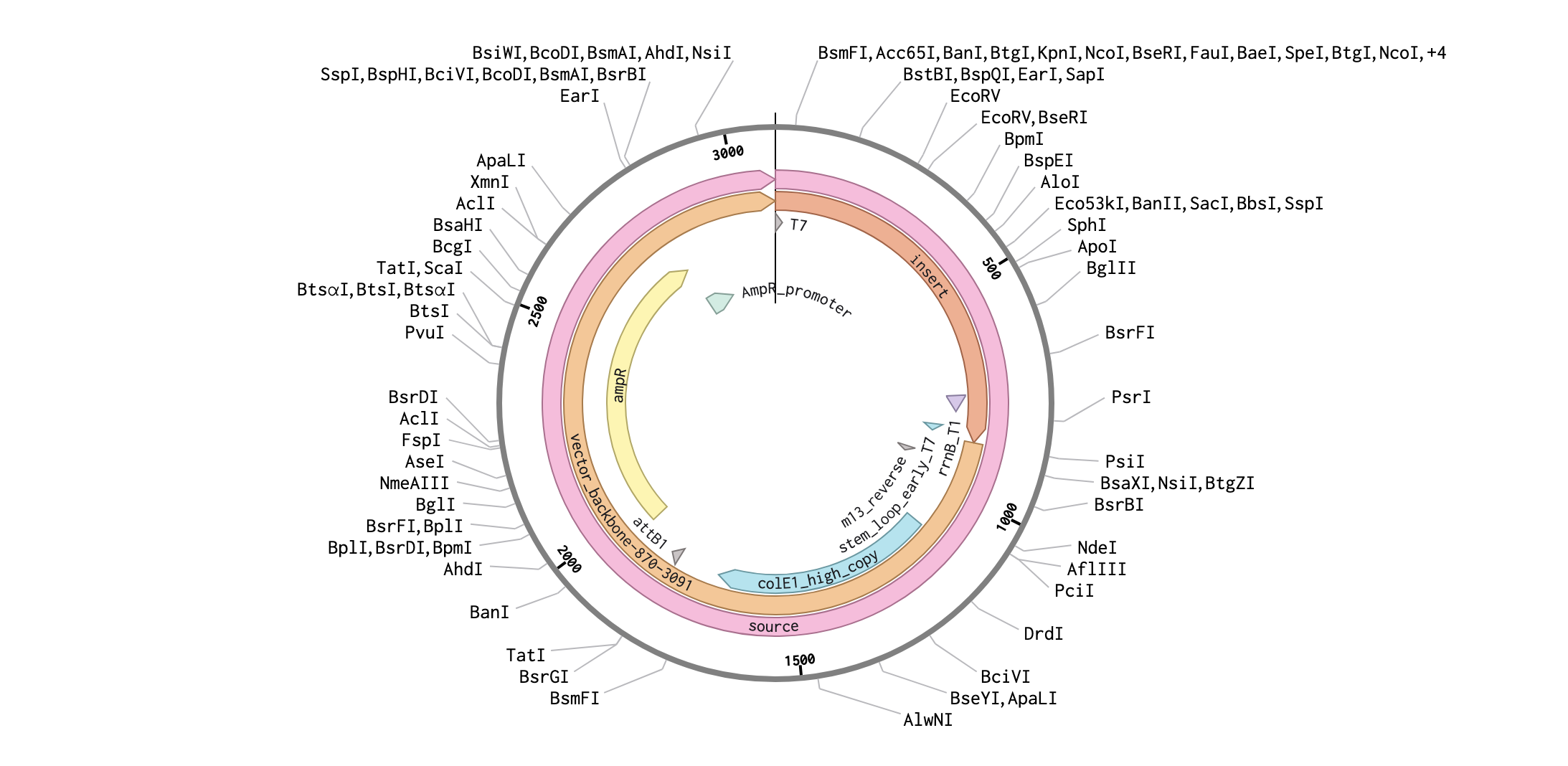
Week 2 HW: DNA read/write/edit
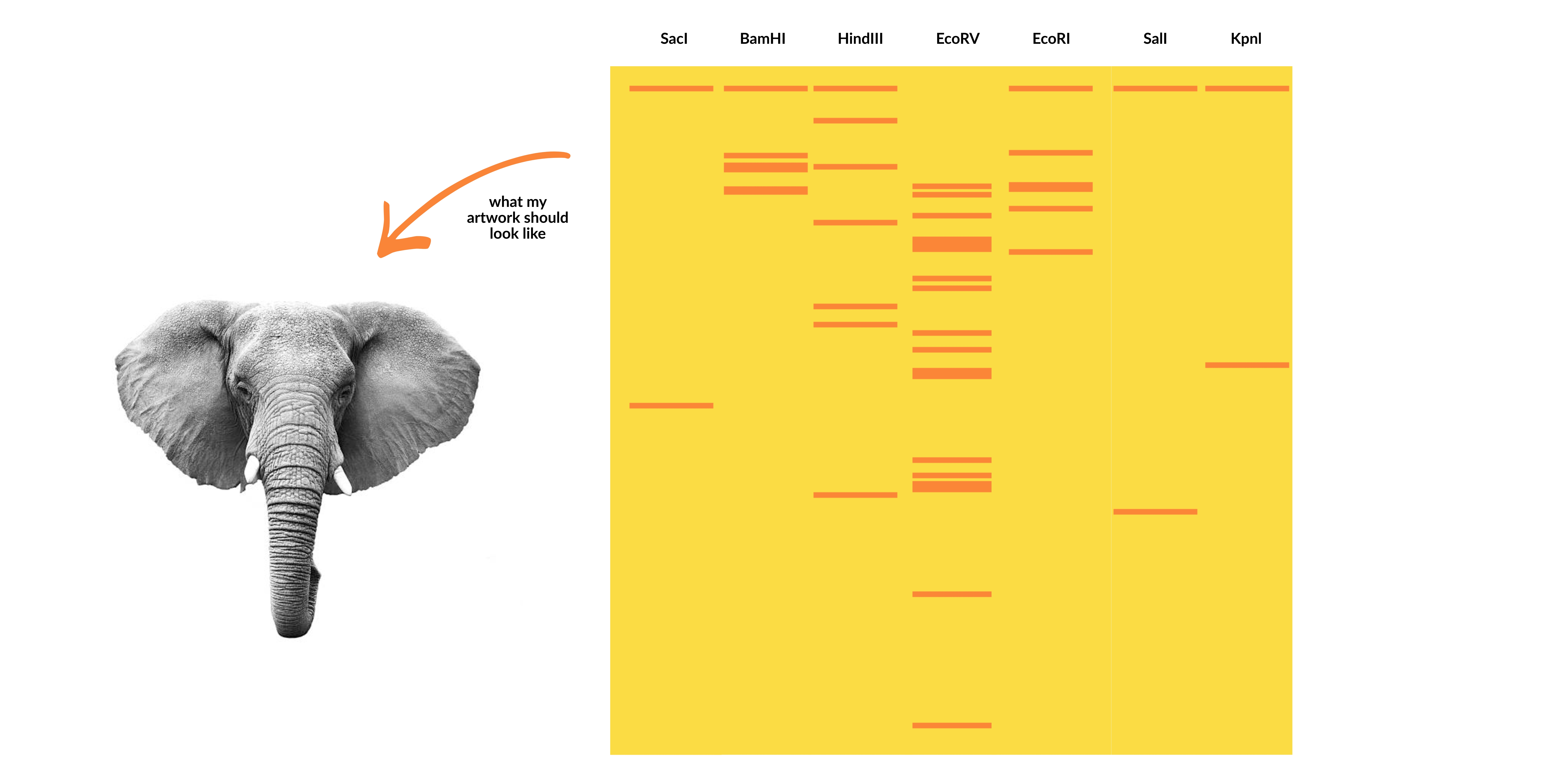
Part I: Benchling & In-silico Gel Art
Create a pattern/image in the style of Paul Vanouse’s Latent Figure Protocol artworks.

