Protein Visualisation
1. PROTEIN OF INTEREST: FIREFLY LUCIFERASE
For this week’s assignments, we will keep focusing on the first final project idea of creating bioluminescent menstrual blood. While it would be interesting to have a closer look at the structure of hemoglobin, the protein that facilitates the transportation of oxygen in erythrocytes (red blood cells) and gives its red color to human blood in visible light, we will keep studying the luciferin 4-monooxygenase. This enzyme, commonly known as firefly luciferase, catalyses the production of light through the oxidation of luciferin. The structure of this protein is simpler than hemoglobin, so this seems to be an ideal option to apply what has been covered in class this week. We might go back to hemoglobin, myoglobin and other proteins determining blood color later depending on how the project develops.
2. IDENTIFICATION OF AMINO ACIDS SEQUENCE
Length: 550 amino acids
Most frequent amino acid: Leucine
Homology: Luciferase has 250 homologs across insects (endopterygota, 95% ; polyneoptera 3%; paraneoptera %) and bacteria (allobacillus, 1%). VISUALISATION
Family: Protein families refers to groups of closely related proteins with high sequence/functional similarity and common ancestry. The firefly luciferase belongs to the acyl-adenylate/thioester-forming superfamily of enzymes, also known as the ANL superfamily or the ATP-dependent AMP-binding enzyme family.
Documentation Sequence Length and Amino Acids Frequency
AA sequence obtained from NCBI and Uniprot database (see WEEK 02 HW)
Colab code used: https://colab.research.google.com/drive/1vlAU_Y84lb04e4Nnaf1axU8nQA6_QBP1
Output: Sequence Length: 550 Amino Acid Frequencies: l: 52 g: 46 v: 44 a: 42 k: 40 i: 37 e: 33 d: 30 f: 30 p: 29 t: 29 s: 29 r: 21 n: 19 y: 19 q: 16 m: 14 h: 14 c: 4 w: 2
Documentation Homology Calculation runned with BLAST: FULL REPORT
3. FIREFLY LUCIFERASE STRUCTURE
RCSB page: https://www.rcsb.org/structure/1LCI
Structure Deposition Date: 1996-06-01
Crystal structure of firefly luciferase throws light on a superfamily of adenylate-forming enzymes
Resolution: 2.00 - 2.20 Å
The resolution is <2.70 Å and therefore considered as good.
Additional molecule(s) in the representation: none
Here is a page which presents the structure of the luciferase complexed with oxyluciferin and AMP (products of the reaction catalyzed by the luciferase): https://www.rcsb.org/structure/2D1R
Associated protein families:
- AMP-binding enzyme-like
- AMP-binding enzyme C-terminal domain-like They are both superfamilies too
Documentation Family Attribution
Calculation with SCOP
Output SCOP search for P08659 uniprot ID: 4 domains: 8025243 1BA3 A:3-434 8037622 1BA3 A:3-434 8055416 1BA3 A:435-520 8055417 1BA3 A:435-520
Output SCOP search for luciferas keyword: 4001964 Dinoflagellate luciferase repeat 4003312 Bacterial luciferase (alkanal monooxygenase)
AI GEMINI Feedback on protein family: AMP-dependent synthetase/ligase family :cross_mark:; Adenylate-forming enzyme superfamily :cross_mark:; AMP-binding enzyme C-terminal domain :check_mark_button:
4. LUCIFERASE VISUALIZATION IN PYMOL
All the visualizations and counts were carried out using the step-by-step guidance of ChatGPT (indeed, very useful). Questions were used as prompts. Once familiar with the software and tasks, further explorations were carried out on my own. Detailed documentation can be found at the end of this section.
