Week 05 HW: Protein Design Part II
Part A: SOD1 Binder Peptide Design
Part 1: Generate Binders with PepMLM
- Begin by retrieving the human SOD1 sequence from UniProt (P00441) and introducing the A4V mutation.
Original sequencesp|P00441|SODC_HUMAN Superoxide dismutase
MATKAVCVLKGDGPVQGIINFEQKESNGPVKVWGSIKGLTEGLHGFHVHEFGDNTAGCTS
AGPHFNPLSRKHGGPKDEERHVGDLGNVTADKDGVADVSIEDSVISLSGDHCIIGRTLVV
HEKADDLGKGGNEESTKTGNAGSRLACGVIGIAQ
Variant: A4V
MATKVVCVLKGDGPVQGIINFEQKESNGPVKVWGSIKGLTEGLHGFHVHEFGDNTAGCTS
AGPHFNPLSRKHGGPKDEERHVGDLGNVTADKDGVADVSIEDSVISLSGDHCIIGRTLVV
HEKADDLGKGGNEESTKTGNAGSRLACGVIGIAQ
- Generate four peptides of length 12 amino acids conditioned on the mutant SOD1 sequence.
Using the mutant SOD1 sequence as input, PepMLM Colab generated four 12-residue candidate binders
| index | Binder | Pseudo Perplexity |
|---|---|---|
| 0 | WRYPVAGARHWE | 18.89836973999799 |
| 1 | KLYYPVVVAWWK | 17.203301905376957 |
| 2 | HRYPVVVAALKE | 11.315016775827807 |
| 3 | WLYGAAVLRHGE | 15.526728984710877 |
- Record the perplexity scores that indicate PepMLM’s confidence in the binders.
PepMLM’s pseudo-perplexity scores indicate the model’s confidence in the generated binders, with lower values corresponding to higher confidence. Among the four generated peptides, HRYPVVVAALKE showed the highest confidence (lowest pseudo-perplexity, 11.315), whereas WRYPVAGARHWE showed the lowest confidence (highest pseudo-perplexity, 18.898). The reference peptide FLYRWLPSRRGG was included for comparison, but no pseudo-perplexity score for it was provided in the displayed output.
Part 2: Evaluate Binders with AlphaFold3
- For each peptide, submit the mutant SOD1 sequence followed by the peptide sequence as separate chains to model the protein-peptide complex.
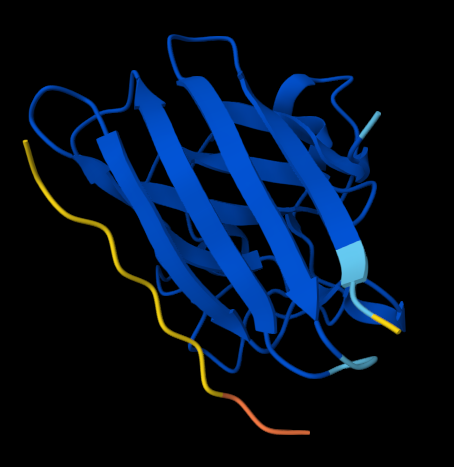

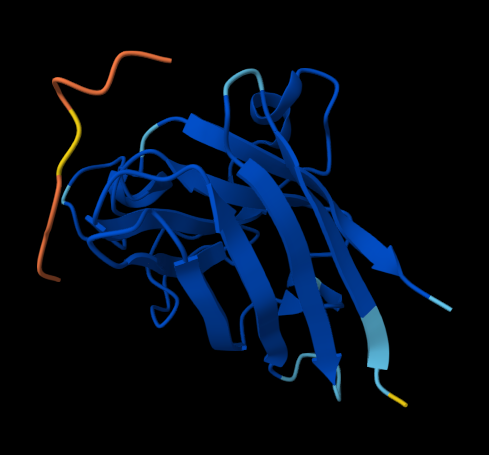

KLVYPVVVAWWK (ipTM = 0.59) FLYRWLPSRRGG (ipTM = 0.33)
Figure 1. AlphaFold-predicted SOD1 A4V complexes shown side by side for comparison. Left: complex with the PepMLM-generated peptide KLVYPVVVAWWK. Right: complex with the known SOD1-binding peptide FLYRWLPSRRGG.
