Week 3 HW: Lab Automation
Python Script for Opentrons Artwork
- Review this week’s recitation and this week’s lab for details on the Opentrons and programming it.
- Generate an artistic design using the GUI at opentrons-art.rcdonovan.com.
I generated a quick design using the above mentioned tool: BioPunk Initials
See the:

- Using the coordinates from the GUI, follow the instructions in the HTGAA26 Opentrons Colab to write your own Python script which draws your design using the Opentrons.
- You may use AI assistance for this coding — Google Gemini is integrated into Colab (see the stylized star bottom center); it will do a good job writing functional Python, while you probably need to take charge of the art concept.
- If you’re a proficient programmer and you’d rather code something mathematical or algorithmic instead of using your GUI coordinates, you may do that instead.
- If the Python component is proving too problematic even with AI and human assistance, download the full Python script from the GUI website and submit that:
- If you use AI to help complete this homework or lab, document how you used AI and which models made contributions.
As its good practice in Software Engineering, not to reinvent the wheel, I had a look at the provided examples, and figured Example 7 would be a good basis for my requirements. Nevertheless, significant updates needed to be done to make the code useful for my purposes. These include:
- Importing necessary libraries
- Converting CSV file reading to Pandas Dataframe
- Removing inversion of y-data
- Changing the color scheme
- Submit your Python file via this form.
Post-Lab Questions
Find and describe a published paper that utilizes the Opentrons or an automation tool to achieve novel biological applications.
Given my background in tissue engineering I picked the paper “Fabrication of cell culture hydrogels by robotic liquid handling automation for high-throughput drug testing” published in “Communications Engineering” (2025)4:222 DOI
The paper introduces HYDRA (HYDrogels by Robotic liquid-handling Automation), a method for fabricating flat and thin hydrogel films directly in standard 96- and 384-well HTS plates using an Opentrons liquid handling machine. The paper tackles one of the main problems in modern drug discovery. Around 50% of compounds passing preclinical assays fail human trials. This has several proposed reasons, the paper tackles one of the widely mentioned problems. In standard HTS cells are grown on plastic dishes that do not resemble the ECM structure of the human body, nor have the dynamic context (mechanical, electrical, etc.) that the ECM provides. Organ on a chip improve the missing biomimicry, but are not compatible with HTS. Additionally, they are too complex too manufacture cheaply and are incompatible with automated screening pipelines. There are existing hydrogel coatings, but they either are so thin that they don’t provide the proper ECM environment mechanically, or so thick that they block high-resolution imaging. Both produce a curved meniscus that makes uniform cell seeding impossible.
This gap, hydrogel coatings thin enough for imaging and thick enough for mechanosensing, is tackled with HYDRA. HYDRA uses fish gelatin for its hydrogel base material, dissolved in PBS at 5-20% w/v and is cross-linked enzymatically with transglutaminase at 0.5-2% w/v. The method was demonstrated on an Opentrons, for easy accessibility, for scalability on a INTRGRA Assist Plus. The robot dispenses precalculated sub-contact volumes 96 well plates: 12 micro liter, 382 well plate: 1 micro liter), and immediately re-aspirates the volume, to archive a thin film. This is possible by using contact angle hysteresis. The volume was optimized using FE simulations. The process takes around 10 minutes for the whole plate, then the plates are gelled, and incubated at 37 degree Celsius, sterilized and swollen and rinsed in PBS. Afterwards cells can be seeded.