Part III: DNA Design Challenge
1. Choose your protein.
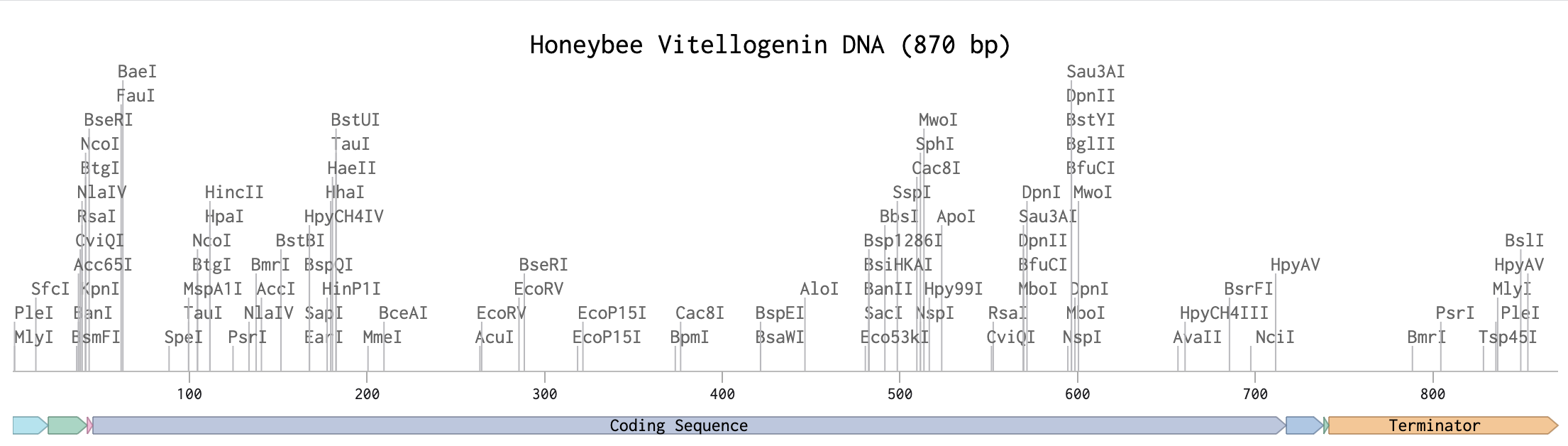
Since my project proposal from last week focuses on honeybee health, I searched for relevant proteins in Apis mellifera. During this process, I identified three candidates that seemed particularly interesting: Defensin-1, Hymenoptaecin and Vitellogenin. Working with Twist Bioscience’s codon optimization tool, I learned that the tool only accepts sequences within a specific length range — proteins that are too short or too long cannot be optimized. After several iterations, vitellogenin was the only protein for which I could successfully perform codon optimization. Vg, a phospholipoglycoprotein synthesized and stored in the honey bee fat body, is an ancient reproduction-associated protein that provides nutrients to eggs in most oviparous animals. Honey bee queens, who produce hundreds of eggs each day, have high levels of Vg gene expression. It is involved in nutrient storage, immune regulation and longevity in honeybees. Its expression is closely linked to colony health and higher vitellogenin levels are associated with improved immune responses and tolerance to Varroa destructor infestation. 1
A bit later, I found that University Münster actually proposed a similar study at iGEM: https://2023.igem.wiki/unimuenster/
Fasta File Text
XP_001122505.3 vitellogenin [Apis mellifera] MLVIILPYLLAARVPSHEATYRDDSDWRRYGPECTYDVLVNMSLSNMDEDARICSVIAFELKCRAKGSDTLNCRFSNGRTARLEDGRGCSNAKRNFAPSTSDRFVDEQPFEIRFNARGIENLVVSRDIARWRLDAMRAIVSQLNVGFELGSGHDRFVAMENSSVGYCEVEVKVSRAGYGGESGGGGLEIALEPERADVAPLSRGSVRIEKVRRPKRCPNRKIYFFGNHRDFSFGSEDIFMDMITSVSRMYISRREMNSFTESTGVMRTSNRPRTMNLHQRIGLSLRNINPARTPIPEIVNPASTSLYAYTNLERIPEYK
1: Amdam, G.V., Fennern, E., Havukainen, H. (2012). Vitellogenin in Honey Bee Behavior and Lifespan. In: Galizia, C., Eisenhardt, D., Giurfa, M. (eds) Honeybee Neurobiology and Behavior. Springer, Dordrecht. https://doi.org/10.1007/978-94-007-2099-2_2
