineering, haptics, mental health.
Main goal: Create bio-integrated, programmable mechanosensory systems that replicate natural touch perception at the cellular level. Applications: immersive VR experiences and therapeutic solutions for touch deprivation in isolation or long-duration space travel.
Documentation: ChatGPT, prompt: “How would you pitch a project on using synthetic biology to mimic the sense of touch to be implemented for instance in VR, or to avoid touch deprivation in case of isolation or during long-duration spaceflight. Please include biology background (mecano receptors in the skin specific to different types of touch)” https://chatgpt.com/s/t_699e3ad59224819180390539c385bc9e
FINAL PROJECT IDEA 06 | CHILDREN OF THE MOON
Fields: Radiation physics, cellular stress biology.
Main goal: Engineering gene circuits that detect ionizing radiation–induced molecular damage and trigger a visible alert (e.g. reversible pigmentation change) and possibly a rapid cellular protective response. Applications: acute radiation syndrom, high-risk occupational exposure, Xeroderma Pigmentosum.
Documentation: ChatGPT, prompt: “How would you pitch a project on using synthetic biology to insert a biosensing mechanism in skin cells that could trigger either an alarm (change of skin pigmentation for instance) or a protective cellular response to an immediate risk of acute irradiation exposure? Please include physics background of radiation contamination monitoring devices, biology background on the effects of radiation exposure at the cellular and molecular levels as well as possible protective strategies, if any.” https://chatgpt.com/s/t_699e4925d558819181d04229c849199b
(c) Flo Razoux with use of AI
All questions were answered using the feedback of Gemini - and ChatGPT for Question 10 - as a research starting point. Questions were used as prompts with an oriented approach (what would Shuguang Zhang answer, how would you explain a 10 year old etc.). Sources were checked and prompts refined from the larger picture into details of interest, as well as when needed if the content was unclear to me.
1. How many molecules of amino acids do you take with a piece of 500 grams of meat? (on average an amino acid is ~100 Daltons)
According to Gemini, meat generally contains approx 25% (+/- 5%) of protein by weight when cooked. Thus, for this calculation, we assume that a 500g piece of meat contains 125g of protein (5x25g). By convention, 1g = 6.022 x 10 ^ 23 Da , so 125g = 7,528 x 10 ^ 25 Da.
Thus, our piece of meat contains 7,528 x 10^23 molecules of amino acids (7,528 x 10^25 /100) : a lot of work awaits pepsin, trypsin and peptidase at the burger shop 🍔
2. Why do humans eat beef/fish but do not become a cow/fish?
Although the idiom “You are what you eat” suggests food directly becomes part of us, proteins ingested during meals are not directly incorporated into the human body: rather, they are first broken down into amino acids through the digestive process and then serve as building blocks for human proteins according to the DNA code. Proteins are species-specific and their production depends on both the genetic code and environmental factors.
3. Why are there only 20 natural amino acids?
The existence of only 20 primary natural amino acids is generally explained by an evolutionary optimization. Organisms evolve to optimize the balance between energy/resource consumption and the benefits derived from biological functions. These specific building blocks were selected over 4 billion years ago for their ability to form stable, functional, and soluble proteins. They provide enough diversity to support all necessary biological functions while remaining cost-efficient and easier to handle by the cell machinery than a larger set of building blocks.
Why twenty amino acid residue types suffice(d) to support all living systems
Teaching the principle of biological optimization
4. Can you make other non-natural amino acids? Design some new amino acids.
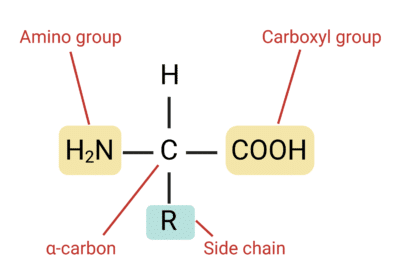
Yes, Shuguang Zhang and George Church have pushed the boundaries of molecular design by developing methods to produce non-natural amino acids. Amino acids are composed of an amino group (-NH2), a carboxyl group (-COOH) and a residue group (-R) that varies. To create new amino acids, one needs to design new residue groups. Example of new residues and how to synthesize them: SKIP for now ⏱
5. Where did amino acids come from before enzymes that make them, and before life started?
Scientists think that life originated from a “primordial soup” in deep-sea hydrothermal vents. Despite extreme heat and high pressure, these environments are teeming with life. These vents emit hot, mineral-rich fluids, creating environments where simple molecules could undergo chemical reactions to form more complex organic compounds, including amino acids. In 1952, Miller and Urey simulated early Earth conditions in an experiment and demonstrated that amino acids can form spontaneously through non-enzymatic pathways. It is also been proposed that amino acids may have an extraterrestrial origin. Amino acids have been found within meteorites that have crashed to Earth and in samples returned directly from asteroids, indicating that the chemical ingredients for life as we know it are widespread in the solar system 👽
A Short Tale of the Origin of Proteins and Ribosome Evolution
Insights into the formation and evolution of extraterrestrial amino acids from the asteroid Ryugu
6. If you make an α-helix using D-amino acids, what handedness (right or left) would you expect?
In humans, the right-handed α-helix is the most common structural arrangement in the secondary structure of proteins. These helices are composed of L-amino acids. L-amino acids and D-amino acids are stereoisomer mirror images of each other, differing in the placement of the amino group (-NH2) on the alpha-carbon (see paper given in reference below). In a Fischer projection, L-amino acids have the amino group on the left (left handed), while D-amino acids have it on the right (right handed). Thus, if building an α-helix using D-amino acids, one would expect the helix to be oriented in the opposite direction and obtain a left-handed α-helix.
Structure and Function of Proteins
7. Can you discover additional helices in proteins?
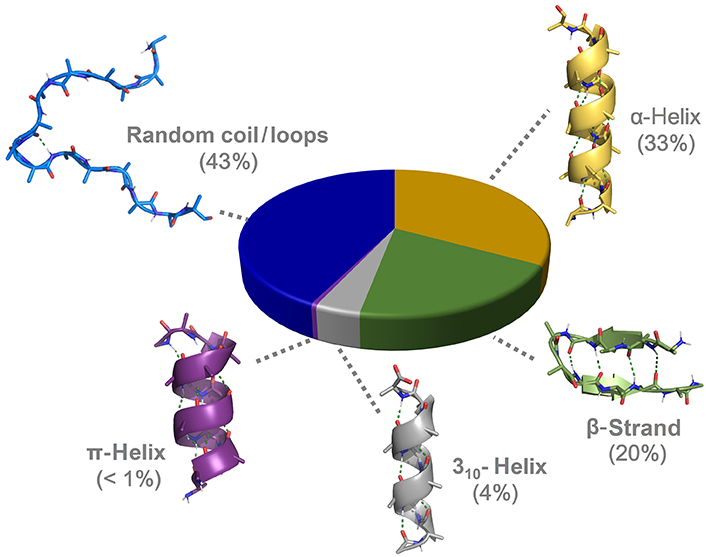
After decades of intensive focus on the study of the α-helices, other helicoidal structures have been and keep being discovered, or rather reclassified after slowly regaining a certain relevance in protein science:
8. Why are most molecular helices right-handed?
During evolution, L-amino acids were preferred for protein synthesis and main metabolism: ribosomes possess a remarkable ability for chiral selectivity and are designed to use L-amino acids for protein synthesis. This explains why most molecular helices found in nature are right-handed (see Question 6 for reference).
To go a bit further: In the early stage of amino acids discovery, scientists actually believed that L-amino acids were solely found in nature and D-amino acids are artificial products. However, with the development of analytical methods in the past decades, D-amino acids have been found in a wide variety of living organisms both in their free form and as isomeric residues in many proteins. Their various biological functions are closely relevant to human physiology and diseases, including cancer. Although not typically formed by the translation machinery, left-handed helices can still be found in nature and are typically built through post-translational modification, non-ribosomal peptide synthetases (NRPS), and the incorporation of glycine and achiral residues.
Natural Occurrence, Biological Functions, and Analysis of D-Amino Acids
d‐amino acids: new functional insights
9. Why do β-sheets tend to aggregate? What is the driving force for β-sheet aggregation?
The aggregation of β-sheets is a phenomenon of molecular self-assembly that is due to hydrophobicity and structural complementation. A β-sheet consists of β-strands that are connected laterally by at least two or three backbone hydrogen bonds, forming a generally twisted, pleated sheet. Because the structure of their edges is complementary (unpaired amino and carboxyl groups just waiting to find their match 💘), adjacent β-sheets naturally bond via their “sticky edges”. But what drives adjacent β-sheets to pile up like pancakes is linked to the amphiphilic nature of the β-sheets. Indeed, one of their sides is hydrophilic and the other is hydrophobic. In the same way that oil droplets gather together when added to water, β-sheets tend to aggregate in order to “hide” their hydrophobic sides from hydrophilic groups or aqueous environment (nucleoplasm, cytoplasm and ground substance).
The Supramolecular Chemistry of β-Sheets
10. Why do many amyloid diseases form β-sheets? Can you use amyloid β-sheets as materials?
Amyloids have been linked to the development of various neurodegenerative diseases such as Alzheimer’s and Parkinson’s. Pathogenic amyloids form when previously healthy proteins misfold: these proteins lose their normal structure and physiological functions, and start forming fibrous deposits within and around cells that cause the progressive disruption of brain functions. Amyloid fibrils form from different proteins, each associated with a particular disease, but they all contain a distinctive dysfunctional β-sheet pattern known as cross-β spine. When the peptides misfold, they align next to each other forming extended β-sheets that present highly stable and ordered structures. Once a small cluster of misfolded peptides is formed, a nucleation effect causes the misfolding and aggregation of other adjacent peptides, leading to an accelerated amplification of the self-propagating aggregation process. Furthermore, the aggregation of cross-β-sheets leads to the creation of steric zippers that makes the amyloids resistant to enzyme degradation (proteolysis) allowing them to accumulate in tissues and organs over time.
.
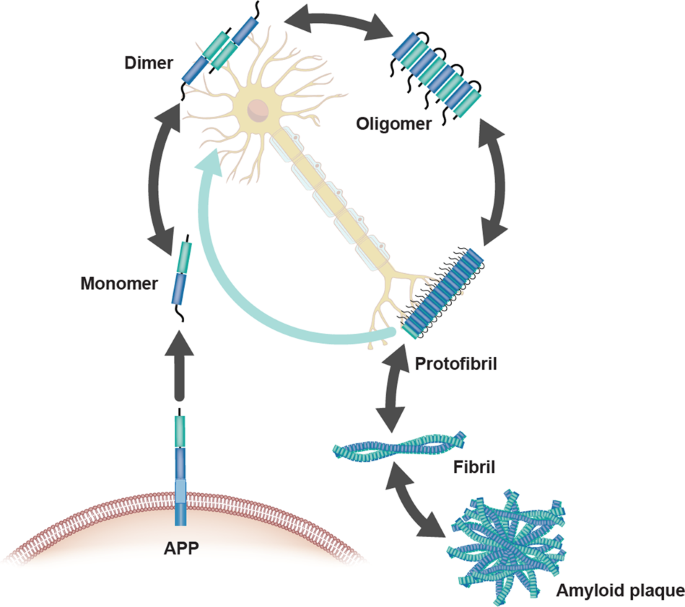
The Amyloid-β Pathway in Alzheimer’s Disease
.
One man’s loss is another man’s gain… The nanocrystal properties of cross-β sheet aggregates make them suitable for the engineering of biomaterials that can better withstand thermal stress and chemical denaturation such as:
.

