Week 2 HW: DNA Read, Write & Edit


Part 0: Basics of Gel Electrophoresis
- Watch Week 2 Lecture (Zoom)
- Watch Week 2 Recitation (Zoom)
- Watch BioBootcamp Day 1 - Day 3 (Zoom)
Part 1: Benchling & In-silico Gel Art
- Make a free account at benchling.com
- Import the Lambda DNA.

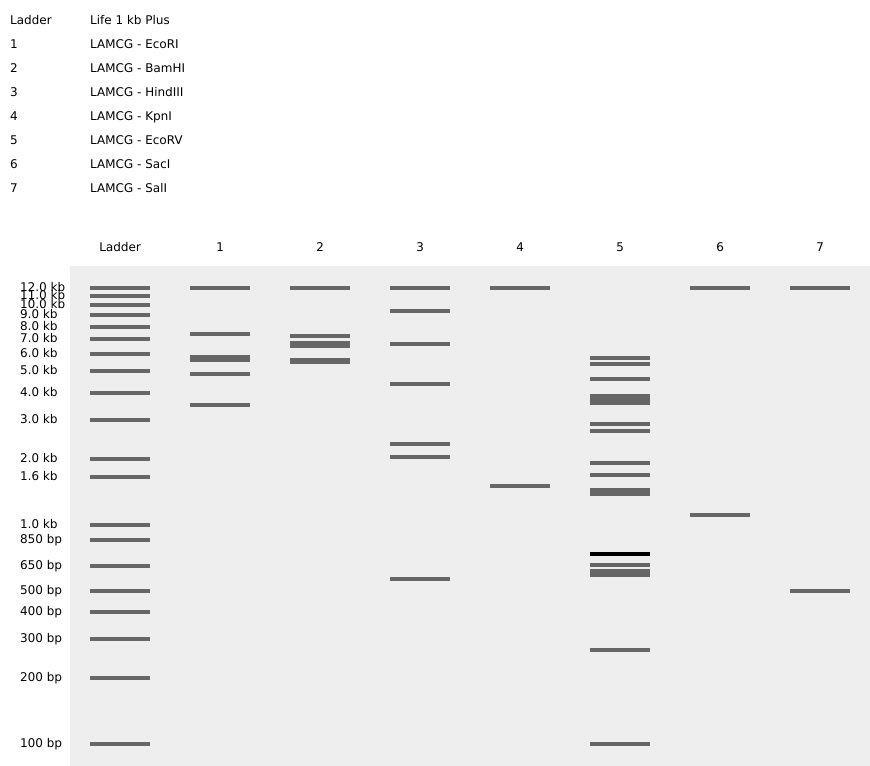
- Simulate Restriction Enzyme Digestion with the following Enzymes:
Artwork
After struggling quite some time with the task of creating artwork with the limited amount of restriction enzymes, in the end decided to stick to a relatively easy and repetitive pattern that with a little imagination has a lot of versatile interpretations:
It can be two friends hanging out
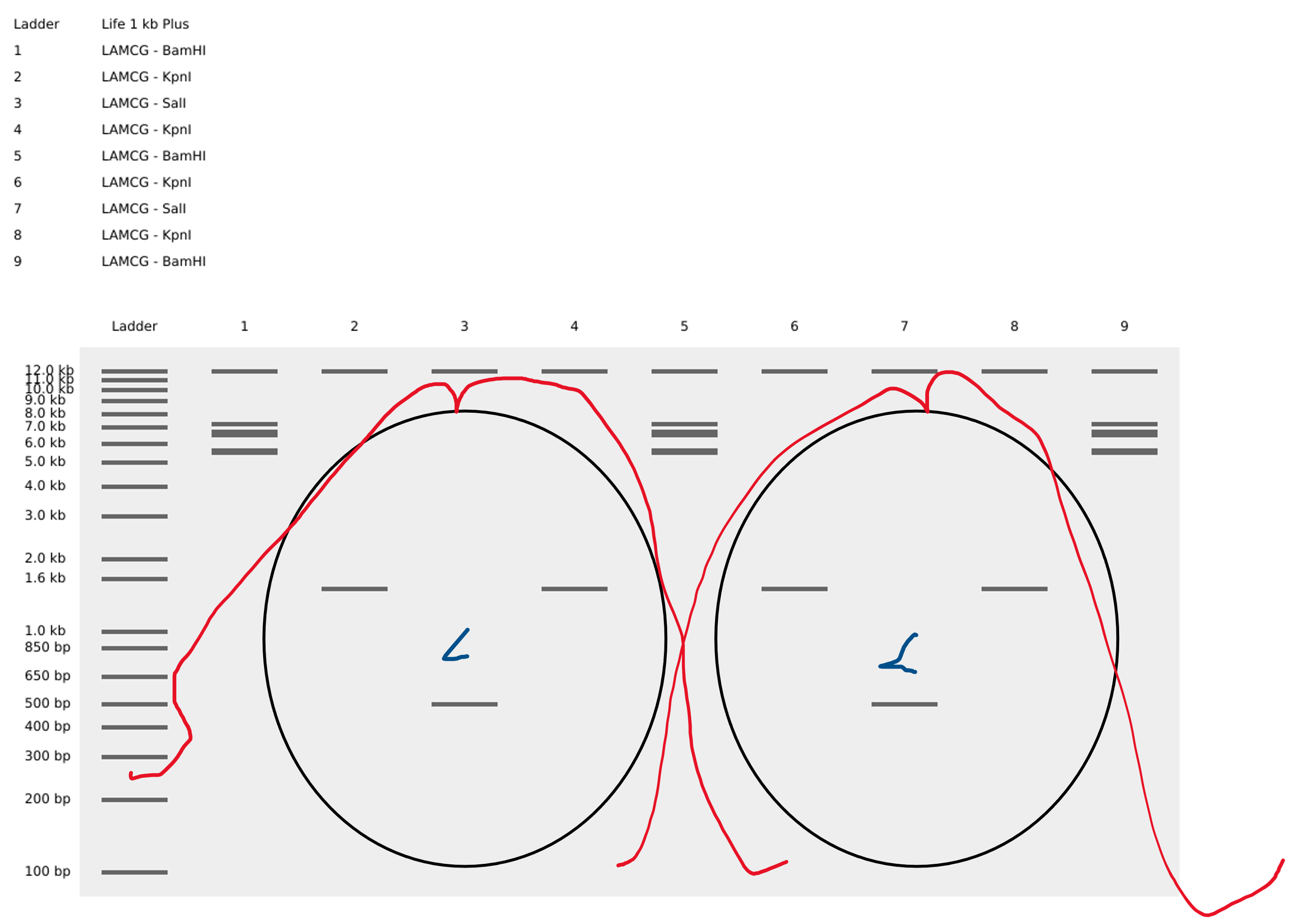
