WEEK 02


(c) Flo Razoux, AI generated


(c) Flo Razoux, AI generated
Part 1: Benchling & In-silico Gel Art
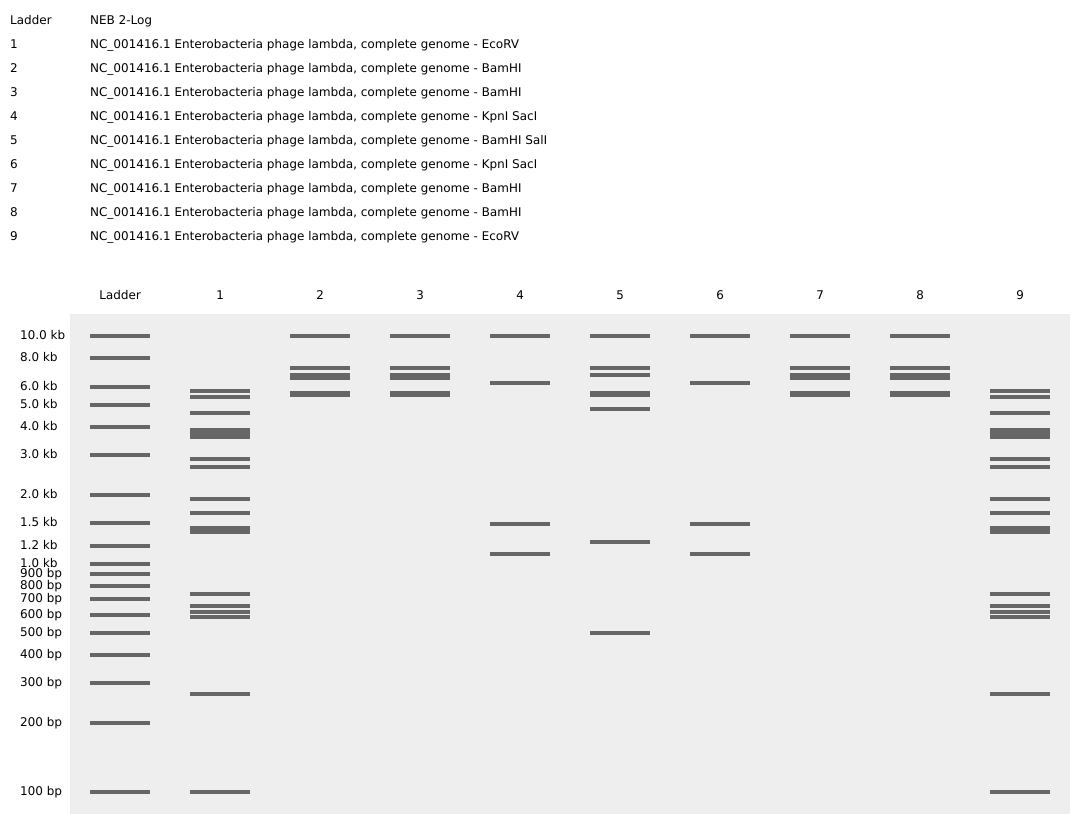

Part 2: Gel Art - Restriction Digests and Gel Electrophoresis
No access to lab
Part 3: DNA Design Challenge
3.1. LUCIFERASE
Bioluminescent art is based on the organic production of visible light by living organisms. This light is produced through the oxidation of luciferin, which is catalyzed by an enzyme called luciferase. For this week’s homework, we will focus on the protein coding for this enzyme, which was first identified in the firefly species (Photinus Pyrlis) [1].