DATASETS
PDB 1LCI: first luciferase crystal structure, open conformation
For luciferin ligand and exploration of other conformations: PDB 2D1S and PDB 4G36
3D STRUCTURE BASIC FORMATS


SECONDARY STRUCTURES
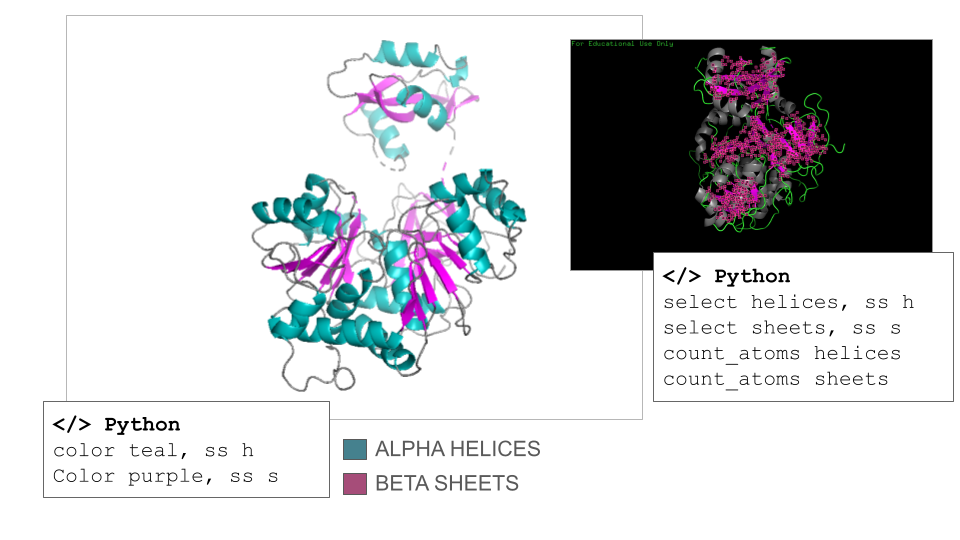


According to ChatGPT, the Firefly Luciferase is an α/β protein, but it is helix-dominated, which is typical for enzymes in the adenylate-forming enzyme family. This fact was surprising to me since I counted 17 helices and 21 sheets on the model when navigating it manually. However, the counts confirmed that the secondary structure of the Firefly Luciferase has more helix than sheets (count_atoms helices: 1367 atoms; count_atoms sheets: 817 atoms).
RESIDUES: HYDROPHATHY PROFILE
There are discrepancies in the classification of the amino acids depending on the sources that can be find online. Thus, I used the amino acids reference chart published by the pharma company Merck KGaA (Darmstadt, Germany) as reference. There are also many ways residues can be represented depending on their classification, so I opted for a binary hydrophobic/hydrophilic approach to better fit the direction of the question.
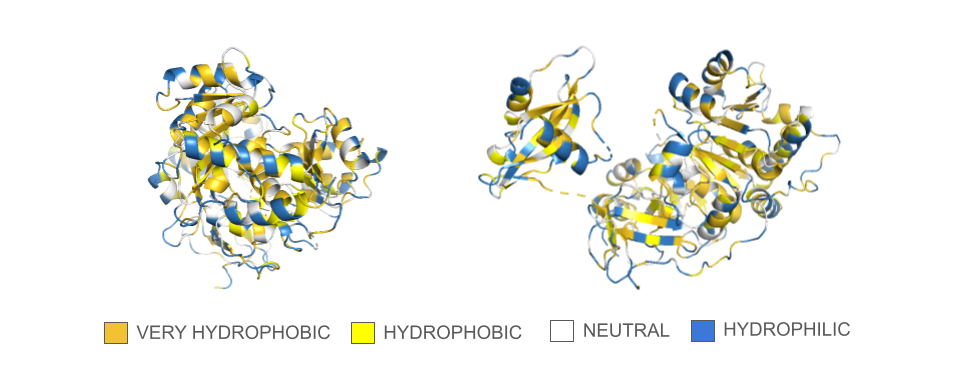

In the spin animation below, one can observe how the protein follows the hydrophobic effect, i.e. how the hydrophobic residues cluster inside the protein. The presence of hydrophilic residues on the surface of the luciferase is coherent with the fact that this enzyme is soluble.
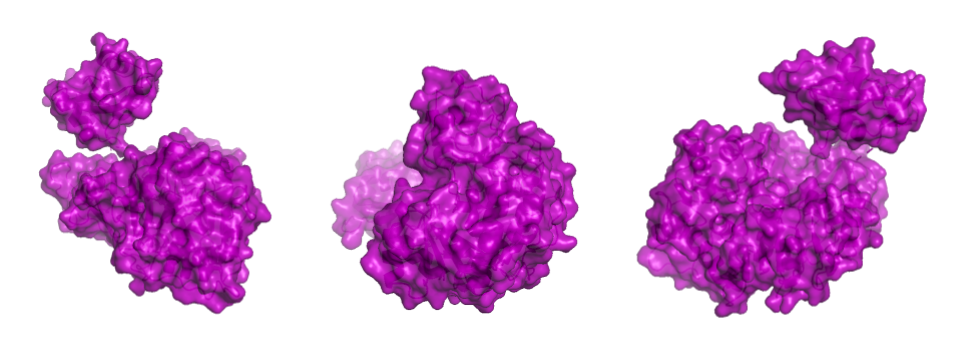
PROTEIN SURFACE
Showing the surface of the protein allows us to see more clearly that the protein is composed of two domains that have a globular shape and present small surface grooves. A large cleft between the N-terminal (residues 1-436, bottom part in the animation) and C-terminal (residues 440–550, on top) domains of the protein is clearly visible when rotating the structure: one can assume that it accommodates the luciferin and the ATP molecule (reference chemical reaction).