Amyloid-induced mineralization: From biological systems to biomimetic materials
.
11. Design a β-sheet motif that forms a well-ordered structure.
SKIP for now ⏱
For this week’s assignments, we will keep focusing on the first final project idea of creating bioluminescent menstrual blood. While it would be interesting to have a closer look at the structure of hemoglobin, the protein that facilitates the transportation of oxygen in erythrocytes (red blood cells) and gives its red color to human blood in visible light, we will keep studying the luciferin 4-monooxygenase. This enzyme, commonly known as firefly luciferase, catalyses the production of light through the oxidation of luciferin. The structure of this protein is simpler than hemoglobin, so this seems to be an ideal option to apply what has been covered in class this week. We might go back to hemoglobin, myoglobin and other proteins determining blood color later depending on how the project develops.
Length: 550 amino acids
Most frequent amino acid: Leucine
Homology: Luciferase has 250 homologs across insects (endopterygota, 95% ; polyneoptera 3%; paraneoptera %) and bacteria (allobacillus, 1%). VISUALISATION
Family: Protein families refers to groups of closely related proteins with high sequence/functional similarity and common ancestry. The firefly luciferase belongs to the acyl-adenylate/thioester-forming superfamily of enzymes, also known as the ANL superfamily or the ATP-dependent AMP-binding enzyme family.
Documentation Sequence Length and Amino Acids Frequency
AA sequence obtained from NCBI and Uniprot database (see WEEK 02 HW)
Colab code used: https://colab.research.google.com/drive/1vlAU_Y84lb04e4Nnaf1axU8nQA6_QBP1
Output: Sequence Length: 550 Amino Acid Frequencies: l: 52 g: 46 v: 44 a: 42 k: 40 i: 37 e: 33 d: 30 f: 30 p: 29 t: 29 s: 29 r: 21 n: 19 y: 19 q: 16 m: 14 h: 14 c: 4 w: 2
Documentation Homology Calculation runned with BLAST: FULL REPORT
RCSB page: https://www.rcsb.org/structure/1LCI
Structure Deposition Date: 1996-06-01
Crystal structure of firefly luciferase throws light on a superfamily of adenylate-forming enzymes
Resolution: 2.00 - 2.20 Å
The resolution is <2.70 Å and therefore considered as good.
Additional molecule(s) in the representation: none
Here is a page which presents the structure of the luciferase complexed with oxyluciferin and AMP (products of the reaction catalyzed by the luciferase): https://www.rcsb.org/structure/2D1R
Associated protein families:
Documentation Family Attribution
Calculation with SCOP
Output SCOP search for P08659 uniprot ID: 4 domains: 8025243 1BA3 A:3-434 8037622 1BA3 A:3-434 8055416 1BA3 A:435-520 8055417 1BA3 A:435-520
Output SCOP search for luciferas keyword: 4001964 Dinoflagellate luciferase repeat 4003312 Bacterial luciferase (alkanal monooxygenase)
AI GEMINI Feedback on protein family: AMP-dependent synthetase/ligase family :cross_mark:; Adenylate-forming enzyme superfamily :cross_mark:; AMP-binding enzyme C-terminal domain :check_mark_button:
All the visualizations and counts were carried out using the step-by-step guidance of ChatGPT (indeed, very useful). Questions were used as prompts. Once familiar with the software and tasks, further explorations were carried out on my own. Detailed documentation can be found at the end of this section.
PDB 1LCI: first luciferase crystal structure, open conformation
For luciferin ligand and exploration of other conformations: PDB 2D1S and PDB 4G36
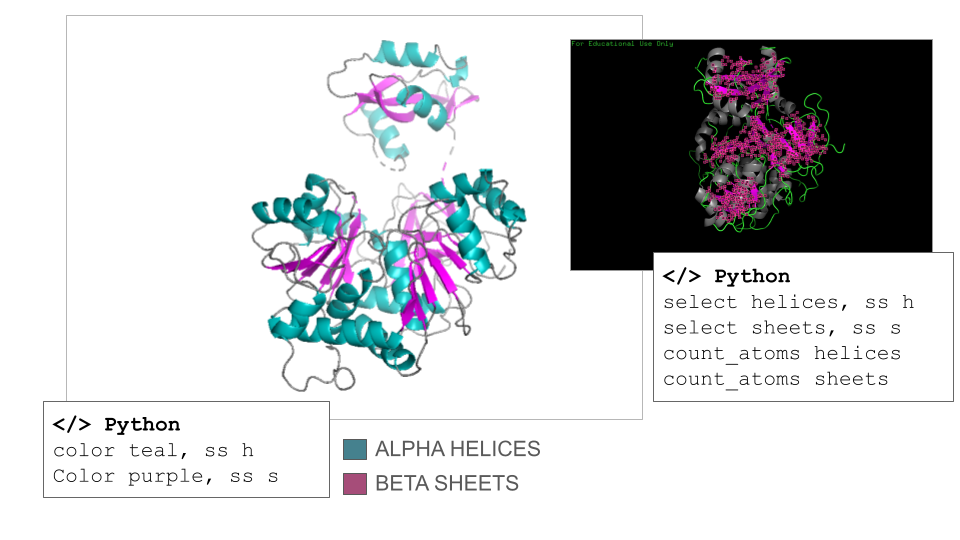




According to ChatGPT, the Firefly Luciferase is an α/β protein, but it is helix-dominated, which is typical for enzymes in the adenylate-forming enzyme family. This fact was surprising to me since I counted 17 helices and 21 sheets on the model when navigating it manually. However, the counts confirmed that the secondary structure of the Firefly Luciferase has more helix than sheets (count_atoms helices: 1367 atoms; count_atoms sheets: 817 atoms).
There are discrepancies in the classification of the amino acids depending on the sources that can be find online. Thus, I used the amino acids reference chart published by the pharma company Merck KGaA (Darmstadt, Germany) as reference. There are also many ways residues can be represented depending on their classification, so I opted for a binary hydrophobic/hydrophilic approach to better fit the direction of the question.
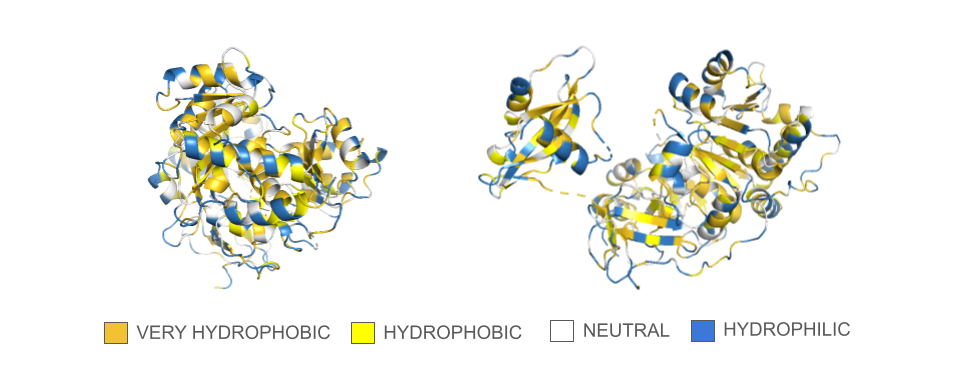

In the spin animation below, one can observe how the protein follows the hydrophobic effect, i.e. how the hydrophobic residues cluster inside the protein. The presence of hydrophilic residues on the surface of the luciferase is coherent with the fact that this enzyme is soluble.
Showing the surface of the protein allows us to see more clearly that the protein is composed of two domains that have a globular shape and present small surface grooves. A large cleft between the N-terminal (residues 1-436, bottom part in the animation) and C-terminal (residues 440–550, on top) domains of the protein is clearly visible when rotating the structure: one can assume that it accommodates the luciferin and the ATP molecule (reference chemical reaction).
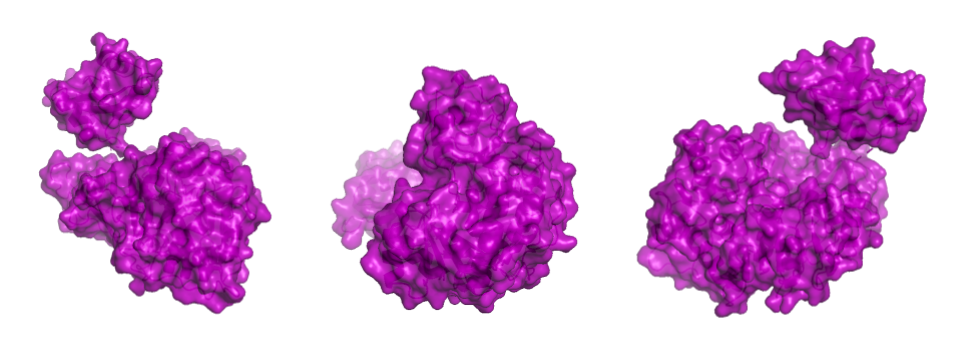

In a second step, the ligand was added into the visualization. Residues reported to be present in the binding pocket and active site of the firefly luciferase were highlighted using the coloring function. One can observe how the ligand and the key residues match beautifully the binding pocket and how inside the cleft, the ligand enters a tunnel-like cavity forming inside the N-terminal domain.
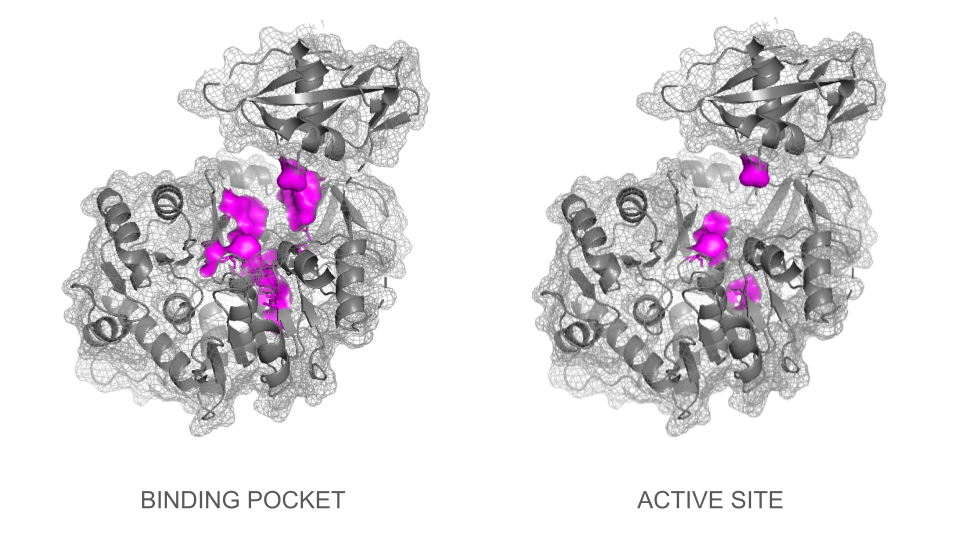