2. Reverse Translate: Protein (amino acid) sequence to DNA (nucleotide) sequence.
GTTCTGTCCCGCATGAATCTTACGCTGGCGAAAATGGAGAAGACCAGCAAACCTCTGCCCATGGTTAACAATCCAGAATCGACTGGGAACCTCGTCTACATTTATTCGAACCCGTTTTCAGACGTAGAAGAGCGCCGCGTCAGTAAGACGGCTATGAATAGCAACCAAATTGTGTCGGACAACAGCCTATCAAGTTCTGAAGAAAAATTAAAACAGGATATCCTGAACTTACGGACAGATATCAGCAGCAGCTCCTCATCCATTAGTTCATCTGAGGAAAATGACTTCTGGCAGCCGAAACCCACCCTGGAGGATGCACCGCAGAATAGCTTGCTGCCTAATTTTGTTGGCTATAAAGGTAAACACATCGGTAAATCCGGAAAAGTGGATGTCATAAATGCAGCCAAGGAACTGATTTTCCAAATCGCCAACGAGCTCGAAGACGCTAGTAATATTCCAGTGCATGCGACGCTGGAAAAATTTATGATTCTGTGCAACCTTATGCGTACCATGAATCGTAAACAGATCAGCGAATTGGAATCTAACATGCAGATCTCGCCGAACGAATTAAAACCGAACGATAAATCTCAGGTGGTAAAGCAAAATACCTGGACCGTGTTTCGTGATGCGATTACACAGACCGGCACTGGCCCGGCCTTCCTGACGATTAAA
3. Codon optimization.
I chose E. coli because it is a standard lab organism, grows fast and widely used for protein production.
GTTCTGTCCCGCATGAACCTGACACTTGCAAAGATGGAAAAGACTAGTAAGCCGCTGCCCATGGTTAACAATCCAGAATCGACTGGGAACCTCGTCTACATTTATTCGAACCCGTTTTCAGACGTAGAAGAGCGCCGCGTCAGTAAGACGGCTATGAATAGCAACCAAATTGTGTCGGACAACAGCCTATCAAGTTCTGAAGAAAAATTAAAACAGGATATCCTGAACTTACGGACAGATATCAGCAGCAGCTCCTCATCCATTAGTTCATCTGAGGAAAATGACTTCTGGCAGCCGAAACCCACCCTGGAGGATGCACCGCAGAATAGCTTGCTGCCTAATTTTGTTGGCTATAAAGGTAAACACATCGGTAAATCCGGAAAAGTGGATGTCATAAATGCAGCCAAGGAACTGATTTTCCAAATCGCCAACGAGCTCGAAGACGCTAGTAATATTCCAGTGCATGCGACGCTGGAAAAATTTATGATTCTGTGCAACCTTATGCGTACCATGAATCGTAAACAGATCAGCGAATTGGAATCTAACATGCAGATCTCGCCGAACGAATTAAAACCGAACGATAAATCTCAGGTGGTAAAGCAAAATACCTGGACCGTGTTTCGTGATGCGATTACACAGACCGGCACTGGCCCGGCCTTCCTGACGATTAAA
4. You have a sequence! Now what?
Once the codon-optimized DNA sequence is obtained, it can be used to produce the protein through transcription and translation. In a cell-dependent system, the DNA is cloned into an expression vector, such as pET-21, and introduced into E. coli, where the bacterial machinery transcribes the DNA into mRNA and translates it into the vitellogenin protein. Alternatively, cell-free systems can carry out transcription and translation in vitro, using extracted enzymes and ribosomes without living cells. In both cases, the DNA sequence serves as a template that determines the amino acid sequence of the resulting protein. 2
I would use the cell-free mechanism “PUREexpress”. Vitellogenin is a very large protein, which can be difficult to express in living cells because of size, folding and potential toxicity. A reconstituted, cell‑free system like “PURExpress” provides a clean, RNase‑ and protease‑poor environment, so long mRNAs and large proteins are less likely to be degraded during expression.3
2: Claassens, N. J., Burgener, S., Vögeli, B., Erb, T. J., Bar-Even, A. (2019) A critical comparison of cellular and cell-free bioproduction systems, Current Opinion in Biotechnology, 60 (221-229) 3: Tuckey, C., Asahara, H., Zhou, Y., Chong, S. (2014) Protein synthesis using a reconstituted cell-free system. Curr Protoc Mol Biol, 108, doi: 10.1002/0471142727.mb1631s108
Part IV: Prepare a Twist DNA Synthesis Order
supportet by AI - “If you have a DNA strang how do you know which is what to annotate like: task instruction”
Part V: DNA Read/Write/Edit
1. What DNA would you want to sequence (e.g., read) and why? This could be DNA related to human health (e.g. genes related to disease research), environmental monitoring (e.g., sewage waste water, biodiversity analysis), and beyond (e.g. DNA data storage, biobank).