[1] De Wet J.R. et al. Firefly luciferase gene: structure and expression in mammalian cells. Mol Cell Biol (1987). https://pmc.ncbi.nlm.nih.gov/articles/PMC365129/
Sources
NCBI database search: https://www.ncbi.nlm.nih.gov/protein/BAF48396.1
Uniprot database search: https://www.uniprot.org/uniprotkb/P08659/entry#sequences
Protein Sequence
550 amino acids
1 medaknikkg papfypledg tageqlhkam kryalvpgti aftdahievn ityaeyfems
61 vrlaeamkry glntnhrivv csenslqffm pvlgalfigv avapandiyn erellnsmni
121 sqptvvfvsk kglqkilnvq kklpiiqkii imdsktdyqg fqsmytfvts hlppgfneyd
181 fvpesfdrdk tialimnssg stglpkgval phrtacvrfs hardpifgnq iipdtailsv
241 vpfhhgfgmf ttlgylicgf rvvlmyrfee elflrslqdy kiqsallvpt lfsffakstl
301 idkydlsnlh eiasggapls kevgeavakr fhlpgirqgy gltettsail itpegddkpg
361 avgkvvpffe akvvdldtgk tlgvnqrgel cvrgpmimsg yvnnpeatna lidkdgwlhs
421 gdlaywdede hffivgrlks likykgyqva paelesillq hpnifdagva glpdddagel
481 paavvvlehg ktmtekeivd yvasqvttak klrggvvfvd evpkgltgkr darkireili
541 kakkggkskl
3.2. Reverse Translate: Protein sequence to DNA sequence
Source
NCBI database search for P.pyralis (firefly) luciferase gene: https://www.ncbi.nlm.nih.gov/nuccore/M15077
Luciferase protein DNA sequence
1 ctgcagaaat aactaggtac taagcccgtt tgtgaaaagt ggccaaaccc ataaatttgg
61 caattacaat aaagaagcta aaattgtggt caaactcaca aacattttta ttatatacat
121 tttagtagct gatgcttata aaagcaatat ttaaatcgta aacaacaaat aaaataaaat
181 ttaaacgatg tgattaagag ccaaaggtcc tctagaaaaa ggtatttaag caacggaatt
241 cctttgtgtt acattcttga atgtcgctcg cagtgacatt agcattccgg tactgttggt
301 aaaatggaag acgccaaaaa cataaagaaa ggcccggcgc cattctatcc tctagaggat
361 ggaaccgctg gagagcaact gcataaggct atgaagagat acgccctggt tcctggaaca
421 attgcttttg tgagtatttc tgtctgattt ctttcgagtt aacgaaatgt tcttatgttt
481 ctttagacag atgcacatat cgaggtgaac atcacgtacg cggaatactt cgaaatgtcc
541 gttcggttgg cagaagctat gaaacgatat gggctgaata caaatcacag aatcgtcgta
601 tgcagtgaaa actctcttca attctttatg ccggtgttgg gcgcgttatt tatcggagtt
661 gcagttgcgc ccgcgaacga catttataat gaacgtaagc accctcgcca tcagaccaaa
721 gggaatgacg tatttaattt ttaaggtgaa ttgctcaaca gtatgaacat ttcgcagcct
781 accgtagtgt ttgtttccaa aaaggggttg caaaaaattt tgaacgtgca aaaaaaatta
841 ccaataatcc agaaaattat tatcatggat tctaaaacgg attaccaggg atttcagtcg
901 atgtacacgt tcgtcacatc tcatctacct cccggtttta atgaatacga ttttgtacca
961 gagtcctttg atcgtgacaa aacaattgca ctgataatga attcctctgg atctactggg
1021 ttacctaagg gtgtggccct tccgcataga actgcctgcg tcagattctc gcatgccagg
1081 tatgtcgtat aacaagagat taagtaatgt tgctacacac attgtagaga tcctattttt
1141 ggcaatcaaa tcattccgga tactgcgatt ttaagtgttg ttccattcca tcacggtttt
1201 ggaatgttta ctacactcgg atatttgata tgtggatttc gagtcgtctt aatgtataga
1261 tttgaagaag agctgttttt acgatccctt caggattaca aaattcaaag tgcgttgcta
1321 gtaccaaccc tattttcatt cttcgccaaa agcactctga ttgacaaata cgatttatct
1381 aatttacacg aaattgcttc tgggggcgca cctctttcga aagaagtcgg ggaagcggtt
1441 gcaaaacggt gagttaagcg cattgctagt atttcaaggc tctaaaacgg cgcgtagctt
1501 ccatcttcca gggatacgac aaggatatgg gctcactgag actacatcag ctattctgat
1561 tacacccgag ggggatgata aaccgggcgc ggtcggtaaa gttgttccat tttttgaagc
1621 gaaggttgtg gatctggata ccgggaaaac gctgggcgtt aatcagagag gcgaattatg
1681 tgtcagagga cctatgatta tgtccggtta tgtaaacaat ccggaagcga ccaacgcctt
1741 gattgacaag gatggatggc tacattctgg agacatagct tactgggacg aagacgaaca
1801 cttcttcata gttgaccgct tgaagtcttt aattaaatac aaaggatatc aggtaatgaa
1861 gatttttaca tgcacacacg ctacaatacc tgtaggtggc ccccgctgaa ttggaatcga
1921 tattgttaca acaccccaac atcttcgacg cgggcgtggc aggtcttccc gacgatgacg
1981 ccggtgaact tcccgccgcc gttgttgttt tggagcacgg aaagacgatg acggaaaaag
2041 agatcgtgga ttacgtcgcc agtaaatgaa ttcgttttac gttactcgta ctacaattct
2101 tttcataggt caagtaacaa ccgcgaaaaa gttgcgcgga ggagttgtgt ttgtggacga
2161 agtaccgaaa ggtcttaccg gaaaactcga cgcaagaaaa atcagagaga tcctcataaa
2221 ggccaagaag ggcggaaagt ccaaattgta aaatgtaact gtattcagcg atgacgaaat
2281 tcttagctat tgtaatatta tatgcaaatt gatgaatggt aattttgtaa ttgtgggtca
2341 ctgtactatt ttaacgaata ataaaatcag gtataggtaa ctaaaaa
3.3. Codon optimization
According to the genetic code, there are fewer amino acids than codon possibilities (see chart below, image credit cdn.prod.website-files.com). Thus, in theory, multiple codons can encode for the same amino acid. But in practice, spatial configuration and kinetics factors affect the translation process. For instance, the use of some codons ressembling the STOP codons can interrupt prematurely the translation process. Thus, codon optimization is an important step when designing a nucleotide sequence.