Loading the data set 2D1S containing both luciferase and ligand was confusing at first: the protein looked different than the 1LCI model. This can be explained by the fact that the enzyme undergoes large conformational changes during the different catalysis states: when the conformation is open, the enzyme allows the ligands to enter the cleft and when it is closed, the catalysis can occur. The C-terminal domain rotates perpendicularly when switching from one state to another, making the structures look quite different even though they are the same protein. Only the open conformation is presented here.
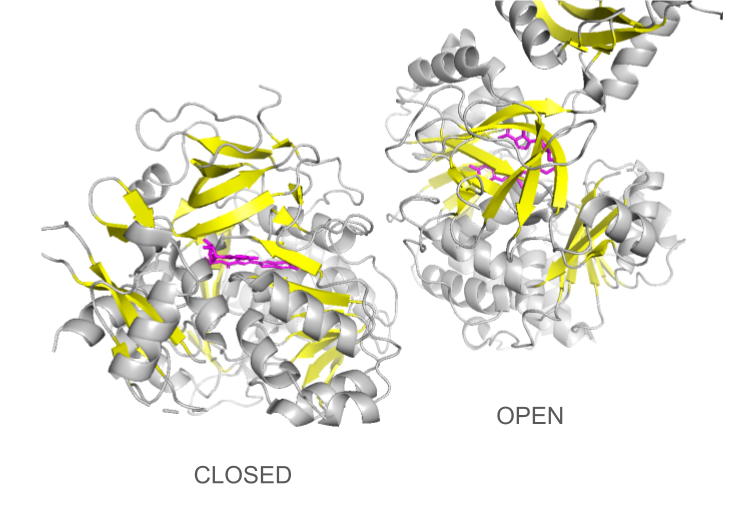

Documentation Visualization in PyMol: Basics
Import dataset: File > Get PDB > Enter 1LCI (PDB ID for Firefly Luciferase) or </> Python command
Useful commands recommended by ChatGPT:
- fetch 1lci (import dataset)
- hide everything
- show cartoon / show ribbon / show sticks / show spheres
- set sphere_scale, 0.25 / set stick_radius, 0.15 (adjust scale to improve 3D visualisation)
- util.cbag (coloring of the “stick and ball” according to PyMol default color coding: carbon, green; basic residues, blue; acidic residues, red; gray for others)
Documentation Visualization in PyMol: Secondary Structures
Useful commands recommended by ChatGPT:
- select helices, ss h / select sheets, ss s (select secondary structures)
- color color_01, ss h / color color_02, ss s (specific coloring of secondary structures)
- count_atoms helices / count_atoms sheets
Documentation Visualization in PyMol: Hydropathy
Useful (adapted) commands recommended by ChatGPT:
- select veryhydrophobic, resn phe+ile+trp+leu+val+met
- color gold, veryhydrophobic
- select hydrophobic, resn tyr+cys+ala
- color yellow, hydrophobic
- select neutral, resn thr+his+gly+ser+gln
- color white, neutral
- select hydrophilic, resn arg+lys+asn+glu+pro+asp
- color sky, hydrophilic
Documentation Visualization in PyMol: Surface
Useful commands recommended by ChatGPT:
- show surface, selection
- set transparency, 0.2
- clip slab, 20 (view of the surface + inside)
- set surface_cavity_mode, 1
- show mesh
Documentation Visualization in PyMol: Binding pocket and active site
Several strategies were tested to visualize the binding pocket and active site. First, visualization of the types of residues generally involved in binding pockets / active sites (charged, hydrophobe, polar, aromatic etc.) and then, types of residues involved specifically in the firefly luciferase. However, all the representations tested were not specific to the binding site.
Useful commands recommended by ChatGPT for part 01:
- select active, resn lys+arg+his+asp+glu
- color color_01, active
- select inactive, resn ala+ile+leu+met+val+gly+pro+cys+ser+phe+trp+tyr+thr+asn+gln
- color colo_02, inactive
In a second step, visualization of the individual residues involved in the active site of the firefly luciferase gave much better results although one needs to mention that the lists provided by Gemini and ChatGPT were different, and varied depending on iterations.
Commands recommended by ChatGPT to visualize binding pocket:
- select luciferin_binding_site, resi 198+214+218+222+244+245+247+286+340+343+344+347+420+421+422+529
- show sticks, luciferin_binding_site
- color color_01, luciferin_binding_site
- show surface, luciferin_binding_site
Commands recommended by ChatGPT to visualize active site:
- select luciferase_catalytic, resi 218+245+343+529
- show sticks, luciferase_catalytic
- color color_01, luciferase_catalytic
Documentation Visualization in PyMol: Adding ligand
Useful commands recommended by ChatGPT:
- fetch 1LCI
- fetch 4G36
- align 4G36, 1LCI
- select luciferin, 4G36 and organic (organic defines the ligand)
- create luciferin_copy, luciferin
- disable 4G36
- show sticks, luciferin_copy
- color color_01, luciferin_copy
Visual tools
Useful commands recommended by ChatGPT:
- mplay/mstop (play and stop movie)
- mclear (delete movie data to save memory when export is done)
- zoom object_of_interest, 8 (zoom power)
- bg_color white (change background color to white)
- set ray_trace_fog (better image quality)
- Color list
Spin animation
Useful (adapted) commands recommended by ChatGPT:
- mset 1 x360 (number of frames)
- util.mroll(1,360,1)
Zoom-through animation
Useful commands recommended by ChatGPT:
- mset 1 x360
- frame 1
- mview store
- frame 360
- move z, -360
- mview store
- mview interpolate
Video export Videos were exported in mp4 format using the screen recording function.
LEGAL ASPECTS: To use the official version of PyMol for free as a student, I had to sign the agreement copied below.
Addendum: 1- By declaring that I’m a ‘full time student’, I understand that being registered as a committed listener of the HTGAA course which is not an internship and involves 15 to 30 hours of weekly academic work fits the required criteria. 2- “Builds” should not be shared publicly. I understand that “builds” are a specific functionality offered by PyMol and that the visualizations that I have shared on this page are not “builds”.
PyMOL Educational Use Declaration for Flo Razoux
- I, Flo Razoux, am either a full-time student or am engaged in teaching full-time students. After being granted access, I will only apply Education-Use-Only PyMOL Builds (“Builds”) for education purposes and specifically including the following: COURSES or DEGREE: MIT Media Lab, Synthetic Biology, Other, 2026
- I will only share the Builds and their download access credentials with my fellow students and/or teachers, and only via private means.
- I will not post the Builds or their download access credentials in a publicly-accessible location, such as a web page, email list, or blog.
- If I apply PyMOL in any for-profit commercial activity or in any non-profit academic research, then I will compile my own builds from the open-source code or purchase an appropriate PyMOL Subscription in order to access the official PyMOL Builds not limited to educational use only.
- Except as otherwise set forth in Sections 1 through 4, I shall not: (i) modify, translate, adapt, create derivative works from or decompile the Builds, or any portion thereof, or create or attempt to create, by disassembling, reverse engineering or otherwise, the source code from the object code supplied hereunder, (ii) rent, lease, sell, transfer, publish, display, distribute, disclose or make the Builds available to third parties or use the builds, or any portion thereof, in a service bureau, time-sharing or outsourcing service or (iii) remove or alter any proprietary rights notices on the Builds. I acknowledge that the restrictions set forth in clauses (i) through (iii) of the immediately preceding sentence shall apply to distributions by Schrodinger, LLC of any third party software or other materials with the Builds.
TO BE COMPLETED ON MARCH 31st
GROUP MEMBERS
Diogo Custodio https://pages.htgaa.org/2026a-diogo-custodio/
Flo Razoux https://pages.htgaa.org/2026a-flo-razoux/
Katharine Kolin https://pages.htgaa.org/2026a/katharine-kolin/
Marisa Satsia https://pages.htgaa.org/2026a-marisa-satsia/
Main goals (tbc)
Proposal (to be updated on March 8th)
Individual and group plan for engineering a bacteriophage (to be updated on March 8th)
Superoxide dismutase 1 (SOD1) is a cytosolic antioxidant enzyme that converts superoxide radicals into hydrogen peroxide and oxygen. In its native state, it forms a stable homodimer and binds copper and zinc. Mutations in SOD1 cause familial Amyotrophic Lateral Sclerosis (ALS). Among them, the A4V mutation (Alanine → Valine at residue 4) leads to one of the most aggressive forms of the disease. The mutation subtly destabilizes the N-terminus, perturbs folding energetics, and promotes toxic aggregation.
CHALLENGE
1. Design short peptides that bind mutant SOD1
2. Then decide which ones are worth advancing toward therapy
SOD1 Original sequence (150 aa):
MATKAVCVLKGDGPVQGIINFEQKESNGPVKVWGSIKGLTEGLHGFHVHEFGDNTAGCTSAGPHFNPLSRKHGGPKDEERHVGDLGNVTADKDGVADVSIEDSVISLSGDHCIIGRTLVVHEKADDLGKGGNEESTKTGNAGSRLACGVIGIAQ
A4V mutated sequence:
Replace A by V on position 4 (note that the initial Methionine in position 1 is not counted)
MATKVVCVLKGDGPVQGIINFEQKESNGPVKVWGSIKGLTEGLHGFHVHEFGDNTAGCTSAGPHFNPLSRKHGGPKDEERHVGDLGNVTADKDGVADVSIEDSVISLSGDHCIIGRTLVVHEKADDLGKGGNEESTKTGNAGSRLACGVIGIAQ
Approach 01: Generation of 4 peptides (length 12 aa, top K 3)
| Index | Peptides | Pseudo Perplexity |
|---|---|---|
| 0 | WLSPVVAAEHKE | 15.485618 |
| 1 | WRYGAAAAEHKE | 9.098425 |
| 2 | WRYYAAAVAHKX | 8.351979 |
| 3 | HHSYAAAVALKK | 12.665942 |
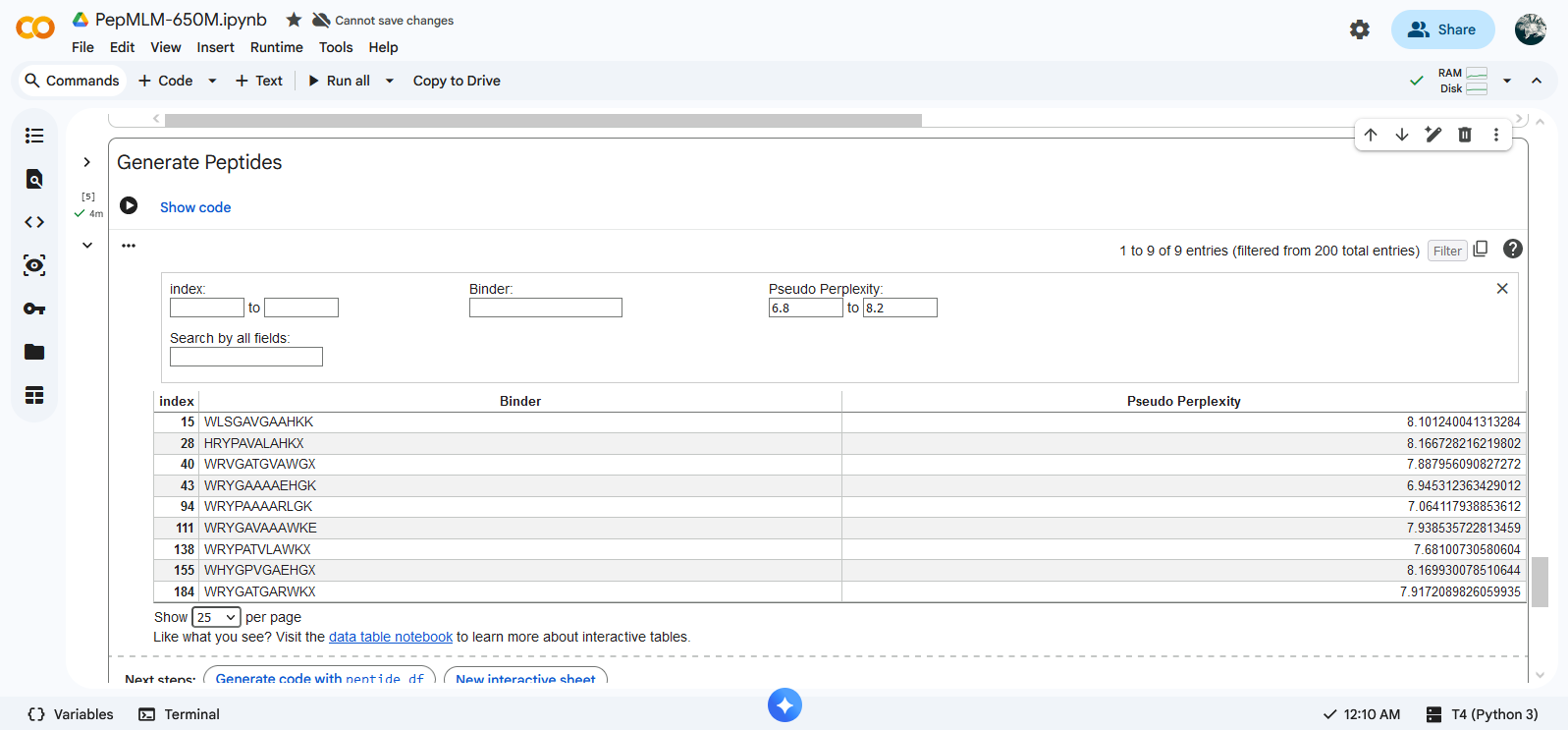
Approach 02: Selection of 4 peptides out of 200 generated (length 12 aa, top K 3)
| Index | Peptides | Pseudo Perplexity |
|---|---|---|
| 47 | WRYPAAAVALKX | 5.951070210421078 |
| 135 | WRSPAAVAAHKX | 4.819682049260351 |
| 147 | WHSGVVALAHKX | 6.103797038822761 |
| 199 | WRYGAVAARLKX | 5.926568160063284 |
For comparison, known peptide binder FLYRWLPSRRGG:
| Index | Binder | Pseudo Perplexity |
|---|---|---|
| N/A | FLYRWLPSRRGG | 20.63523127283615 |
Pseudo Perplexity measures how confident the model is in the sequence. A lower PPL (≈ 3–6) means that the sequence fits welllearned binding patterns and thus the model is confident that the peptide will bind the target. A higher PPL (> 10–15) means that the sequence looks different than the learned patterns and thus, the model is less confident about the probability for the peptide to bind the target.
In theory, the high value of the pseudo perplexity score (>15) of the known binder FLYRWLPSRRGG can sound surprising. But in reality, the authors of the reference paper showed that known binders can have relatively high perplexity values. While the confidence scores can be useful for ranking candidates, they have their own limitations: the score evaluates familiarity, not whether the peptide truly binds the protein of interest. See documentation for more details on outliers.
UPDATE: AlphaFold basic functions don’t allow the integration of undefined amino acids (X) so a new series of peptides was generated with Colab.
| Reference | Peptides | Pseudo Perplexity |
|---|---|---|
| Binder | FLYRWLPSRRGG | 20.64 |
| 0 | WLSPVVAAEHKE | 15.49 |
| 1 | WRYGAAAAEHKE | 9.10 |
| 3 | HHSYAAAVALKK | 12.67 |
| 15 | WLSGAVGAAHKK | 8.10 |
| 43 | WRYGAAAAEHGK | 6.95 |
| 94 | WRYPAAAARLGK | 7.06 |
| 111 | WRYGAVAAAWKE | 7.94 |
AlphaFold Visualization:
Except for peptides 3 and 43 which form an alpha helice secondary structure, all peptides present an unfolded structure that can appear more or less floppy or following the shape of the protein. None of the peptides are buried in the mutant SOD1: if binding to the protein, this appear to be a surface-bound for all of them. None of the peptide appear to interact with the N-terminal nor the dimer interface. All peptides seem to interact, at least partially with the beta-barrel. Beside the known binder, peptides 43, 94 and 111 appear to also interact with the electrostatic loop (loop containing a small portion of alpha helice).
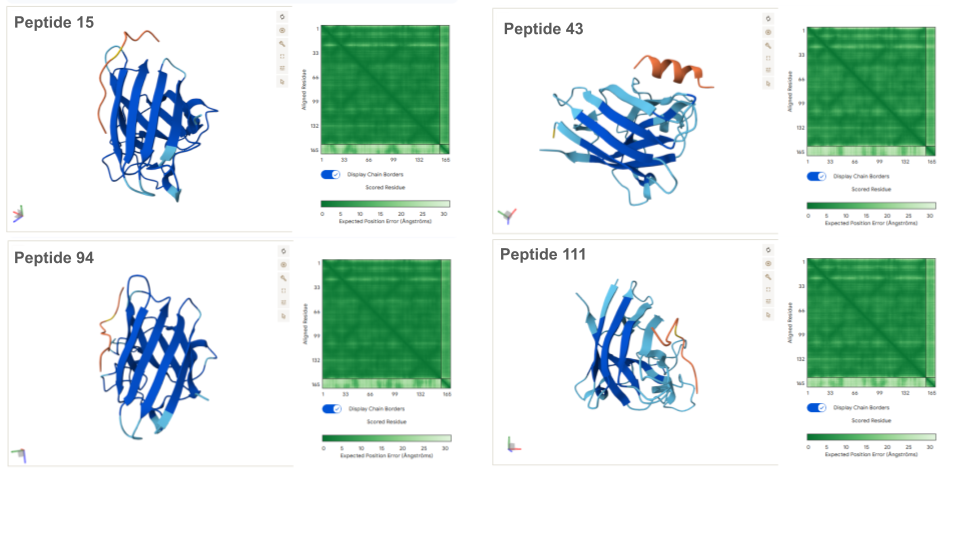
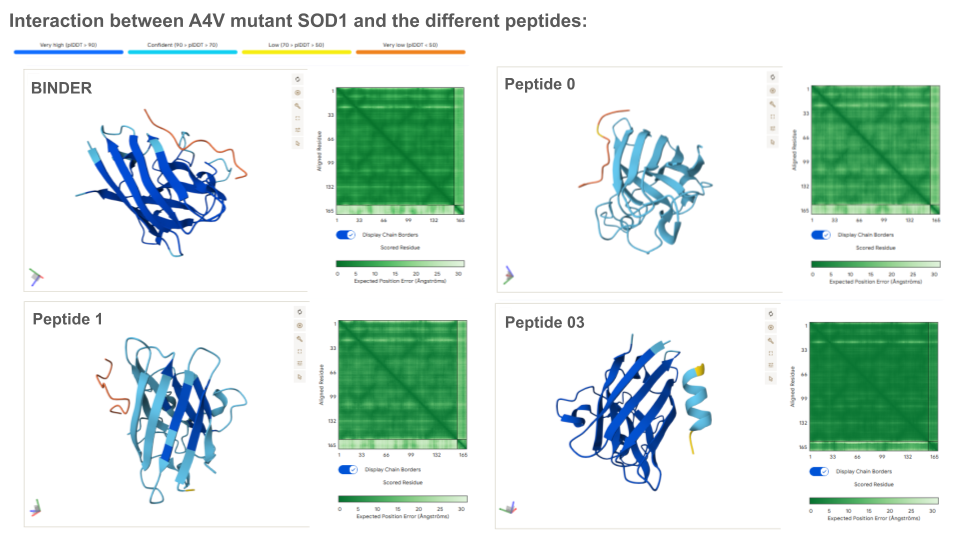