 It can be DNA (or at least a rought estimation of the firrst two loops)
It can be DNA (or at least a rought estimation of the firrst two loops)
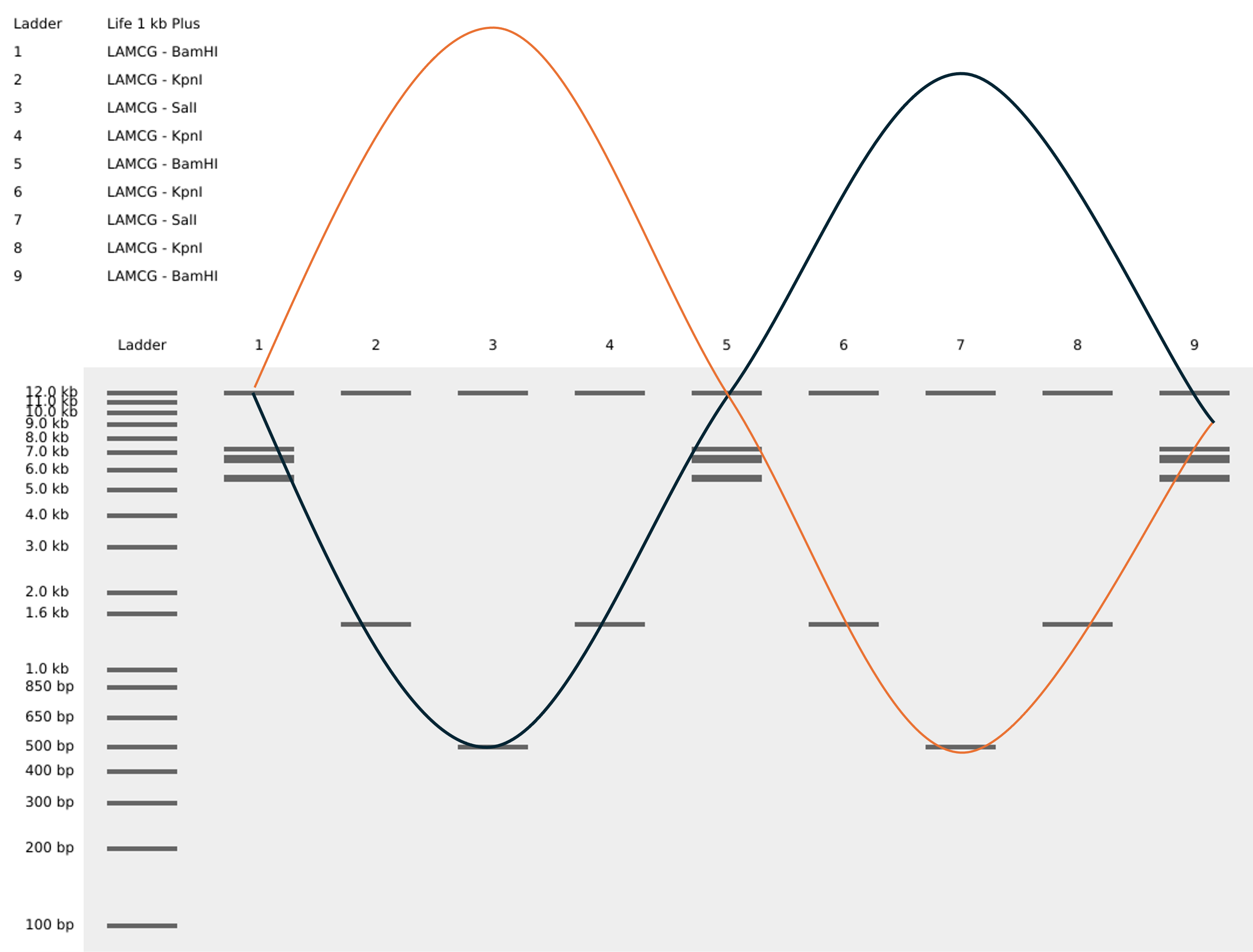
 To help you visualize it a bit better i created some generative AI art
To help you visualize it a bit better i created some generative AI art
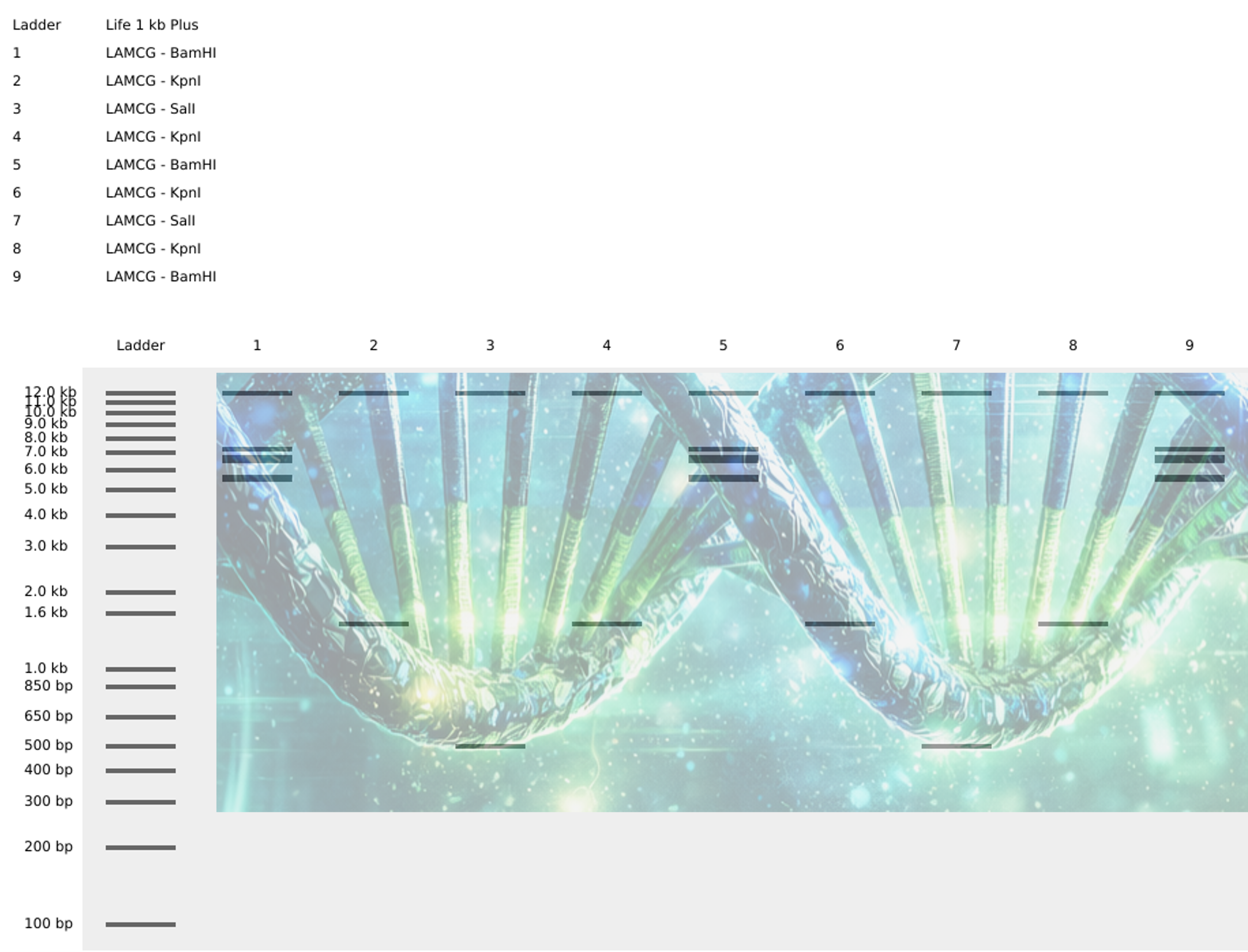

To create the loop like structure, I used the Restriction enzymees BamHI - KpnI - SalI - KpnI - BamHI
Part 2: Gel Art - Restriction Digests and Gel Electrophoresis
not relevant as I don’t have access to a lab
Part 3: DNA Design Challenge
3.1. Choose your protein.
For Week 2 Homework I choose the Growth/differentiation factor 8 (short GDF-8), also known as human myostatin protein. I choose myostatin for the inital reason, that it was the first protein that came to mind. Being known for the viral video of Jo Zayner injecting the DIY-Gene Therapy to knock out the myostatin associated gene, or the many pictures of muscled animals, like cattle and dogs.
Digging further, myostatin seemed to be a good choice, not only for it’s fame. It is a well studied protein, with a clear function to negatively regulate muscle growth, as seen with the example of the “jacked” bagle or the cattle. Furthermore myostatin is not only interesting to biohackers and instagram scrollers, it has actual therapeutical interest, and is actively researched to combat muscular dystrophy. In the up and coming field of Longevity, myostatin is researched to mitigate age-related muscle loss.