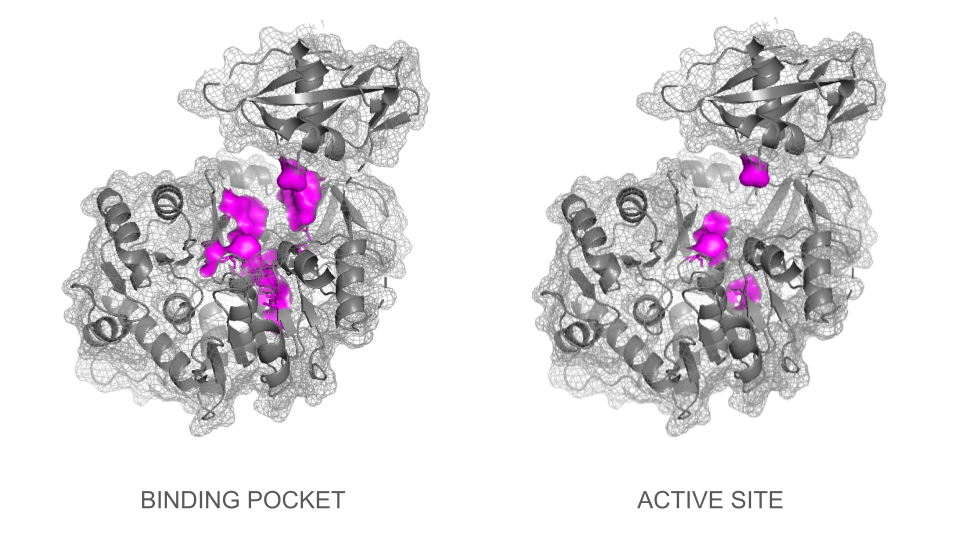
BINDING POCKET & ACTIVE SITE
In a second step, the ligand was added into the visualization. Residues reported to be present in the binding pocket and active site of the firefly luciferase were highlighted using the coloring function. One can observe how the ligand and the key residues match beautifully the binding pocket and how inside the cleft, the ligand enters a tunnel-like cavity forming inside the N-terminal domain.

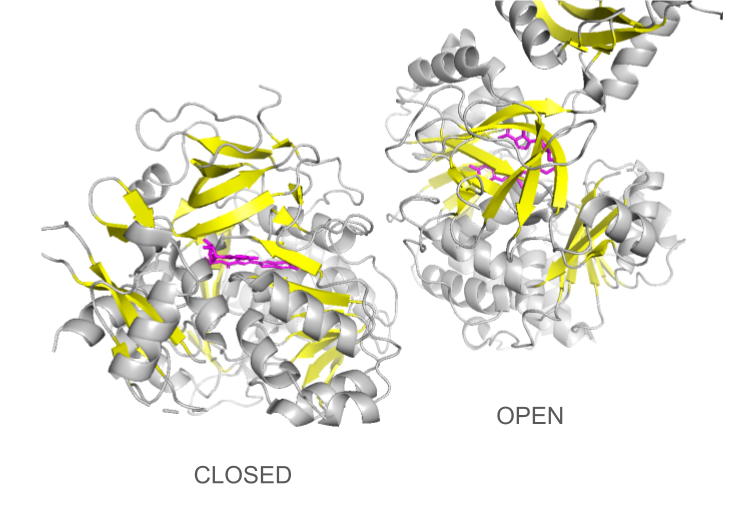
CONFORMATIONAL CHANGES
Loading the data set 2D1S containing both luciferase and ligand was confusing at first: the protein looked different than the 1LCI model. This can be explained by the fact that the enzyme undergoes large conformational changes during the different catalysis states: when the conformation is open, the enzyme allows the ligands to enter the cleft and when it is closed, the catalysis can occur. The C-terminal domain rotates perpendicularly when switching from one state to another, making the structures look quite different even though they are the same protein. Only the open conformation is presented here.

DOCUMENTATION
Documentation Visualization in PyMol: Basics
Import dataset: File > Get PDB > Enter 1LCI (PDB ID for Firefly Luciferase) or </> Python command
Useful commands recommended by ChatGPT:
- fetch 1lci (import dataset)
- hide everything
- show cartoon / show ribbon / show sticks / show spheres
- set sphere_scale, 0.25 / set stick_radius, 0.15 (adjust scale to improve 3D visualisation)
- util.cbag (coloring of the “stick and ball” according to PyMol default color coding: carbon, green; basic residues, blue; acidic residues, red; gray for others)
Documentation Visualization in PyMol: Secondary Structures
Useful commands recommended by ChatGPT:
- select helices, ss h / select sheets, ss s (select secondary structures)
- color color_01, ss h / color color_02, ss s (specific coloring of secondary structures)
- count_atoms helices / count_atoms sheets
Documentation Visualization in PyMol: Hydropathy
Useful (adapted) commands recommended by ChatGPT:
- select veryhydrophobic, resn phe+ile+trp+leu+val+met
- color gold, veryhydrophobic
- select hydrophobic, resn tyr+cys+ala
- color yellow, hydrophobic
- select neutral, resn thr+his+gly+ser+gln
- color white, neutral
- select hydrophilic, resn arg+lys+asn+glu+pro+asp
- color sky, hydrophilic
Documentation Visualization in PyMol: Surface
Useful commands recommended by ChatGPT:
- show surface, selection
- set transparency, 0.2
- clip slab, 20 (view of the surface + inside)
- set surface_cavity_mode, 1
- show mesh
Documentation Visualization in PyMol: Binding pocket and active site
Several strategies were tested to visualize the binding pocket and active site. First, visualization of the types of residues generally involved in binding pockets / active sites (charged, hydrophobe, polar, aromatic etc.) and then, types of residues involved specifically in the firefly luciferase. However, all the representations tested were not specific to the binding site.