Prediction of the different A4V Mutant SOD1-peptide Bindings:
| Reference | ipTM |
|---|---|
| Binder | 0.3 |
| 0 | 0.27 |
| 1 | 0.28 |
| 3 | 0.73 |
| 15 | 0.46 |
| 43 | 0.35 |
| 94 | 0.42 |
| 111 | 0.31 |
ipTM indicates the probability of binding. Typically, if ipTM is >0.8, the probability of binding is high, and low if it’s <0.6. Thus, it is surprising that the binder only exhibits a ipTM of 0,3 with SOD1, which is either lower or similar to all generated peptides. This can maybe be explained by the fact that the peptide binds the surface vs being typically buried inside the protein, or/and because the peptide establish an usual bound with the electrostatic loop. With an ipTM = 0.73, peptide 3 exhibits the highest binding score: it might be interesting to predict its therapeutic property to evaluate if it is worth advancing toward therapy.
All peptides evaluated exhibit soluble and non-hemolytic properties.
Except for peptide 111, they all appear to bind only weakly to the mutant SOD1 target.
Interestingly, Peptiverse prediction about binding affinity:
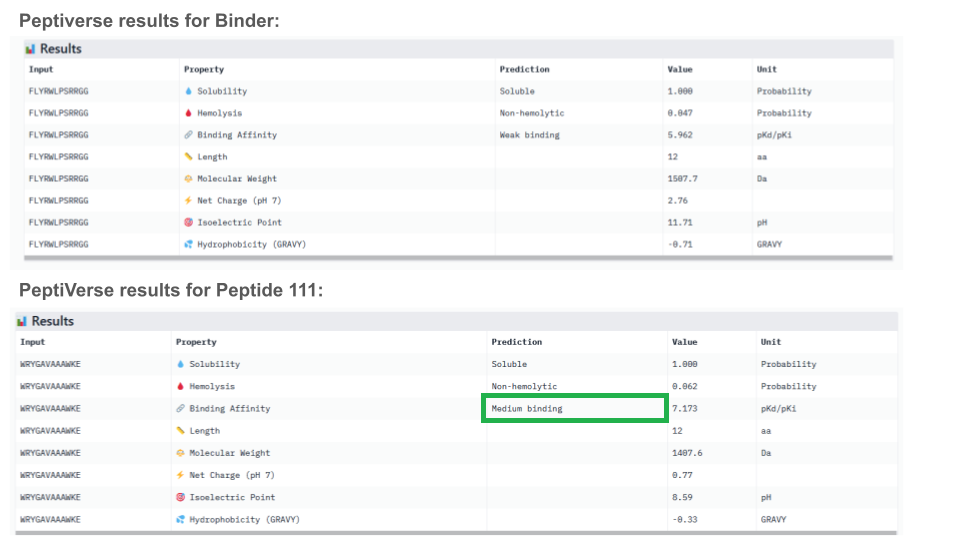