Sequence
>sp|O14793|GDF8_HUMAN Growth/differentiation factor 8 OS=Homo sapiens OX=9606 GN=MSTN PE=1 SV=1 MQKLQLCVYIYLFMLIVAGPVDLNENSEQKENVEKEGLCNACTWRQNTKSSRIEAIKIQI LSKLRLETAPNISKDVIRQLLPKAPPLRELIDQYDVQRDDSSDGSLEDDDYHATTETIIT MPTESDFLMQVDGKPKCCFFKFSSKIQYNKVVKAQLWIYLRPVETPTTVFVQILRLIKPM KDGTRYTGIRSLKLDMNPGTGIWQSIDVKTVLQNWLKQPESNLGIEIKALDENGHDLAVT FPGPGEDGLNPFLEVKVTDTPKRSRRDFGLDCDEHSTESRCCRYPLTVDFEAFGWDWIIA PKRYKANYCSGECEFVFLQKYPHTHLVHQANPRGSAGPCCTPTKMSPINMLYFNGKEQII YGKIPAMVVDRCGCS
3.2 Reverse Translate: Protein (amino acid) sequence to DNA (nucleotide) sequence.
The Central Dogma discussed in class and recitation describes the process in which DNA sequence becomes transcribed and translated into protein. The Central Dogma gives us the framework to work backwards from a given protein sequence and infer the DNA sequence that the protein is derived from. Using one of the tools discussed in class, NCBI or online tools (google “reverse translation tools”), determine the nucleotide sequence that corresponds to the protein sequence you chose above.
Using the Reverse Translate tool from Bioinformatics.org, I got the following results:
>reverse translation of sp|O14793|GDF8_HUMAN Growth/differentiation factor 8 OS=Homo sapiens OX=9606 GN=MSTN PE=1 SV=1 to a 1125 base sequence of most likely codons. atgcagaaactgcagctgtgcgtgtatatttatctgtttatgctgattgtggcgggcccg gtggatctgaacgaaaacagcgaacagaaagaaaacgtggaaaaagaaggcctgtgcaac gcgtgcacctggcgccagaacaccaaaagcagccgcattgaagcgattaaaattcagatt ctgagcaaactgcgcctggaaaccgcgccgaacattagcaaagatgtgattcgccagctg ctgccgaaagcgccgccgctgcgcgaactgattgatcagtatgatgtgcagcgcgatgat agcagcgatggcagcctggaagatgatgattatcatgcgaccaccgaaaccattattacc atgccgaccgaaagcgattttctgatgcaggtggatggcaaaccgaaatgctgctttttt aaatttagcagcaaaattcagtataacaaagtggtgaaagcgcagctgtggatttatctg cgcccggtggaaaccccgaccaccgtgtttgtgcagattctgcgcctgattaaaccgatg aaagatggcacccgctataccggcattcgcagcctgaaactggatatgaacccgggcacc ggcatttggcagagcattgatgtgaaaaccgtgctgcagaactggctgaaacagccggaa agcaacctgggcattgaaattaaagcgctggatgaaaacggccatgatctggcggtgacc tttccgggcccgggcgaagatggcctgaacccgtttctggaagtgaaagtgaccgatacc ccgaaacgcagccgccgcgattttggcctggattgcgatgaacatagcaccgaaagccgc tgctgccgctatccgctgaccgtggattttgaagcgtttggctgggattggattattgcg ccgaaacgctataaagcgaactattgcagcggcgaatgcgaatttgtgtttctgcagaaa tatccgcatacccatctggtgcatcaggcgaacccgcgcggcagcgcgggcccgtgctgc accccgaccaaaatgagcccgattaacatgctgtattttaacggcaaagaacagattatt tatggcaaaattccggcgatggtggtggatcgctgcggctgcagc
3.3. Codon optimization.
Once a nucleotide sequence of your protein is determined, you need to codon optimize your sequence. You may, once again, utilize google for a “codon optimization tool”. In your own words, describe why you need to optimize codon usage. Which organism have you chosen to optimize the codon sequence for and why?
Organisms have different procivities for using certain codons. Some codons are used more frequently in one orgamis. So for the same amino acide certain codons are “preferred” over others. If the inserted DNA matches the preferred codons of the organism more translation happens. In my case, as the protein I’m trying to express is of human origin, though expressed in e.coli (common for early experiments), many of the codons common in homo sapiens, are rare in e.coli.