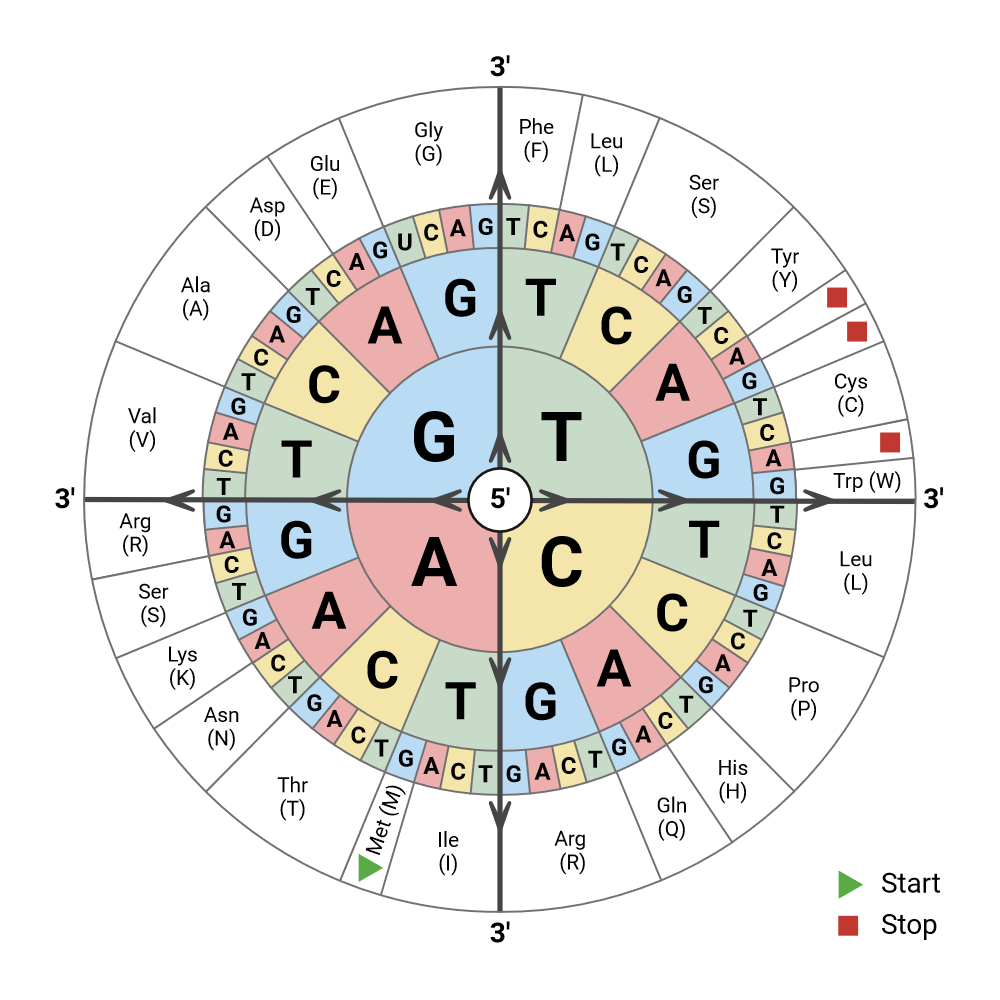

Luciferas protein DNA sequence with Codon-Optimization
TO BE COMPLETED ON MARCH 31st.