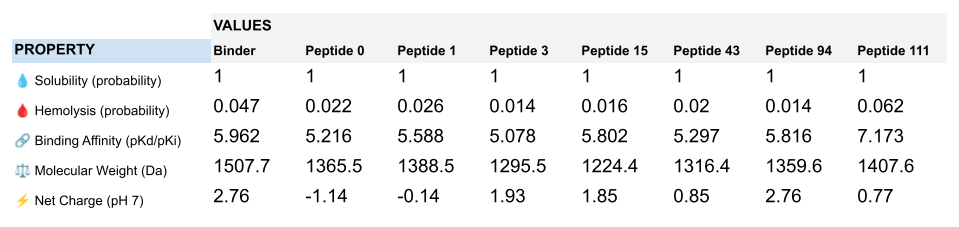

When considering how strongly the peptide binds to the target, peptide 111 seems to be the best candidate to proceed with. However, one can first question the discrepancy between the results obtained with the different prediction tools. And second, when checking more properties needed for the peptide to become a real drug, PeptiVerse indicates that this peptide presents a low membrane permeability and a fouling behavior. Thus, peptide 111 might not be the ideal candidate to be advanced for therapy. This result aligns with the purpose of PeptiVerse explained in the reference paper, which is to filter out bad candidates early and possibly, explains compromises that may have been made in the process: e.g. weaker binding property of the known SOD1 binder.
Peptide 111 extended analysis:
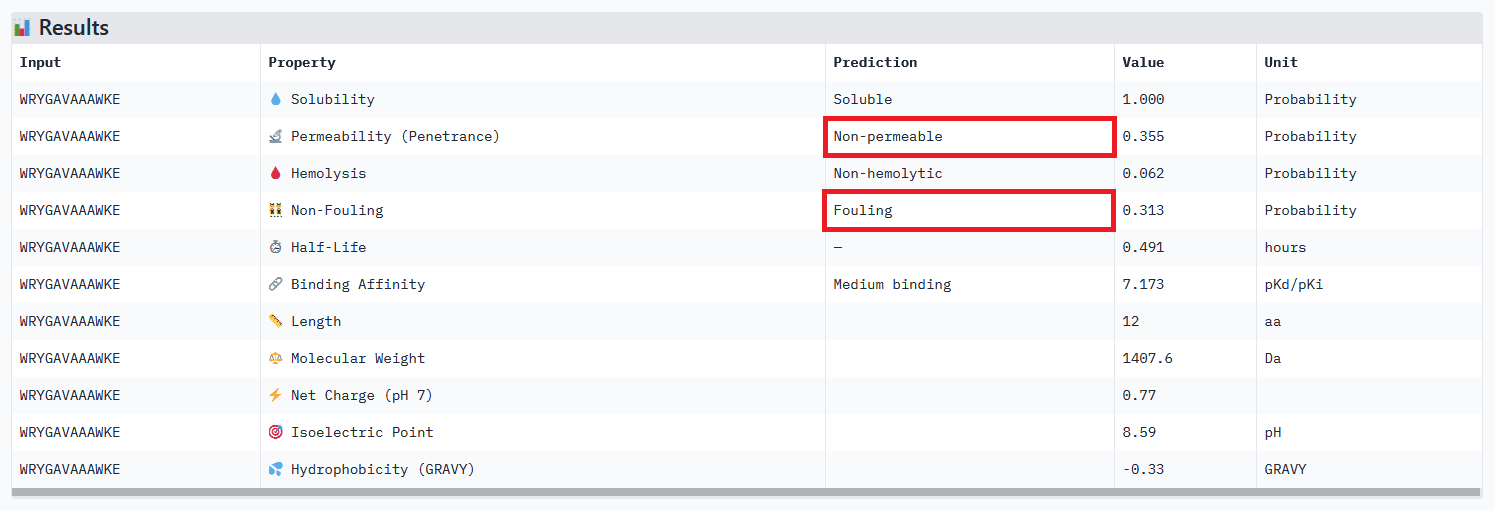

My strategy to find a potential candidate would be to (1) repeat the steps of peptides generation and select hundreds of candidates, (2) narrow down the selection by ensuring coherent results between the different prediction tools, checking all therapeutic properties and evaluating how the binding site may affect therapeutic efficacy (see next section).
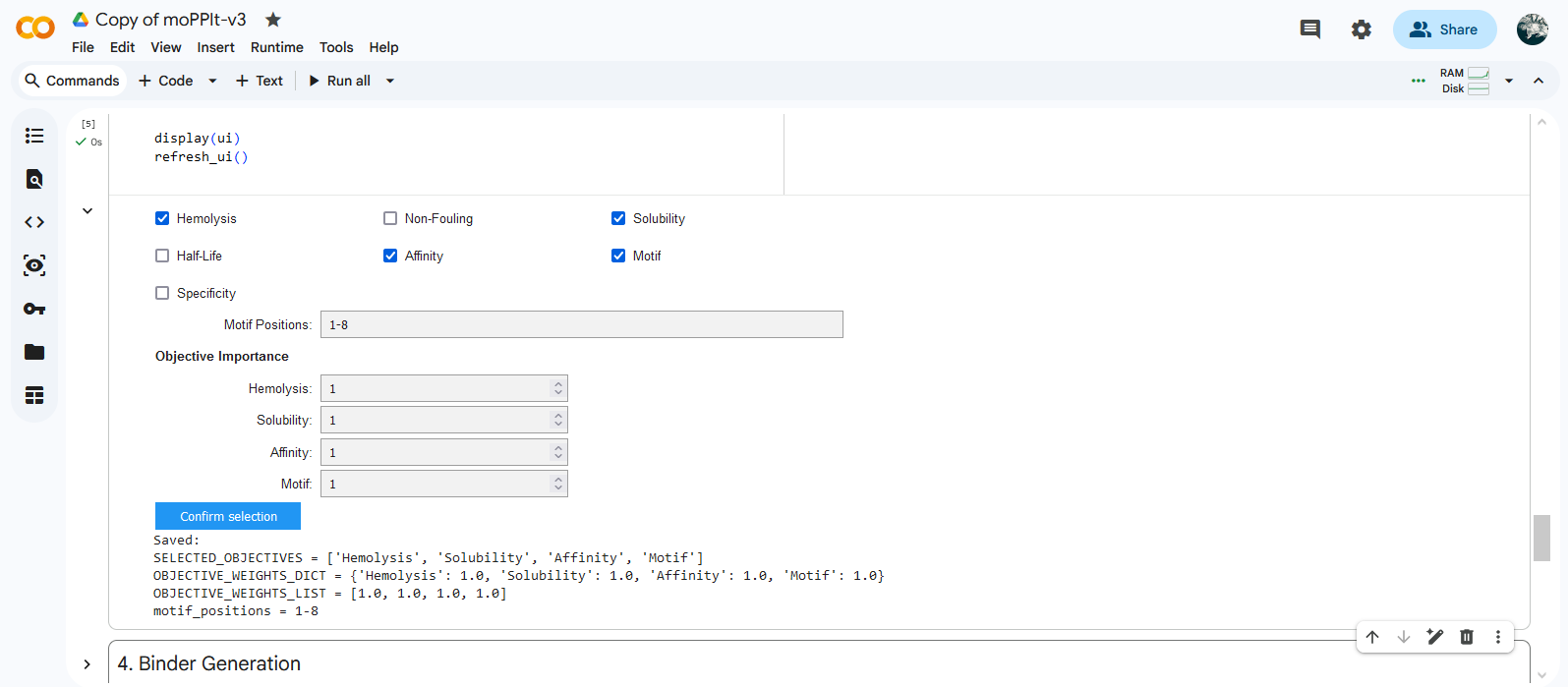
Colab moPPIt was used to generate peptides binding to a 8 aa length motif focused on the residue 4 corresponding to the mutation site (motif: 1-8, specificity: off). Affinity guidance, solubility and hemolysis guidance parameters were enabled.
Optimized Peptide 01: RKTTCQLTKEQG
Optimized Peptide 02: TEKSEEFKKKII
Optimized Peptide 03: RNETCVQKSKGF
moPPIt values:
| Binder | Hemolysis | Solubility | Affinity | Motif |
|---|---|---|---|---|
| RKTTCQLTKEQG | 0.97 | 0.92 | 5.82 | 0.88 |
| TEKSEEFKKKII | 0.98 | 0.75 | 5.67 | 0.42 |
| RNETCVQKSKGF | 0.97 | 0.83 | 6.70 | 0.88 |
Analysis of the optimized peptides in PeptiVerse
Interpretation:
Analysis of the optimized peptides in AlphaFold
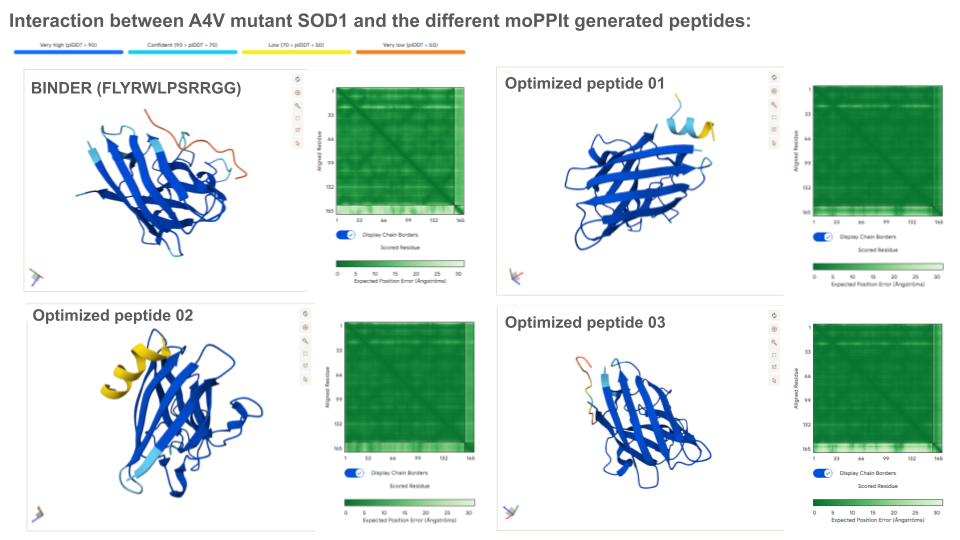

Interpretation:
CONCLUSION
While its binding affinity is evaluated as weak in PeptiVerse, 5.96 seems sufficient for it to bind to the target protein. Besides, the know binder exhibits all therapeutic characteristics needed: soluble, permeable, non-hemolytic and non-fouling.
EVALUATION OF THE KNOWN BINDER (FLYRWLPSRRGG) IN PEPTIVERSE:
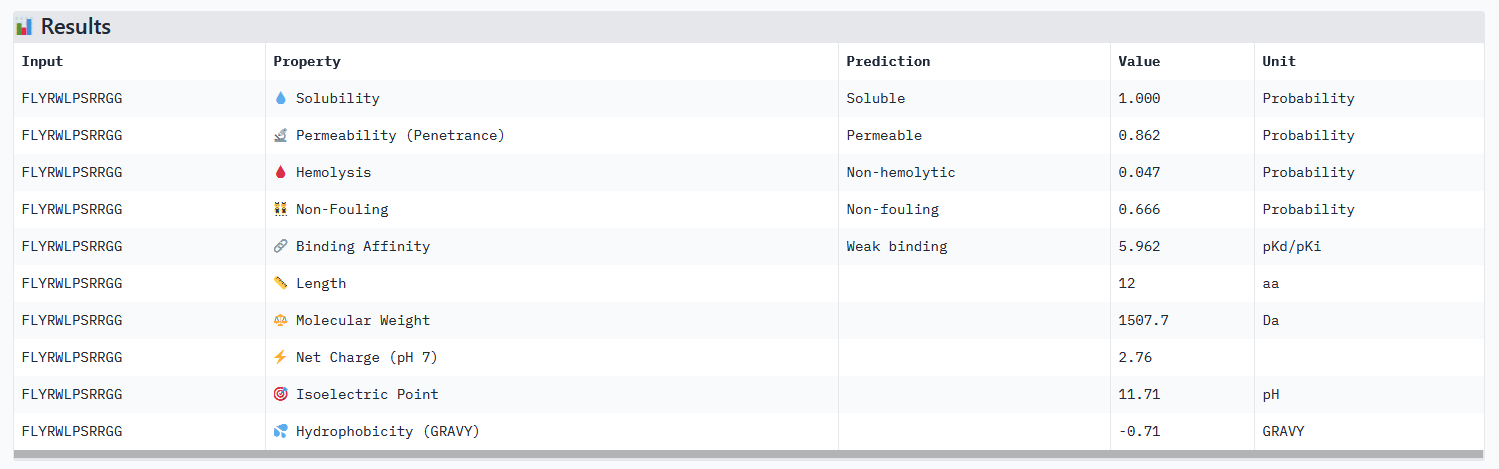