Using the Codon Optimization tool from VectorBuilder, and choosing e.coli K-12 as the organism i get the following optimized codon:
ATGCAGAAACTGCAGCTGTGCGTTTACATTTATCTGTTCATGCTGATTGTGGCCGGCCCGGTGGATCTGAACGAAAACAGTGAACAGAAAGAAAACGTGGAAAAAGAAGGTCTGTGCAACGCCTGTACCTGGCGCCAGAATACCAAATCGAGCCGCATTGAAGCCATTAAAATTCAGATCCTGTCAAAACTGCGTCTGGAAACCGCGCCGAATATTAGCAAAGATGTGATCCGTCAGCTGCTGCCGAAAGCCCCGCCGCTGCGTGAACTGATTGATCAGTATGATGTGCAGCGCGATGATAGCAGCGATGGCAGCCTGGAAGATGATGATTATCACGCGACCACCGAAACCATTATTACCATGCCGACCGAAAGCGATTTTCTGATGCAGGTGGATGGCAAACCGAAATGCTGCTTCTTCAAATTTAGCTCGAAAATTCAATATAATAAAGTGGTGAAAGCGCAGCTGTGGATCTATCTGCGCCCGGTGGAAACCCCGACCACCGTGTTTGTGCAGATTCTGCGCCTGATTAAACCGATGAAAGATGGCACCCGCTACACCGGCATTCGCAGCCTGAAACTGGATATGAACCCGGGCACCGGCATCTGGCAGAGCATTGATGTGAAAACCGTTCTGCAGAATTGGCTGAAACAGCCGGAAAGCAACCTGGGCATTGAAATTAAAGCCCTGGATGAAAATGGCCATGATCTGGCAGTGACCTTTCCGGGCCCGGGCGAAGATGGCCTGAATCCGTTCCTGGAAGTGAAAGTGACCGATACCCCGAAACGCAGCCGCCGCGACTTTGGCCTGGATTGCGATGAACACAGCACCGAAAGCCGCTGCTGCCGCTACCCGCTGACCGTGGATTTTGAAGCGTTCGGCTGGGATTGGATTATTGCGCCGAAACGCTATAAGGCGAACTACTGCAGCGGTGAATGCGAATTTGTGTTTCTGCAGAAATATCCGCACACCCATCTGGTGCACCAGGCAAACCCGCGCGGCAGCGCGGGCCCGTGCTGTACCCCGACCAAAATGAGCCCGATTAACATGCTGTATTTTAACGGCAAAGAACAGATTATCTATGGCAAAATCCCGGCGATGGTTGTGGATCGCTGCGGTTGTAGC while avoiding cleavage sites of restriction enzymes: BamHI HindIII
Whether the e.coli is the proper host for this application is debateable, as e.coli lacks the post-translational modification capabilities. Other hosts like CHO cells or human cells, could prove to be a better choice, if one aims for a properly folded, functioning protein.
3.4. You have a sequence! Now what?
What technologies could be used to produce this protein from your DNA? Describe in your words the DNA sequence can be transcribed and translated into your protein. You may describe either cell-dependent or cell-free methods, or both.
Let me focus on cell-dependent methods. There are several expression systems like e.coli, yeast and mamalian cells. Each system has their pros and cons. E.coli is a procaryote, it’s cheap, fast and well established, though it lacks post translational modification abilities. Therefore the myostatin might be misfolded. Addtionally the cells have to be lysed to get to the myostatin. Yeast is eukaryotic, therefore it has some post-translational folding capabilities, additionally secretion is possible. Mammalian cells offer human-like folding, and secretion, though they grow slower, are a-lot harder to handle, offer lower yields and are significantly more expensive than e.coli or yeast.