Useful commands recommended by ChatGPT for part 01:
- select active, resn lys+arg+his+asp+glu
- color color_01, active
- select inactive, resn ala+ile+leu+met+val+gly+pro+cys+ser+phe+trp+tyr+thr+asn+gln
- color colo_02, inactive
In a second step, visualization of the individual residues involved in the active site of the firefly luciferase gave much better results although one needs to mention that the lists provided by Gemini and ChatGPT were different, and varied depending on iterations.
Commands recommended by ChatGPT to visualize binding pocket:
- select luciferin_binding_site, resi 198+214+218+222+244+245+247+286+340+343+344+347+420+421+422+529
- show sticks, luciferin_binding_site
- color color_01, luciferin_binding_site
- show surface, luciferin_binding_site
Commands recommended by ChatGPT to visualize active site:
- select luciferase_catalytic, resi 218+245+343+529
- show sticks, luciferase_catalytic
- color color_01, luciferase_catalytic
Documentation Visualization in PyMol: Adding ligand
Useful commands recommended by ChatGPT:
- fetch 1LCI
- fetch 4G36
- align 4G36, 1LCI
- select luciferin, 4G36 and organic (organic defines the ligand)
- create luciferin_copy, luciferin
- disable 4G36
- show sticks, luciferin_copy
- color color_01, luciferin_copy
Visual tools
Useful commands recommended by ChatGPT:
- mplay/mstop (play and stop movie)
- mclear (delete movie data to save memory when export is done)
- zoom object_of_interest, 8 (zoom power)
- bg_color white (change background color to white)
- set ray_trace_fog (better image quality)
- Color list
Spin animation
Useful (adapted) commands recommended by ChatGPT:
- mset 1 x360 (number of frames)
- util.mroll(1,360,1)
Zoom-through animation
Useful commands recommended by ChatGPT:
- mset 1 x360
- frame 1
- mview store
- frame 360
- move z, -360
- mview store
- mview interpolate
Video export Videos were exported in mp4 format using the screen recording function.
LEGAL ASPECTS: To use the official version of PyMol for free as a student, I had to sign the agreement copied below.
Addendum: 1- By declaring that I’m a ‘full time student’, I understand that being registered as a committed listener of the HTGAA course which is not an internship and involves 15 to 30 hours of weekly academic work fits the required criteria. 2- “Builds” should not be shared publicly. I understand that “builds” are a specific functionality offered by PyMol and that the visualizations that I have shared on this page are not “builds”.
PyMOL Educational Use Declaration for Flo Razoux
- I, Flo Razoux, am either a full-time student or am engaged in teaching full-time students. After being granted access, I will only apply Education-Use-Only PyMOL Builds (“Builds”) for education purposes and specifically including the following: COURSES or DEGREE: MIT Media Lab, Synthetic Biology, Other, 2026
- I will only share the Builds and their download access credentials with my fellow students and/or teachers, and only via private means.
- I will not post the Builds or their download access credentials in a publicly-accessible location, such as a web page, email list, or blog.
- If I apply PyMOL in any for-profit commercial activity or in any non-profit academic research, then I will compile my own builds from the open-source code or purchase an appropriate PyMOL Subscription in order to access the official PyMOL Builds not limited to educational use only.
- Except as otherwise set forth in Sections 1 through 4, I shall not: (i) modify, translate, adapt, create derivative works from or decompile the Builds, or any portion thereof, or create or attempt to create, by disassembling, reverse engineering or otherwise, the source code from the object code supplied hereunder, (ii) rent, lease, sell, transfer, publish, display, distribute, disclose or make the Builds available to third parties or use the builds, or any portion thereof, in a service bureau, time-sharing or outsourcing service or (iii) remove or alter any proprietary rights notices on the Builds. I acknowledge that the restrictions set forth in clauses (i) through (iii) of the immediately preceding sentence shall apply to distributions by Schrodinger, LLC of any third party software or other materials with the Builds.