The hydrogel thin films are between 10-50 micrometer thickness, with tunable stiffness. Lastly the authors validated several imaging platforms (digital holographic, widefield fluorescence, and high-resolution confocal microscopy) see Fig. 1
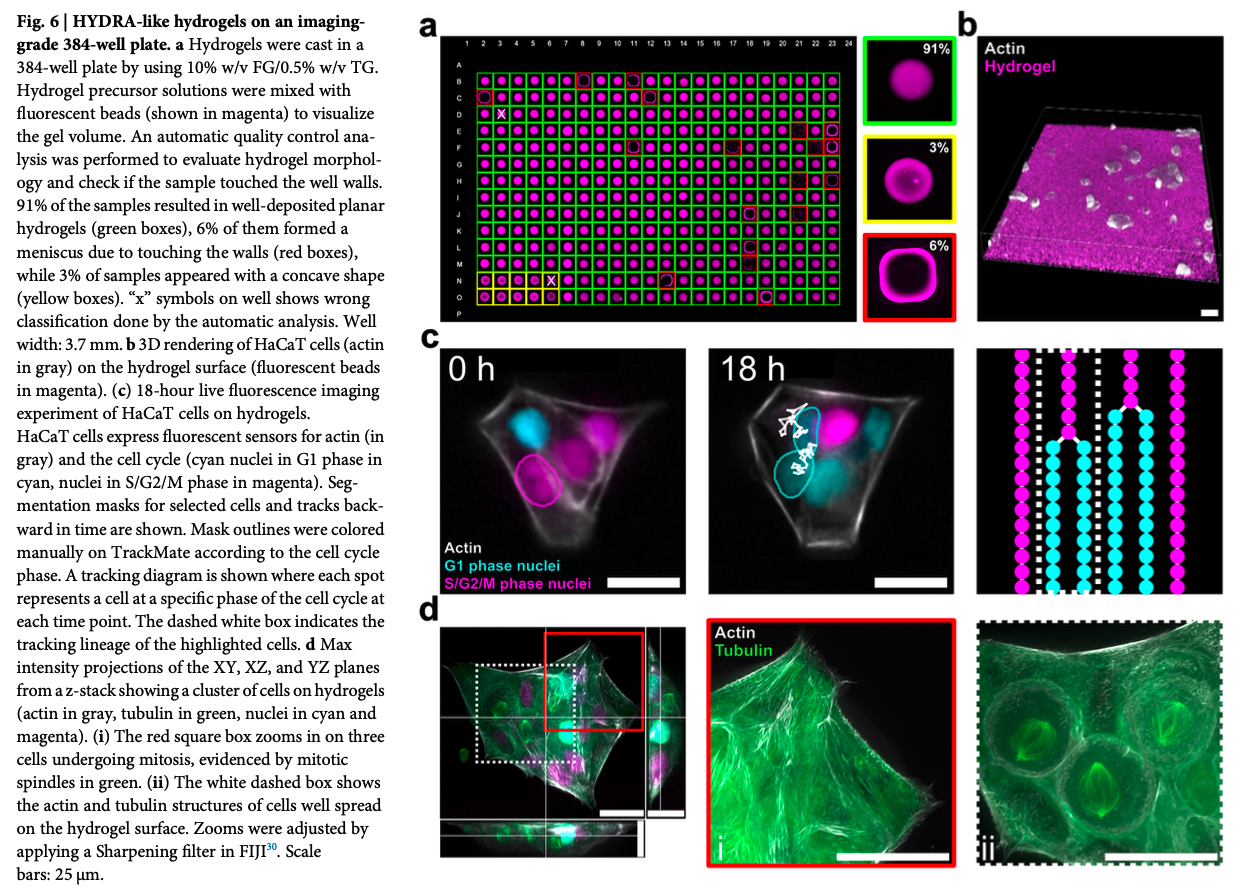

Write a description about what you intend to do with automation tools for your final project. You may include example pseudocode, Python scripts, 3D printed holders, a plan for how to use Ginkgo Nebula, and more. You may reference this week’s recitation slide deck for lab automation details.
Developmental biology aims to understand how tissues and cells develop from dynamic, history-dependent processes governed by interacting biochemical, electrical, mechanical cues in a temporal manner. While these properties are well established theoretically, in practice experiments lack behind by what can be executed manually. Even if automation is deployed it only addresses one of the drawbacks of manual experimentation. This forced experiments to make simplifications, such as searching only low-parameter search spaces, coarse temporal guiding and open loop design of experiments. This results in heuristic sampling and interpretation of results with lacking search spaces. Using automation directly addresses these limitations by enabling precise temporal control, systematic exploration of parameter rich search spaces, and adaptive, feedback-driven intervention. This is especially relevant in organoid and organ-on-chip systems.
Mimicking the ECM in developmental biology
Many developmental signals only have their proper effect through the timing, duration, and frequency, rather than concentration alone. Temporal regimes are rarely explored experimentally, e.g. competence windows, pulsed signaling, and oscillatory dynamics due to the impracticality of executing. Automated liquid-handling systems can change that by enabling stable, repeatable temporal execution.
Exploring the design space
Cell fate and tissue morphology have very high-dimensional development search spaces. This comes from the interaction of multiple biochemical and mechanical variables that evolve over time. Contrasting this intuition or slow sequential tuning is currently used. Automation enables systematic, multi-parameter exploration of said search space. The goal of automation is to find non-linear responses and regime changes that are not obvious in manual experiments.
Guiding Experiments
Development proceeds through continuous feedback between tissue state and signaling environment, yet most experiments are designed in an open-end fashion, with data analysis and redesign being manual and in a discrete fashion. Automation should allow experiments to close the loop, where readouts inform the next steps of the experiment. The goal is to adaptively explore tissue formation.
Final Project Ideas
As explained in this week’s recitation, add 1-3 slides in your Node’s section of this slide deck with 3 ideas you have for an Individual Final Project. Be sure to put your name, city, and country on your slide!
- slides are added
Use of Generative AI
Generative AI was used as a conceptual drafting aid during the development of this project. Specifically, it supported the structuring and refinement of complex ideas related to automated biological experimentation and experimental design in tissue engineering, as well as clarifying concepts around temporal control, high-dimensional experimentation, and reproducibility in laboratory automation. The AI was used to iteratively improve clarity of language and to explore alternative conceptual framings. No generative AI was used for the implementation or coding components of the project, and all substantive ideas, technical decisions, and final judgments were made by the author.