To get the DNA into the orgamism:
- Perform PCR on the optimized myostatin DNA strand, to generate many copies of said gene. make sure the DNA strand is flanked with BanHI and HindIII sites.
- Open the plasmid at the previously avoided restriction sites (BamHI HindIII), creating compatible sticky ends.
- Cut the DNA strand with the same restriction enzymes
- Mix plasmid and DNA strand, introduce ligase enzymes and ATP, to connect the matching sticky ends.
- Transform the plasmid into e.coli using common transformation methods, e.g. heat-shock or electroporation
- (optional) screen for uptake of plasmid
Having chosen e.coli here the DNA would transscribe into mRNA, which in turn would translate into myostating facilitated by ribosomes. For lab-sized fermentation, a batch system is sufficient, on industrial scale a fed-batch system would be used.
Part 4: Prepare a Twist DNA Synthesis Order
4.1. Create a Twist account and a Benchling account
- Create Twist Account
- Create Benchling Account
4.2. Build Your DNA Insert Sequence
Following the example on the course site, this is the linear map of the sequence:
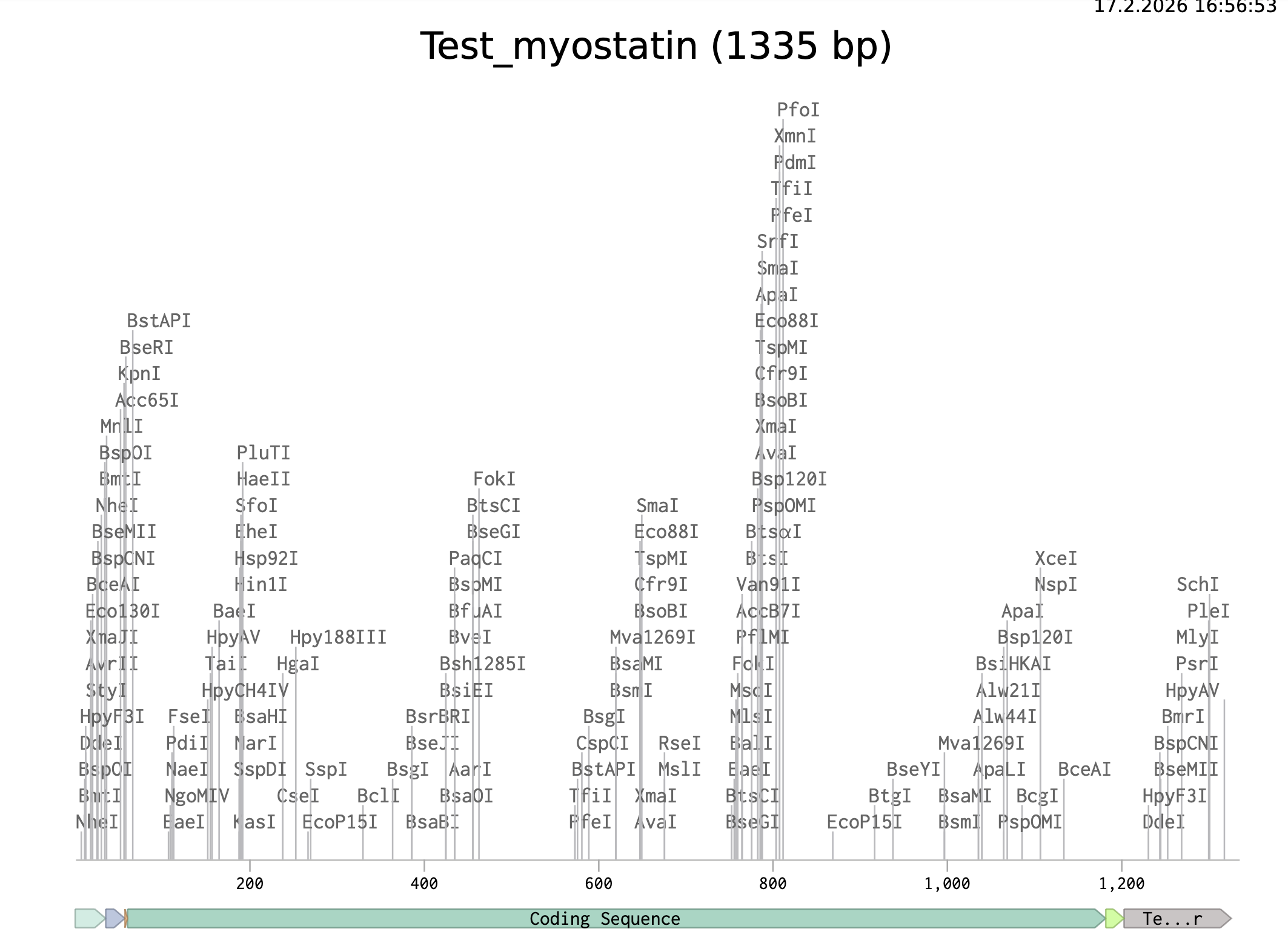


>Test_myostatin TTTACGGCTAGCTCAGTCCTAGGTATAGTGCTAGCCATTAAAGAGGAGAAAGGTACCATGCAGAAACTGCAGCTGTGCG TTTACATTTATCTGTTCATGCTGATTGTGGCCGGCCCGGTGGATCTGAACGAAAACAGTGAACAGAAAGAAAACGTGGA AAAAGAAGGTCTGTGCAACGCCTGTACCTGGCGCCAGAATACCAAATCGAGCCGCATTGAAGCCATTAAAATTCAGATC CTGTCAAAACTGCGTCTGGAAACCGCGCCGAATATTAGCAAAGATGTGATCCGTCAGCTGCTGCCGAAAGCCCCGCCGC TGCGTGAACTGATTGATCAGTATGATGTGCAGCGCGATGATAGCAGCGATGGCAGCCTGGAAGATGATGATTATCACGC GACCACCGAAACCATTATTACCATGCCGACCGAAAGCGATTTTCTGATGCAGGTGGATGGCAAACCGAAATGCTGCTTC TTCAAATTTAGCTCGAAAATTCAATATAATAAAGTGGTGAAAGCGCAGCTGTGGATCTATCTGCGCCCGGTGGAAACCC CGACCACCGTGTTTGTGCAGATTCTGCGCCTGATTAAACCGATGAAAGATGGCACCCGCTACACCGGCATTCGCAGCCT GAAACTGGATATGAACCCGGGCACCGGCATCTGGCAGAGCATTGATGTGAAAACCGTTCTGCAGAATTGGCTGAAACAG CCGGAAAGCAACCTGGGCATTGAAATTAAAGCCCTGGATGAAAATGGCCATGATCTGGCAGTGACCTTTCCGGGCCCGG GCGAAGATGGCCTGAATCCGTTCCTGGAAGTGAAAGTGACCGATACCCCGAAACGCAGCCGCCGCGACTTTGGCCTGGA TTGCGATGAACACAGCACCGAAAGCCGCTGCTGCCGCTACCCGCTGACCGTGGATTTTGAAGCGTTCGGCTGGGATTGG ATTATTGCGCCGAAACGCTATAAGGCGAACTACTGCAGCGGTGAATGCGAATTTGTGTTTCTGCAGAAATATCCGCACA CCCATCTGGTGCACCAGGCAAACCCGCGCGGCAGCGCGGGCCCGTGCTGTACCCCGACCAAAATGAGCCCGATTAACAT GCTGTATTTTAACGGCAAAGAACAGATTATCTATGGCAAAATCCCGGCGATGGTTGTGGATCGCTGCGGTTGTAGCCAT CACCATCACCATCATCACTAACCAGGCATCAAATAAAACGAAAGGCTCAGTCGAAAGACTGGGCCTTTCGTTTTATCTG TTGTTTGTCGGTGAACGCTCTCTACTAGAGTCACACTGGCTCACCTTCGGGTGGGCCTTTCTGCGTTTATA