None of the generated peptides in this assignement combined all parameters at once. Concerning optimized peptide 01: a weaker permeability is not a deal breaker for potential therapeutic use, but given that the binding affinity isn’t strong I would exclude it from the list of potential therapeutic candidates for now. My strategy for the next steps would be to generate more peptides using moPPIt while testing different motifs: testing different length of the motif on a specific site (e.g. around the mutation site) and different site location. As a matter of fact, the electrostatic loop has been reported to be involved in the misfolding of SOD1 and the formation of toxic oligomers. Because it has a highly exposed surface and structural flexibility, this site is a motif top candidate (see documentation).
Source Protein Sequence: UniProt P00441
Simple explanations of the reference paper:
https://chatgpt.com/s/t_69b036610424819197b52653c5b55f6b
https://chatgpt.com/s/t_69b03ad48f748191aa89855f9d3830e4
https://chatgpt.com/s/t_69b03d68d6908191a0c3e4b251c01972
Strategies for creating peptides + explanation top K value:
https://chatgpt.com/s/t_69b0555069408191ba5e1253c8c345f2
https://chatgpt.com/s/t_69b05b9e75bc8191876d8d01f138fbf8
Perplexity score: https://chatgpt.com/s/t_69b05b9e75bc8191876d8d01f138fbf8
Creation of the peptides: Peptides were created using PepMLM Colab The slider functions were blocked so the parameters (length, top K and number of binders) were changed manually in the code. Peptides were selected according to perplexity values using the interactive table function in the Generate Peptide block.

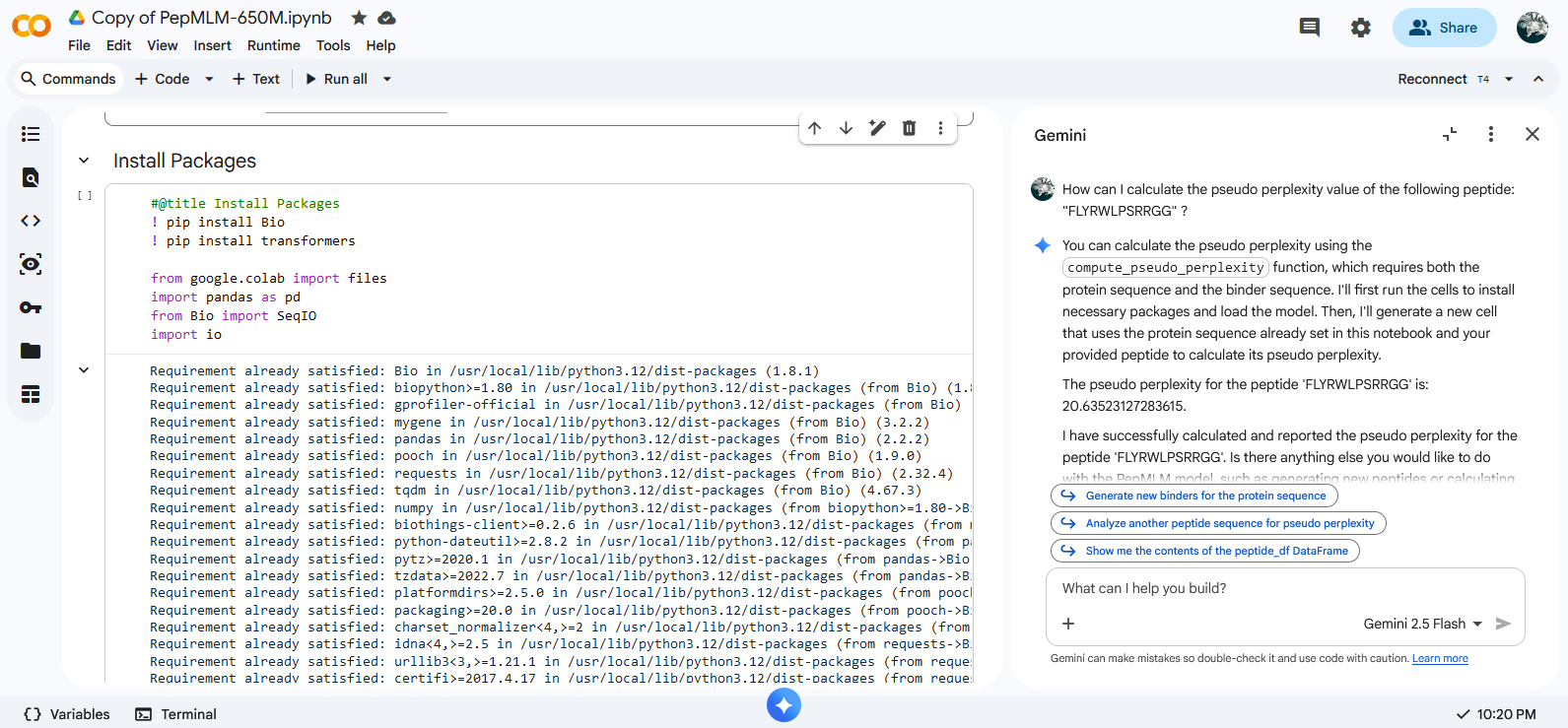
The pseudo perplexity value of the known SOD1-binding peptide FLYRWLPSRRGG was calculated by Gemini within Colab using compute_pseudo_perplexity function.

More about outliers binders: https://chatgpt.com/s/t_69b09f5842848191ad68e48392e7c919
Mutant A4V SOD1 sequence and the different peptide sequences were submitted as separate chains to model the protein-peptide complex in AlphaFold. SOD1 structure
Explanation parameters of the analysis results in AlphaFold:
https://chatgpt.com/s/t_69b1d7639604819191417d9ae3069f9f
https://chatgpt.com/s/t_69b486d55cf48191b082ed6264b81c3a
Data
Mutant SOD1 - Peptide 00 complex https://alphafoldserver.com/fold/43ecc9ff963fde21
Mutant SOD1 - Peptide 01 complex https://alphafoldserver.com/fold/86c286d21a2176a
Mutant SOD1 - Peptide 03 complex https://alphafoldserver.com/fold/58da5ac311db6c0a
Mutant SOD1 - Peptide 15 complex https://alphafoldserver.com/fold/19055fa61aaf71a5
Mutant SOD1 - Peptide 43 complex https://alphafoldserver.com/fold/556af6d37be44d8f
Mutant SOD1 - Peptide 94 complex https://alphafoldserver.com/fold/3dba002c05b228b1
Mutant SOD1 - Peptide 111 complex https://alphafoldserver.com/fold/4018c707b09f0f5a
Peptides were evaluated using the PeptiVerse platform.
Simple explanations of the reference paper: https://chatgpt.com/s/t_69b489a6aa048191bbbe0e45a0227899
Optimized peptides were generated using Colab moPPIt.
Simple explanations of the reference paper: https://chatgpt.com/s/t_69b53a1913b08191bbeffd3d085f973d
Run time: 30 min for 3 peptides (no access to A100 or L4 GPU).

Gemini coded an extra cell to download results.

Feedback ChatGPT on motif: https://chatgpt.com/s/t_69b7054bb99881918c31e0f77b6662d4
SKIP FOR NOW ⏱
TO BE UPDATED on March 31st.
The Phusion HF PCR Master Mix is a ready-to-use “all-in-one” cocktail designed for DNA amplification. It contains:
What needs to be added before running the PCR: template, primers and water.
Sources: New England Biolabs, Fisher Scientific
Primer annealing is a crucial PCR step during which primers bind to their complementary sequences on a single-stranded DNA template. This step occurs after the high-temperature denaturation phase, when the temperature is lowered to a specific point that allows stable hydrogen bonding between the primer and the template, providing a starting point for DNA synthesis by polymerase enzyme.
The annealing temperature (Ta) must be high enough to ensure specificity (binding only to the target) and low enough to allow binding to occur efficiently. Ta is determined from the melting temperature (Tm) of the primers, which is the temperature where half of the primer–DNA duplex separates and is usually set 3-5°C below the lower primer Tm.
Key factors that influence Ta include:
Sources: Qiagen, ThermoFisher
Basic Protocol: Polymerase Chain Reaction
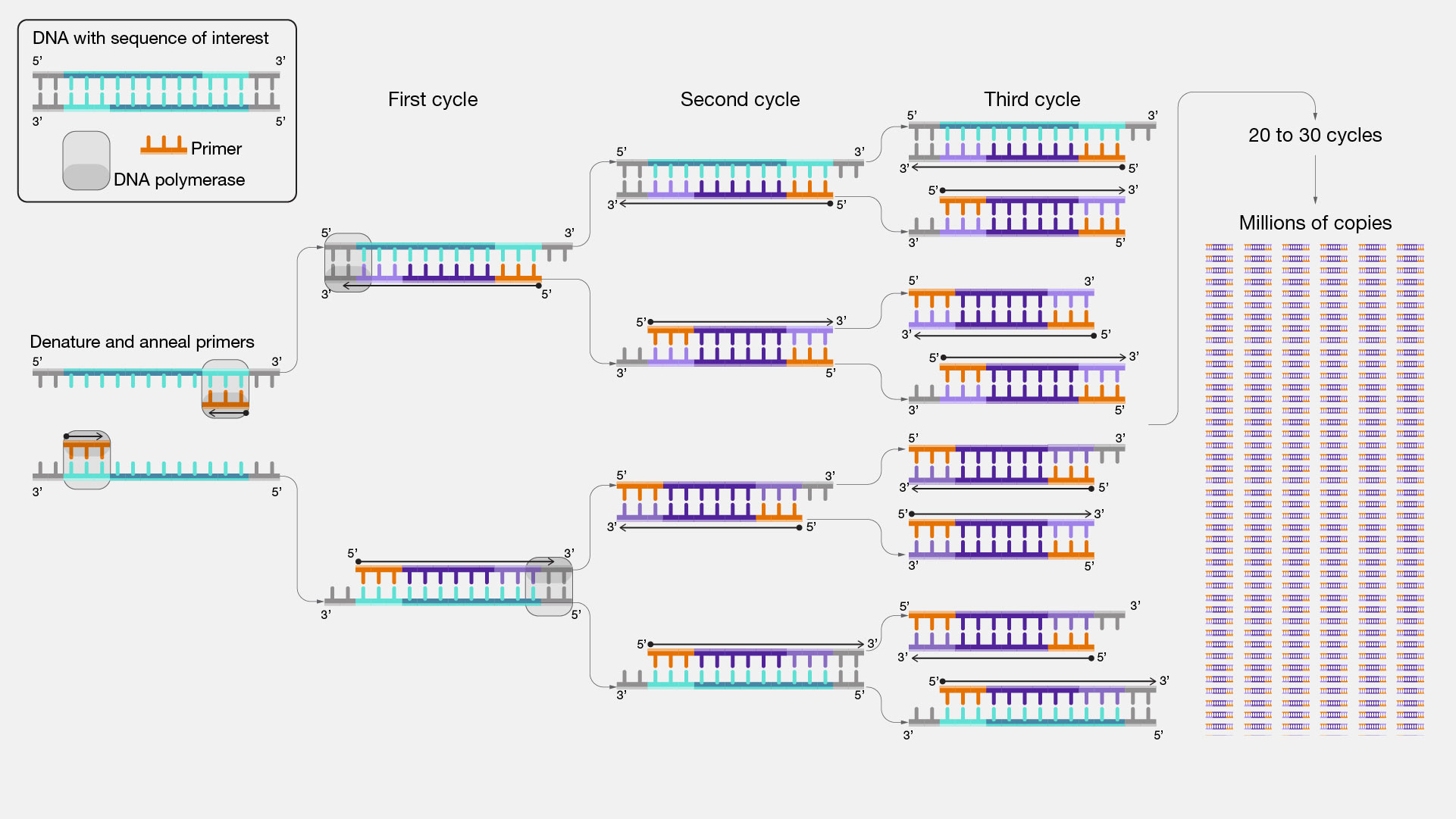
PCR creates millions of copies of a specific DNA fragment using primers(short DNA fragment) and a DNA polymerase enzyme. Key feature: The DNA fragments are defined by primer design.
Basic Protocol: Restriction Enzyme Digest