- Record the ipTM score and briefly describe where the peptide appears to bind. Does it localize near the N-terminus where A4V sits? Does it engage the β-barrel region or approach the dimer interface? Does it appear surface-bound or partially buried?
WRYPVAGARHWE — ipTM = 0.35. The peptide appears loosely surface-bound on the SOD1 surface, with no clear evidence of a well-defined buried binding mode. It does not convincingly localize near the N-terminus/A4V region and instead appears to contact the β-barrel surface in a weakly resolved manner.
KLVYPVVVAWWK — ipTM = 0.59. This peptide showed the strongest predicted interface among the tested candidates. It appears mainly surface-bound and extended along the β-barrel region, rather than deeply buried in a pocket. It does not clearly localize near the N-terminus where A4V sits, and no strong interaction with the dimer-interface region is evident.
HRYPVVVAALKE — ipTM = 0.48. The peptide appears surface-associated with low-to-moderate interface confidence. It does not seem to bind near the N-terminal A4V region and instead contacts an exposed outer region of SOD1, consistent with a surface-bound interaction rather than a partially buried one.
WLYGAAVLRHGE — ipTM = 0.31. This peptide shows a very weak predicted interface and appears largely extended and surface-associated, without a defined binding pocket. It does not localize near the A4V-containing N-terminus, nor does it show a clear approach to the dimer interface. The interaction appears predominantly surface-bound.
FLYRWLPSRRGG — ipTM = 0.33. The known binder also showed a low-confidence interface in this AlphaFold prediction. The peptide appears loosely surface-bound rather than buried, with no strong evidence of localization near the N-terminus/A4V site or a clearly resolved interaction at the dimer-interface region.
- In a short paragraph, describe the ipTM values you observe and whether any PepMLM-generated peptide matches or exceeds the known binder.
Overall, the predicted SOD1 A4V–peptide complexes showed low-to-moderate ipTM values, indicating that none of the modeled interfaces was predicted with high confidence. Among the PepMLM-generated candidates, KLVYPVVVAWWK produced the highest ipTM (0.59), followed