In Twist having chosen the Clonal Gene, with the vector: pTwist Amp High Copy, this is the output from Twist

 Following is the final Clonal Gene in Benchling
Following is the final Clonal Gene in Benchling
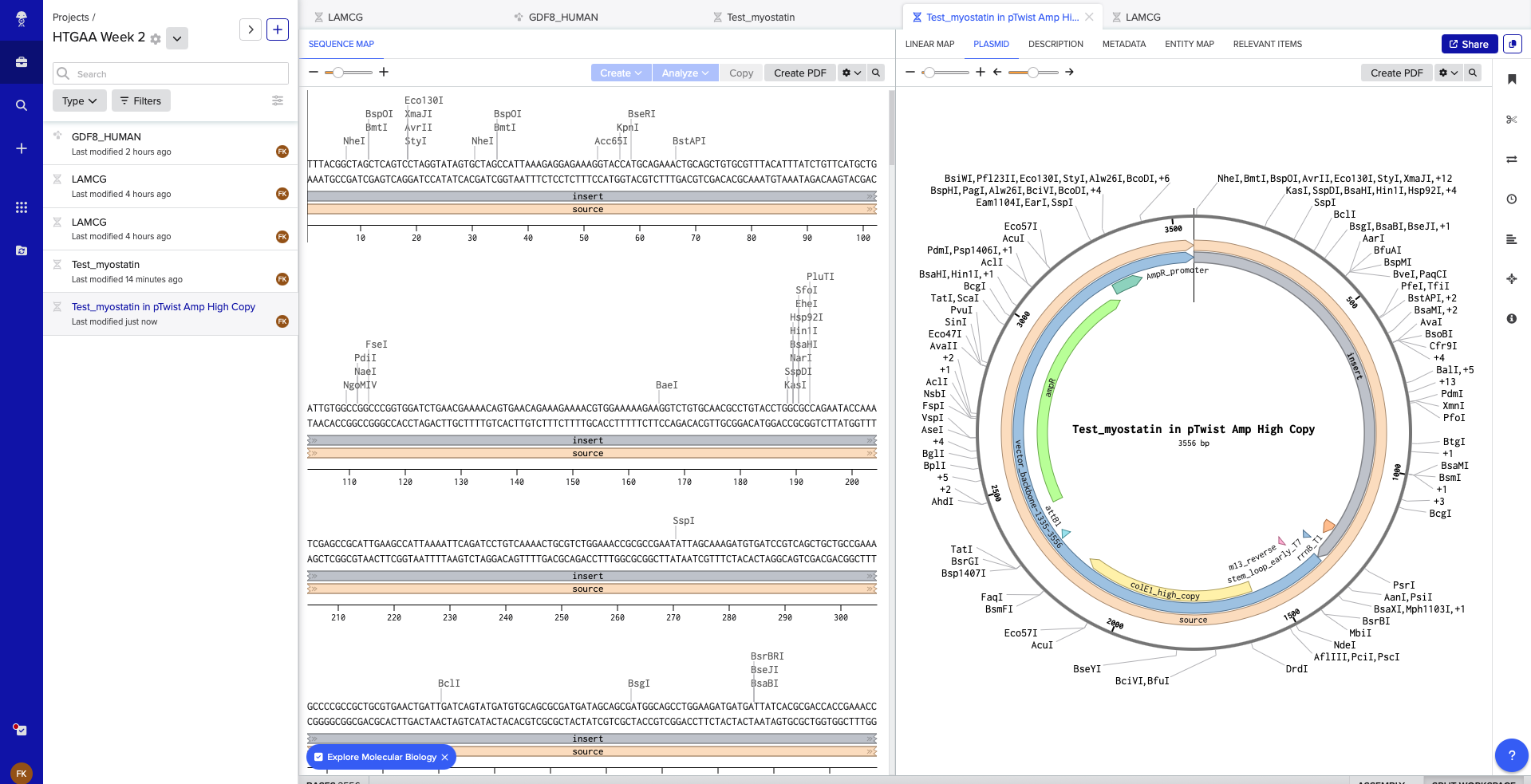

Part 5: DNA Read/Write/Edit
5.1 DNA Read
(i) What DNA would you want to sequence (e.g., read) and why? This could be DNA related to human health (e.g. genes related to disease research), environmental monitoring (e.g., sewage waste water, biodiversity analysis), and beyond (e.g. DNA data storage, biobank).
I’m interested in genetic origin for muscle growth. People have different outcomes for the similar inputs, I’m interested into the marginal influence Gene’s have on ones physiological development. While overall health is important, this also has clinical application, e.g. for patients with muscle loss diseases, age-related muscle atrophy. Therefore a number of genes can be studied. Using search and LLMs, these are relevant proteins for muscle growth,
- Myostatin: given that my Design a Gene Challenge was about this, it makes sense to study this gene. I suspect mutations in the gene lead to enhanced or reduced function.
- ACTN3: Determines between “fast” or “slow” muscle fiber twitches. They determine whether one is dispositioned for heavy lifting or endurance.
- IGF1: This gene expresses a growth factor that aids muscle repair and their growtn after exercise. People with different versions might respond differently to certain training styles.
(ii) In lecture, a variety of sequencing technologies were mentioned. What technology or technologies would you use to perform sequencing on your DNA and why?
I’d choose NGS sequencing, as I’m interested in targeted genes. NGS allows for a good middle ground of cost, and accuracy, and also has established protocols for variant detetction.
Also answer the following questions:
Is your method first-, second- or third-generation or other? How so?
Second-generation sequencing. DNA fragments are sequenced simultaniously in massively paralellized fashion.
What is your input? How do you prepare your input (e.g. fragmentation, adapter ligation, PCR)? List the essential steps.