Restriction Enzyme Digest cuts DNA genome or plasmids at specific recognition sites. If a restriction enzyme cuts a plasmid, the circular DNA becomes linear. Key feature: The DNA fragments are defined by existing recognition sites in the DNA. ‘Sticky End’ vs ‘Blunt End’ Restriction Enzymes
Both methods produce linear DNA fragments, require a DNA template, use enzymes, often require gel electrophoresis afterwards to evaluate the size of the produced DNA fragments and are both commonly used in DNA cloning protocols.
However, the two methods are fundamentally different:
To make sure digested and PCR-amplified DNA fragments are suitable for Gibson Assembly, one mainly needs to verify that (1) the design of the overhangs is correct and compatible, and (2) make sure to use the best quality PCR products.
(1) Overhangs Design
Gibson Assembly relies on homologous overlaps between adjacent fragments. Each primers should present a 18–22 bp core binding region, and a ~20–40 bp overlap with the next fragment. One needs to make sure that these overlaps are identical in sequence and correctly oriented (5′→3′ continuity across fragments). Furthermore, the Tm should be similar across overlaps, and primers designed to avoid cross homology, secondary structures or strong hairpins (check lab protocol for guidelines).
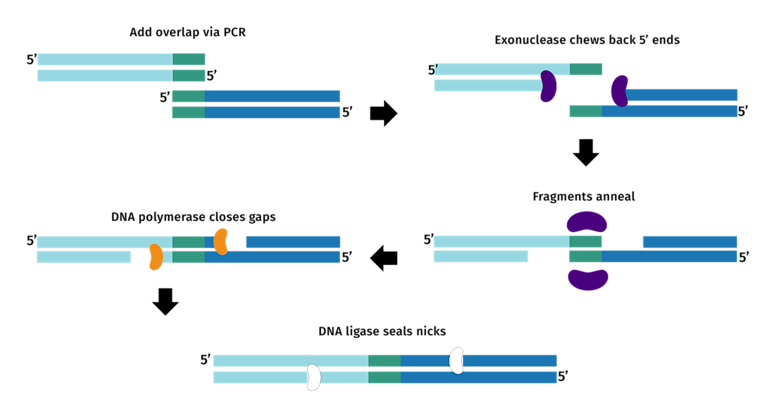
(2) Quality of the PCR Products
Template plasmid removal: the original DNA template needs to be destroyed using the enzyme Dpnl (recognition and cleavage of specific methylated sites present in the template).
DNA purification using the Zymo Research DNA Clean & Concentrator kit to remove unwanted components such as primers, enzymes, dNTPs. Contaminations such as salt and ethanol should also be avoided because Gibson enzymes (exonuclease, ligase) are very sensitive to contaminants.
(3) Verification Steps
Gel electrophoresis to confirm that the size of the fragments is correct: one clean band for the backbone and another clean one for the insert.
DNA quantification and fragment stoichiometry: one needs to make sure to respect 2:1 insert:backbone molar ratio to avoid empty plasmids or incorrect assemblies. DNA concentrations can be checked using Nanodrop/Qbit
Transformation refers to the process of introducing foreign genomic material into bacterial cells (remark: one talk about transfection when working with mammalian cells). Transformation can be induced either by heat shock (like in the present protocol) or by electroporation (electrical shock).
Chemical transformation workflow

According to Ron Weiss lecture, Intracellular Analog Neural Networks (IANNS) offer several advantages as compared with classical Boolean genetic circuits (combination of simple on/off switches):
Continuous signal: IANNs allow the integration of continuous signals: this processing is much closer to what happens in cell biology. For instance, a Boolean circuit can assess whether the protein CasE is present or not, while the IANNs can compute the concentration of CasE in the cell.
Non-linear computing: IANNs allow weighted summations and substractions through universal digital logic (AND + NOT gates). In other words, the computation allows some inputs to have more importance than others. This processing mimics the complexity of analog neural networks and by extension, also intracellular processes in a way which is closer than a circuit only made of on/off gates.
Programmability: Nodes can be composed into multi-layer networks (e.g. bandpass circuits). This feature allows IANNs to compute complex tasks with high efficiency, and for tuning the circuits without having to rethink everything.
The example of cancer cell classification given by Ron Weiss during his lecture is interesting for my final project: the same approach can be implemented to “assess the microbiotic landscape(s) of perimenopause”*. Biological states such as cancer or hormonal transitions don’t involve one but multiple parameters. IANNs can be used to track these changes with more precision and at a much earlier time point than a common single biomarker screening.
In Multi-input RNAi-based logic circuit for identification of specific cancer cells Weiss and al. use IANNs to determine whether a cell is cancerous or not. The authors use the levels of different types of microRNAs known to be involved in cancer as inputs (X1: biomarker 01, X2: biomarker 02, X3: biomarker 03). After integration of these inputs through the hidden layers of the circuit, a specific response is triggered depending on the classification of the cell. For instance, the “cancer cell” signature could correspond to intracellular biomarkers X1 being high, X2 high and X3 low. If the output equals 1 (or “true”) for “cancer cell”, the circuit triggers apoptosis (cell death) in these cells. If the miRNAs levels do not fit the cancer signature, nothing happens.
Possible design limitations:
Below is the diagram of the perceptron corresponding to the template given for the Neuromorphic Wizard software: it depicts a two-layers perceptron where the X1 input is DNA encoding for the CasE endoribonuclease (layer 1), the X2 input is DNA encoding for the Csy4 endoribonuclease (layer 2) and the bias output is the expression of the fluorescent protein mNeonGreen (mNG). Marker X1: eBFP2 ; Marker X2: mKO2 ; ERN01_rec_ERN02: endoribonuclease 01 regulates endoribonuclease 02; Y: output; Tx: transcription; Tl: translation.

Neuromorphic Wizard Template Build and Prediction:


Below is the diagram of the perceptron I submitted on Friday 20th. It depicts a circuit similar to the template but with a swap between X1 and X2: the X1 input is DNA encoding for the Csy4 endoribonuclease (layer 1) and the X2 input is DNA encoding for the CasE endoribonuclease (layer 2). Marker X1: mKO2 ; Marker X2: eBFP2 ; ERN01_rec_ERN02: endoribonuclease 01 regulates endoribonuclease 02; Y: output; Tx: transcription; Tl: translation.

Circuit 01 Build, Layout and Prediction:



Below is the diagram that depicts a circuit similar to the template but with one added layer in which X3 inputs an endoribonuclease (PgU) that regulates X2 output (Case). Marker X1: mKO2 ; Marker X2: eBFP2; Marker X3: mMaroon1 ; ERN01_rec_ERN02: endoribonuclease 01 regulates endoribonuclease 02; Y: output; Tx: transcription; Tl: translation.

Circuit 02 Build and Prediction:


The installation of the software was an assignment in itself and included watching again the BioClub recitation and a change of device. Reopening the software worked smoothly though. Note for later use:
At first, because a proper explanation of the building blocks was missing, it was difficult to understand the correspondance between the design of the perceptron and the Neuromorphic Wizard build. I worked directly on the software and exchanged a few blocks following an intuitive logic until I obtained a prediction instead of an error message. This led me to create and submit Circuit 01 on March 20th but I was missing the corresponding perceptron.
In a second step, after rewatching very carefully Ron Weiss’s lecture and the MIT recitation, I finally managed to gather enough elements to understand the building blocks and draw the diagrams corresponding to the builds of the template and Circuit 01. From there, I draw Circuit 02 and obtained the corresponding prediction in the Neuromorphic Wizard software. Total DNA concentration above 650 mg did not trigger an error message but changing the levels to stay under that threshold did.
Afterwards, I tried to incorporate a negative feedback loop into Circuit 01 : csy4_rec_Case but got a error message. Same when trying to translate my final project idea into a diagram.
Next steps:
Fungal materials are biomaterials made out of fungi, a kingdom of organisms that includes yeasts, molds and mushrooms. The fruiting body of Fomes fomentarius (aka “tinder fungus”) is known for its ancient use as a fire-starter and has been used for centuries to make amadou, a buckskin-like fabric. However, the traditional craft of amadou-making is slowly dying and the vast majority of fungal materials are now made of mycelium, the root-like part of mushrooms and molds that consist of a vast network of microscopic thread-like filaments.
Contemporary examples: Mycelium-based materials can be used to make textiles, packaging, isolation panels, furniture, building materials and even funerary vessels. Yeasts (unicellular micro-organisms classified as fungi) have also been reported to be used to create glue and other adhesive materials. And while they are less popular, fruity bodies are also used to make packaging, leather-like textiles and waterproof sealants.
Contemporary mycelium-based material designs

Mycelium-based materials serve as seemingly non-toxic, sustainable, biodegradable, and low-density alternatives to a wide array of synthetic and traditional materials such as plastic, polystyrene or even wood.
Composite-mycelium materials grow a network that binds to a matrix and can be molded: they basically assemble themselves using waste products. They represent an interesting alternative to packaging materials such as plastic or polystyrene.
Construction is currently one of the most polluting industry sectors. While fungal materials represent a more sustainable alternative than concrete, their low density can be an asset when used as isolation panels but a drawback for structures that require more mechanical stability.
Another weakness of mycelium-based materials is their lack of resistance to moisture. AI mycelium-based furniture and constructions look esthetically and conceptually attractive, but how will they age over time?
Fungi are eukaryotes, meaning their cellular machinery is more similar to plants and animals. Bacteria can be useful for the fast production of simple molecules but fungi allows the production of more complex proteins and in higher quantities.
As explained by Ren Ramlan during recitation, engineering fungi would help overcome some of these current limitations. For instance, introducing mutations into genes involved in the regulation of chitin, glucans or other proteins composing the cell walls of the hifae may lead to produce mycelium-based materials with stronger mechanical or waterproof properties. Being said, some conceptual paradoxes can also appear when thinking about engineering mycelium: if we design mycelium-based materials to be stronger, they become harder to break down and thus, less eco-friendly.
Schematic representation of mycelium physiology at different scales from Advanced Materials From Fungal Mycelium: Fabrication and Tuning of Physical Properties:

When it comes to scaling up mycelium-based production, scientists working in the industrial sector may want to engineer mycelium to achieve better control over material production and quality (e.g. reduce contamination risks). Whether mass production can ever be truly sustainable without changing our consumption practices, however, remains an open question.
Period poverty affects over 500 million people world-wide and leads to missed education/work and health risks from using improper materials. And when available, single-use menstrual products create significant environmental damages including plastic pollution and an annual release of hundreds of thousands of tons of waste.
If invited to create a mycelium-based product, I would first aim to evaluate whether it is possible or not to create basic and safe DIY mycelium-based menstrual pads.
In a second step, I would design and engineering a mycelium-based tampon that presents reduced risk for toxic shock syndrome. I would try to stay as close as possible to the natural material, but explore how to engineer the fibers so that they can capture and possibly digest or neutralize toxins produced by bacterial strains such as Staphylococcus aureus or Streptococcus pyogenes.
The World’s Last Amadou Makers
Mycelium: Rethinking Materials in Design
Breaking Down the Barriers: The Future of Mycelium-Based Materials
Period Poverty and Stigma: A Matter of Human Rights
Toxic Shock Syndrome: A Literature Review
Engineering yeast for bioplastic production
Mycelium-based composites: An updated comprehensive overview
Reference paper: Cell-Free Gene Expression: Methods and Applications
According to the course material, CFPS systems offers more flexibility because there is no need anymore to maintain cells alive. For instance, it allows the production of toxic proteins that would otherwise kill the cells producing them. Using easier cell machinery systems can also give scientists more flexibility in adapting their experimental design. For instance, it allows them to incorporate non-natural amino acids into the reaction. And finally, CFPS systems also offer much easier storage conditions because it is possible to freeze-dry the reactions on tiny pellets.
Cell-free protein synthesis also offers more control during production: there is no unknown cellular process occurring in parallel, nor interference with other proteins naturally produced by the cells. The conditions of the environment (e.g. ions concentrations, pH, enzymes etc.) can be defined more precisely. Finally, it makes it much easier to isolate the target protein. These assets are useful in the pharmaceutical industry for producing large batches of hormones (e.g. insulin), vaccines or medicines.
To go further…
A critical comparison of cellular and cell-free bioproduction systems
Cell-Free Synthesis: Expediting Biomanufacturing of Chemical and Biological Molecules
Components of a CFPS reaction mixed in a test tube:

The CFPS system mixture contains:
The translation and transcription processes require a lot of energy. Without a continuous supply in ATP, the protein synthesis quickly shuts down. Thus, energy regeneration is needed to provide a continuous supply of energy and avoid the accumulation of by-products (e.g. phosphate) that can interfere with the protein synthesis.
There are many different ways to regenerate energy in cell-free systems: see references. Choosing glucose as an energy source can be interesting in the prototyping phase of a project because it is a highly cost-effective and an efficient way to ensure a continuous supply of ATP via glycolysis in both E. coli and yeast extracts. It can also be combine with creatine phosphate in a dual system: creatine phosphate acts as a high-energy phosphate donor for rapid ATP regeneration via creatine kinase, while glucose feeds the glycolytic pathway to produce ATP and consume inorganic phosphate.

From The cost-efficiency realization in the Escherichia coli-based cell-free protein synthesis systems
References
ATP Regeneration from Pyruvate in the PURE System
Cell-Free PURE System: Evolution and Achievements
Development of prokaryotic cell-free systems for synthetic biology (includes PANOx energy regeneration system)
Prokaryotic cell-free systems are fast and efficient for producing simple proteins. For instance, E.coli systems are ideal for producing simple proteins such as Green Fluorescence Protein (GFP) in labs and in the industry.
On the other hand, eukaryotic systems are slower and more expensive, but they can be used for the production of proteins which require complex folding, disulfides bridges, and post-translational modifications (e.g. glycosylation, lipidation) such human anti-bodies.
Reference
Protein Synthesis in Prokaryotes vs. Eukaryotes: What’s the Difference?
Cell-Free Protein Synthesis: Pros and Cons of Prokaryotic and Eukaryotic Systems
Membrane proteins are composed of transmembrane domains which makes them hydrophobic and thus hard to express in traditional cell-based systems: they need a special environment to avoid aggregation, and enable the proteins to fold and function correctly. However, it is possible to recreate such membrane-like environments by adding specific supplements to the reaction depending on the protein (Is it small and simple, or bigger and complex?) and what we want to do with it (do we only want to extract the protein or does it have to be functional?).
Cell-free systems derived from prokaryotic, as well as eukaryotic extracts that lack endogenous microsomes (natural membrane fragments from the cells) be can be supplemented with:

From Cell-free Membrane Protein Synthesis
For my final project “Sensing perimenopause: a bioluminescent art installation”, one design strategy is to use the G protein–coupled receptor to elicit a bioluminescent response to an environmental change, e.g. a change in extracellular levels of glycogen in the vaginal secretions. If using a cell-free system, I might want to use eukaryotic cell extracts and add liposomes to ensure that the receptor can sense the glycogen levels in real-time.
To go further…
Cell-Free Protein Synthesis: A Promising Option for Future Drug Development
Membrane protein synthesis in cell-free systems: From bio-mimetic systems to bio-membranes
Membrane protein production in Escherichia coli cell-free lysates
Membrane protein synthesis in cell-free systems: From bio-mimetic systems to bio-membranes
Cell-free synthesis of membrane proteins: Tailored cell models out of microsomes
Achieving optimal protein yield is a major challenge in cell-free protein synthesis systems. Below is a table that lists three common issues that can lead to low protein production and how to solve them.
| Issue | Troubleshoot |
|---|---|
| Inadequate design (Transcription/Translation) | Adapt expression system (e.g. switch to eukaryotic for complex proteins) Codon optimization * Check DNA design (e.g. plasmid sequence, promoter strength) * Adapt temperature and ions concentration (possibly run a screening test to find the optimal conditions) * Energy depletion: adapt energy regeneration system |
| Misfolding/aggregation | Adapt expression system (e.g. switch to rabbit reticulocyte or wheat germ) * Add chaperones * Adjust temperature and chemical conditions * Use solubility-enhancing tags and supplement with solubilizing agents to avoid aggregation |
| Degradation/purity | mRNA degradation: lower temperature to 20C to slow down phage polymerase * Protein degradation: add protease inhibitors * Lysis: check lysis time, temperature and buffer composition * Purification: check affinity tags, resin compatibility and resin amount if using column * Elution: check buffer pH, concentration of eluting agent and possibly increase incubation time |
References
How to Troubleshoot Low Protein Yield After Elution
Solved: Low Yields in Cell-Free Protein Synthesis
Troubleshooting Protein Folding Issues in Cell-Free Synthesis: Tips from Industry Experts
TO BE ADAPTED/COMPLETED ON APRIL 19th Below is only a first step using AI generated answer (Gemini). In progress: currently checking the litterature and adapting my answer according to my findings
Biosensor types To be confirmed after for class on biosensors
GLYCOGEN measurements Membrane proteins that can detect glycogen levels in the environment or on the cell surface are generally designed as engineered biosensors, as natural extracellular glycogen sensors are rare. Key proteins and strategies identified in research include:
Membrane protein-based biosensors https://pmc.ncbi.nlm.nih.gov/articles/PMC5938585/
A generic method for fluorescence monitoring glycogen through patent blue V triggered supramolecular switching https://www.sciencedirect.com/science/article/abs/pii/S0925400522002726
A red fluorescent genetically encoded biosensor for in vivo imaging of extracellular l-lactate dynamics https://www.nature.com/articles/s41467-025-64484-x
Protein Targeting to Glycogen (PTG): A Promising Player in Glucose and Lipid Metabolism https://www.mdpi.com/2218-273X/12/12/1755
A yeast FRET biosensor enlightens cAMP signaling https://www.molbiolcell.org/doi/10.1091/mbc.E20-05-0319
Novel method for detection of glycogen in cells https://pmc.ncbi.nlm.nih.gov/articles/PMC5444244/
In vivo biochemistry: Applications for small molecule biosensors in plant biology https://pmc.ncbi.nlm.nih.gov/articles/PMC3679211/
ACID LACTIC measurements (Next step)
BIOLUMINESCENCE
Membrane proteins play a crucial role in bioluminescence, acting as transporters for substrates, anchoring luciferases for localized signaling, or as part of energy transfer complexe.
Key membrane proteins and related mechanisms include:
In marine organisms, these membrane proteins are essential for the high-intensity, controlled flashes of light often seen in deep-sea creatures, where localization of the reaction to the membrane maximizes light output efficiency.
Bioluminescent and Fluorescent Proteins: Molecular Mechanisms and Modern Applications https://pmc.ncbi.nlm.nih.gov/articles/PMC9820413/
Bioluminescence Assay for Detecting Cell Surface Membrane Protein Expression https://pmc.ncbi.nlm.nih.gov/articles/PMC3064531/
Reference eGFP sequence:
MVSKGEELFTG VVPILVELDG DVNGHKFSVS GEGEGDATYG KLTLKFICTT GKLPVPWPTL VTTLTYGVQC FSRYPDHMKQ HDFFKSAMPE GYVQERTIFF KDDGNYKTRA EVKFEGDTLV NRIELKGIDF KEDGNILGHK LEYNYNSHNV YIMADKQKNG IKVNFKIRHN IEDGSVQLAD HYQQNTPIGD GPVLLPDNHY LSTQSALSKD PNEKRDHMVL LEFVTAAGIT LGMDELYKLE HHHHHH
According to Gemini:
Schematic diagram of the chromophore formation in maturing eGFP:

Source image: What is the maturation time for fluorescent proteins?
Calculation
When entering the full sequence in the online calculator one obtains a theoretical general molecular weight of ~ 28.006 kDa (“Th.Av. Mw” = 28006.60 Da)

Integration of the fluorophore maturation: Th.Av. Mw - (H2O Mw + H2 Mw) = 28006.60 - (18.02 + 2) = 28006.60 - 20.02 = 27986.58 Da
In conclusion, the molecular weight of eGFP is estimated to 27.987 kDa.

Contribution
Impressions
A fun, satisfying, and addictive project that also reveals some very interesting layers.
My experience felt more competitive than collaborative, with a general tendency to push through one’s own idea vs. letting things develop collectively into a creative direction (e.g. exquisite corpse concept).
Direct communication with my “competitors” (via DMs on the forum) helped orientate the creative flux but despite the frustration, I found the dynamic of the “territorial tensions” more interesting.
The experience made me curious about how the collected data might inform topics such as patterns (spacial colonization, spontaneously emerging pixel creativity…), competitive dynamics, and flocking behaviors. It also made me wonder whether - and how - collective intelligence can manifest in a society which is more driven by productivity than observation.
https://forum.htgaa.org/t/global-pixel-artwork-cooperation-guidelines/559/16?u=2026a-flo-razoux