by HRYPVVVAALKE (0.48), whereas WRYPVAGARHWE (0.35) and WLYGAAVLRHGE (0.31) showed weaker predicted interfaces. The known SOD1-binding peptide FLYRWLPSRRGG gave an ipTM of 0.33. Therefore, the best PepMLM-generated peptide, KLVYPVVVAWWK, exceeded the known binder in this AlphaFold-based comparison, and HRYPVVVAALKE and WRYPVAGARHWE also matched or surpassed it, whereas WLYGAAVLRHGE did not.
Part 3: Evaluate Properties of Generated Peptides in the PeptiVerse
Using PeptiVerse, let’s evaluate the therapeutic properties of your peptide! For each PepMLM-generated peptide:
- Results
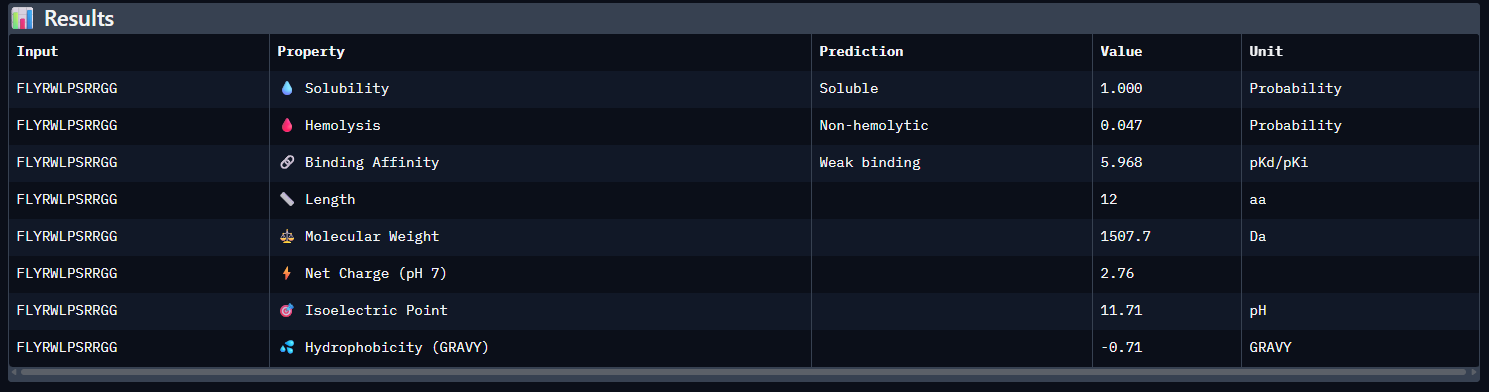

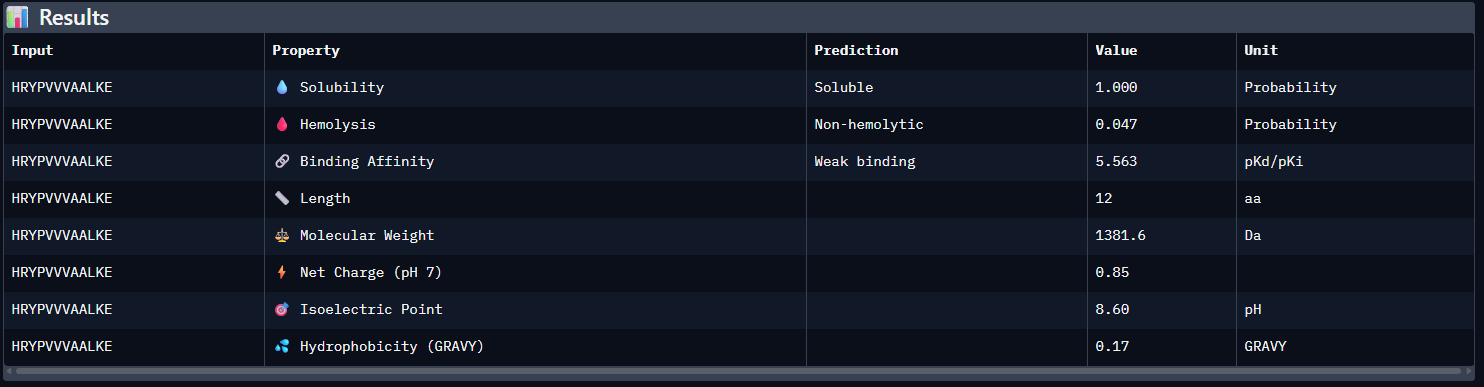

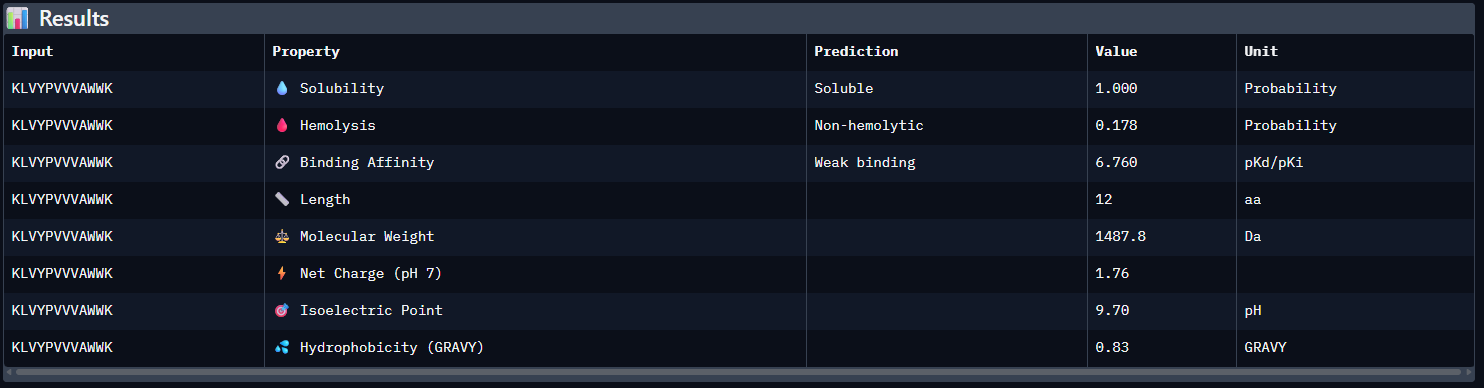

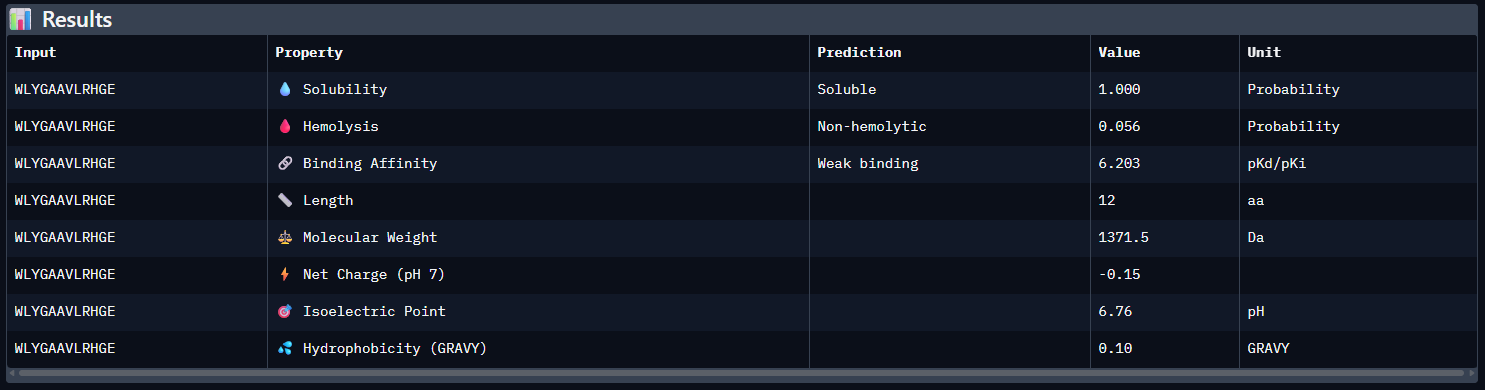

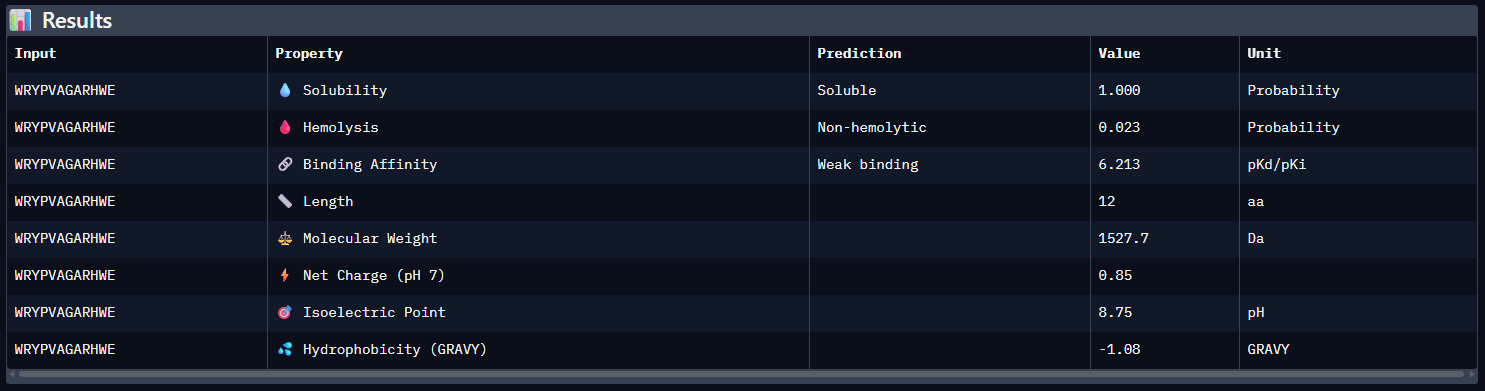

- Compare these predictions to what you observed structurally with AlphaFold3. In a short paragraph, describe what you see. Do peptides with higher ipTM also show stronger predicted affinity? Are any strong binders predicted to be hemolytic or poorly soluble? Which peptide best balances predicted binding and therapeutic properties?