With NGS library prep is necessary. First DNA is extracted. Then the DNA is fragmented into smaller chunks (around 300-500 bp). Next adapter ligation is used to attache each DNA chunk to the coded oligonucleotide on the flow plate. PCR afterwards is optional
What are the essential steps of your chosen sequencing technology, how does it decode the bases of your DNA sample (base calling)?
NGS sequencing takes DNA and fragments it, then ligates it to adapter sequences, which add to a sequencing library. The fragments bind to a flow cell, then aplified to create clusters of identical DNA strands. The gene strand is sequenced by synthesizing a complementary strand using labeled nucleotides, and only one base is incorporated per cycle. After incorporation the flow cell is imaged, and the specific fluoresence is used to identify the added base.
What is the output of your chosen sequencing technology?
A FASTQ files, with a large number of small DNA fragment readouts. Additionally these fragements are then aligned and referenced. These can then be screened for differences.
5.2 DNA Write
(i) What DNA would you want to synthesize (e.g., write) and why? These could be individual genes, clusters of genes or genetic circuits, whole genomes, and beyond. As described in class thus far, applications could range from therapeutics and drug discovery (e.g., mRNA vaccines and therapies) to novel biomaterials (e.g. structural proteins), to sensors (e.g., genetic circuits for sensing and responding to inflammation, environmental stimuli, etc.), to art (DNA origamis). If possible, include the specific genetic sequence(s) of what you would like to synthesize! You will have the opportunity to actually have Twist synthesize these DNA constructs! :)
A synthetic DNA construct that can be injected locally, that will express miRNAs that silence myostatin mRNA, to allow for temporary enhanced muscle growth. This would pose a therapeutical intervention, either for biohackers or patients with degenerative muscle diseases.
(ii) What technology or technologies would you use to perform this DNA synthesis and why? Also answer the following questions:
I’d probably choose Phosphoramidite Chemistry, as its the current gold standard and ample for smaller DNA fragments.
What are the essential steps of your chosen sequencing methods? What are the limitations of your sequencing method (if any) in terms of speed, accuracy, scalability?
I’m a bit confused why sequencing instead of synthesis methods are mentioned. So assuming that synthesis is meant, I’ll talk about the synthesis based on Phosphoramidite Chemistry The essential steps with PC is repeating the so called Coupling cycle, that is repeated for each base. First the protecting group gets removed (deetritylation), theen the next phosphoramidite nucleeatide is added (coupling), the unreacted 5’ groups get blocked (cappping), lastly unstable phosphite is converted to stable phosphate (oxidation). This way fragemnts up to 200bp are synthesised and later assembled.
5.3 DNA Edit
(i) What DNA would you want to edit and why? In class, George shared a variety of ways to edit the genes and genomes of humans and other organisms. Such DNA editing technologies have profound implications for human health, development, and even human longevity and human augmentation. DNA editing is also already commonly leveraged for flora and fauna, for example in nature conservation efforts, (animal/plant restoration, de-extinction), or in agriculture (e.g. plant breeding, nitrogen fixation). What kinds of edits might you want to make to DNA (e.g., human genomes and beyond) and why?
Going with the theme a reversible knockout for myostatin expression. It could be an alternative treatment for patients with degenerative muscular diseases as well as biohackers.
(ii) What technology or technologies would you use to perform these DNA edits and why? To ensure reversability, I’d consider base editors, introducing stop codons into the myostatin coding sequence. I’d chose base editors as they make single-nucleotide changes without cutting DNA, to reduce the chance of unwanted off-target effects, compared to classical CRISPR-Cas9 technology. Also answer the following questions:
How does your technology of choice edit DNA? What are the essential steps?
First Guide RNA detects a specific myostatin sequence, then guides the Base editor there. Next a base conversion occures, where e.g. a cytosine base editor converts a C to a T. With this edit normal codons are converted to pre-mature stop codons. This leads to the myostatin being misformed and ideally not bioactive.
What preparation do you need to do (e.g. design steps) and what is the input (e.g. DNA template, enzymes, plasmids, primers, guides, cells) for the editing?
Select a target sequence to turn a normal codon into a premature stop codon, e.g. CGA to TGA. Next you have to design a Guide RNA to position the base editor at the chosen target. Next check whether there are any off target effects. I’d need the DNA of the base editor, the DNA Template for the guide RNA, an plasmid that integrates the base editor with the guideRNA, restricition and ligation enzymes. For injection into humans, probably also the targeted muscle cells as well as culture media and transfection equipment.
What are the limitations of your editing methods (if any) in terms of efficiency or precision?
Base editing has some key limitation, the most severe one being the low efficiency compared to classical CRISPR methods. Human cells are notoriously difficult to work with, muscle cells are some of the hardest to edit in humans. The delivery of the editor to muscle cells poses another challenge. The base editors have limited target options, based on their capability to make edits in a narrow basepair window. Off target effects are also a concern. Also it needs to be checked, that the editor only comes in contact with the intended C to T conversion as it converts Cs indiscriminantly.
Use of Generative AI
Generative AI was used as a drafting aid throughout the development of this homework assignment. Specifically, it supported the structuring and refinement of complex ideas at the of DNA Design as well as aiding the understanding of DNA Read, Write and Edit technologies. The AI was used to iteratively clarify language and explore alternative framings, while all substantive ideas, judgments, and final decisions were made by the author.
