Week 03 HW: Lab-Automation
1. Python Script for Opentrons Artwork
This artwork was generated using the HTGAA26 Opentrons Colab environment. opentron_code The design was implemented programmatically using geometric constructions and multi-color pipetting logic.

To properly render Devanagari text (e.g., “चित्”) using PIL in Google Colab, system-level fonts must be installed before executing the Opentrons script. The Noto Sans Devanagari font was installed using the following commands in a separate Colab cell:
- !sudo apt-get update -qq
- !sudo apt-get install -y -qq fonts-noto-core fonts-noto-unhinted
- !fc-list | grep -i “Devanagari” | head -n 20
2. Post-Lab Questions
2.1 Find and describe a published paper that utilizes the Opentrons or an automation tool to achieve novel biological applications.
A relevant example of automation enabling novel biological applications is Nielsen et al., 2016. “Genetic circuit design automation.” Science
In this study, the authors developed an automated design–build–test workflow to systematically engineer genetic logic circuits in living cells. Rather than constructing gene circuits through manual trial-and-error, they integrated computational modeling, standardized genetic parts, and high-throughput experimental validation to design functional Boolean logic gates such as AND, OR, NOR, and more complex multi-layer circuits.
The key innovation was the automation of circuit design and screening. Computational tools were used to predict circuit behavior based on promoter strength, repressor activity, and regulatory architecture. These predictions were then experimentally validated using high-throughput plate-based assays, allowing many circuit variants to be constructed and tested in parallel. Automated measurement of reporter outputs (e.g., fluorescence) enabled quantitative evaluation of logic performance, signal thresholds, and leakiness.
Automation significantly reduced the combinatorial complexity inherent in multi-input genetic circuit design. Instead of manually constructing and testing a few variants, the workflow enabled systematic exploration of many possible architectures, improving robustness and reproducibility. This approach demonstrated that genetic logic circuits can be engineered in a scalable and programmable manner, similar to electronic circuit design.
This paper shows how automation transforms synthetic biology from artisanal genetic assembly into an engineering discipline with predictive modeling and systematic validation.
2.2 Write a description about what you intend to do with automation tools for your final project. You may include example pseudocode, Python scripts, 3D printed holders, a plan for how to use Ginkgo Nebula, and more. You may reference this week’s recitation slide deck for lab automation details.
For my final project, I intend to engineer a bacterial therapeutic system for oncology applications. The core design involves a plasmid encoding a NAND logic gate integrating multiple tumor-associated inputs, including hypoxia, elevated lactate levels, and ultrasound stimulation. Only when specific tumor microenvironment conditions are satisfied would the circuit activate expression of a therapeutic cassette (e.g., the eriBCDEN complex).
Automation would be essential to systematically design and validate this multi-input logic system. I propose implementing an automated design–build–test workflow focused on high-throughput screening of circuit variants.
Combinatorial plasmid assembly Promoter variants responsive to hypoxia, lactate, and ultrasound would be modularly assembled using combinatorial DNA assembly methods (e.g., Golden Gate). An acoustic liquid handler (e.g., Echo) could transfer defined promoter and RBS fragments into specified wells to systematically generate circuit variants.
RBS strength tuning Ribosome binding site variants of defined translation strengths would be introduced to modulate repressor expression levels. This step allows fine control of expression thresholds and minimization of basal leakiness, which is critical for achieving accurate NAND logic behavior.
High-throughput culture setup Following transformation and colony selection, bacterial variants would be distributed into 96-well plates using liquid handling robotics. This enables parallel testing of multiple architectures under standardized growth conditions.
Controlled environmental testing Each well would be exposed to defined combinations of normoxia/hypoxia conditions, graded lactate concentrations, and ultrasound stimulation (applied externally). This systematic input matrix allows evaluation of all Boolean input states.
Automated reporter quantification A fluorescent reporter would be used during circuit prototyping prior to therapeutic deployment. Fluorescence measurements using a plate reader (e.g., PHERAstar) would quantify output across all input combinations, enabling assessment of Boolean fidelity, dynamic range, activation thresholds, and leakiness.
Because multi-input genetic logic circuits require careful balancing of transcriptional and translational parameters, manual testing would be slow and prone to variability. Automation enables parallelized combinatorial screening and quantitative validation before integrating the therapeutic cassette.
By integrating automation into the circuit development pipeline, this approach would accelerate optimization of tumor-specific logic and improve safety and precision in engineered bacterial cancer therapies.
3. Final Project Ideas
Gated Siderophore Bacteriotherapy: My First is a programmable bacterial therapy that targets tumors by expressing a salmochelin siderophore cassette only under highly controlled conditions. The control logic is a two-input gate: hypoxia provides spatial specificity, and ultrasound provides clinician timing. Mechanistically, both inputs are implemented through DNA-binding repressors that toggle promoter accessibility—this week I analyzed the lambda cI DNA-binding domain (PDB 1LMB) as a structural model for repressor–operator control, which maps directly onto the TlpA39–P_tlpA thermal switch used for ultrasound activation. Next, I’m integrating the dual-repressor logic into a single promoter architecture and validating it with sequence/structure design tools.
Neuroengineering - Metabolic Calcium Control: My second project is a closed-loop neuroengineering circuit to keep neuronal activity in a safe range. The input is lactate, a simple metabolic signal that rises in stressed tissue. I use a lactate-responsive promoter to drive a nanobody-based controller that tunes calcium entry when activity becomes too strong. I’ll test it in C. elegans touch neurons using the mec-4d degeneration model, where calcium dynamics can be imaged in vivo. The goal is a genetic feedback system that links metabolism to stable neural signaling.
Ultrasound-Triggered Genetic Switches: My third project is the enabling technology behind my tumor-targeting bacteria: using ultrasound as a non-invasive control signal. The core idea is to build biological transducers—such as gas vesicles and mechanosensitive channels—that convert focused ultrasound into a reliable genetic switch. That switch becomes an external “ON command” you can combine with internal signals like hypoxia to build multi-input logic in living cells. So this project turns ultrasound into a general remote-control layer, and my bacteriotherapy project is the first concrete use case.
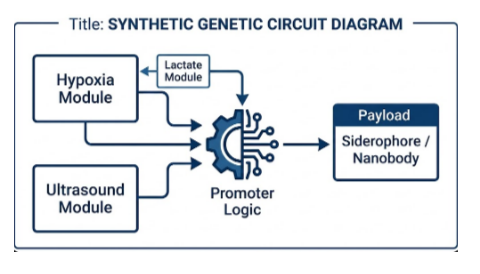

Figure: High-level architecture of the genetic control system. Hypoxia, ultrasound, and lactate sensing modules feed into a promoter-logic layer that gates expression of the therapeutic payload (either a siderophore cassette or a nanobody), enabling multi-input control over when and where the output is produced.