Compared with the AlphaFold3 models, the sequence-based peptide property predictions show only partial agreement with the structural results. The peptide with the highest structural confidence, KLVYPVVVAWWK (ipTM = 0.59), also has the strongest predicted binding affinity among the candidates (pKd/pKi = 6.760), so in that case the two methods are consistent. However, this trend is not perfect across all peptides: for example, WRYPVAGARHWE has a low AlphaFold3 interface score (ipTM = 0.35) but still a moderately favorable predicted affinity (6.213), while the known binder FLYRWLPSRRGG showed both low structural confidence (ipTM = 0.33) and the weakest predicted affinity (5.968). Importantly, all peptides were predicted to be soluble and non-hemolytic, so none of the better binders appears disqualified by poor solubility or overt hemolysis risk. Among them, KLVYPVVVAWWK appears to best balance predicted binding and therapeutic properties, since it combines the highest ipTM, the strongest predicted affinity, full solubility, and a non-hemolytic prediction, although its hemolysis probability (0.178) is somewhat higher than that of the other candidates and would still merit attention in follow-up validation.
- Choose one peptide you would advance and justify your decision briefly.
I would advance KLVYPVVVAWWK. Among the tested peptides, it showed the highest AlphaFold3 interface confidence (ipTM = 0.59) and the strongest predicted binding affinity (pKd/pKi = 6.760), making it the most consistent top candidate across both structural and sequence-based evaluations. It was also predicted to be soluble and non-hemolytic, which supports its therapeutic potential. Although its hemolysis probability was somewhat higher than that of the other candidates, it remained below the threshold for a hemolytic prediction, so overall it provided the best balance between predicted binding performance and developability.
Part 4: Generate Optimized Peptides with moPPIt
After generation, briefly describe how these moPPit peptides differ from your PepMLM peptides. How would you evaluate these peptides before advancing them to clinical studies?
| Peptide | Method | Target motif | Affinity score | ipTM | Interpretation |
|---|---|---|---|---|---|
| KLVYPVVVAWWK | PepMLM | None | 6.760 | 0.59 | Best overall PepMLM candidate |
| HRYPVVVAALKE | PepMLM | None | 5.563 | 0.48 | Intermediate |
| WRYPVAGARHWE | PepMLM | None | 6.213 | 0.35 | Weak interface |
| WLYGAAVLRHGE | PepMLM | None | 6.203 | 0.31 | Weak interface |
| FLYRWLPSRRGG | Known binder | None | 5.968 | 0.33 | Reference; weaker than best PepMLM candidate |
| RYTDIQQYCGKW | moPPIt | 29–35 | 6.423 | 0.42 | Moderate but not strong |
| GQSDYCTRQGKI | moPPIt | 29–35 | 5.933 | 0.52 | Best moPPIt structural result so far |
| KRGKTCLECYQY | moPPIt | 29–35 | 7.286 | 0.28 | Strong property scores but poor structural support |
| GCGYSRSYTKYE | moPPIt | 107–115 | 7.286 | 0.44 | Good score profile, but only moderate structural support |
| GDRSEYCSQKKQ | moPPIt | 107–115 | 6.418 | 0.53 | Best moPPIt structural result; moderate interface confidence |
| EQSRYGHKQDER | moPPIt | 107–115 | 5.221 | 0.36 | High motif score but weak structural support |
Note: Residues 29–35; 107-115 were selected as a hypothesis-driven target motif based on their apparent mutational sensitivity in the ESM2 deep mutational scan. This choice does not demonstrate that the region is surface-exposed or experimentally validated as a peptide-binding site, but it highlights a segment that may be structurally or functionally important and therefore worth testing in a controlled design setting.
Before any clinical development, these peptides would require stepwise preclinical evaluation. First, their intended mechanism of action would need to be clarified: whether they are meant to bind mutant SOD1 merely as recognition molecules, to block a pathogenic interaction surface, to interfere with dimerization, or to reduce misfolding or aggregation. They would then need to be tested experimentally in biochemical and cellular assays to confirm real binding to mutant SOD1, measure affinity and selectivity relative to wild-type SOD1, and determine whether binding produces a meaningful functional effect. This should be followed by preclinical studies addressing stability, protease susceptibility, uptake or delivery, toxicity, hemolysis, immunogenicity risk, pharmacokinetics, and efficacy in relevant animal models of SOD1-associated disease before considering any first-in-human study. In other words, these computational results could justify preclinical follow-up, but they are far from sufficient to support direct clinical advancement.