I would like to sequence Varroa mite DNA because Varroa destructor is a key global parasite of honey bees and a major cause of colony losses. Sequencing its genome and mitochondrial markers would help identify treatment‑resistance mutations, track the spread of different mite lineages between regions, and link mite genotypes to disease outcomes in colonies. This information can directly support better Varroa monitoring, more targeted control strategies, and breeding of honey bees that are more resilient to the specific Varroa populations in their environment, ultimately benefiting pollination, food security, and ecosystem health.4
4: Grindrod, I., Martin, SJ. (2021) Parallel evolution of Varroa resistance in honey bees: a common mechanism across continents? Proc Biol Sci, 288(1956), doi: 10.1098/rspb.2021.1375.
2. In lecture, a variety of sequencing technologies were mentioned. What technology or technologies would you use to perform sequencing on your DNA and why?
Chosen sequencing technology
I would use Illumina next-generation sequencing to analyze honeybee genes associated with Varroa mite resistance because it provides highly accurate and cost-efficient sequencing for comparing many samples or studying specific gene regions.5
5: Hu, T., Chitnis, N. , Monos, D., Dinh, A. (2021) Next-generation sequencing technologies: An overview, Human Immunology, 82(11), 801-811, https://doi.org/10.1016/j.humimm.2021.02.012.
Generation of technology
This method belongs to the second generation of sequencing technologies because it sequences millions of short DNA fragments in parallel, unlike first-generation Sanger sequencing or third-generation long-read single-molecule sequencing.
Input and preparation steps
- The input is genomic DNA extracted from honeybees or mites. Preparation involves:
- DNA extraction from samples
- DNA fragmentation into short pieces
- Adapter ligation to fragment ends
- PCR amplification of fragments
- Loading fragments onto a sequencing flow cell
How bases are decoded (sequencing principle)
Fragments bind to the flow cell and are amplified into clusters. During sequencing, fluorescently labeled nucleotides are incorporated one base at a time. A camera records the color signal after each cycle, and software converts these signals into DNA base sequences — this process is called base calling.
Output of sequencing
The output consists of millions of short DNA reads, typically stored in FASTQ files containing:
- DNA sequences
- quality scores for each base
These reads are then assembled or mapped to a reference genome to analyze genetic variation related to disease resistance.
3. What DNA would you want to synthesize (e.g., write) and why? These could be individual genes, clusters of genes or genetic circuits, whole genomes, and beyond. As described in class thus far, applications could range from therapeutics and drug discovery (e.g., mRNA vaccines and therapies) to novel biomaterials (e.g. structural proteins), to sensors (e.g., genetic circuits for sensing and responding to inflammation, environmental stimuli, etc.), to art (DNA origamis). If possible, include the specific genetic sequence(s) of what you would like to synthesize! You will have the opportunity to actually have Twist synthesize these DNA constructs!
For my project, I would like to synthesize DNA that enables the production of a honeybee protein relevant to resistance against Varroa mite infection, specifically a codon-optimized fragment of the vitellogenin gene for expression in E. coli. Producing this protein in a laboratory system would allow further investigation of its structure and function and could support future research on improving honeybee resilience, which is crucial for pollination, biodiversity, and food production.
4. What technology or technologies would you use to perform this DNA synthesis and why? What are the essential steps of your chosen sequencing methods? What are the limitations of your sequencing method (if any) in terms of speed, accuracy, scalability?
Modern DNA synthesis relies on chemical oligonucleotide synthesis combined with enzymatic assembly. Short DNA fragments are chemically synthesized and then assembled into longer genes. This method is efficient, scalable, and allows full customization of DNA sequences, including codon optimization and removal of unwanted restriction sites.
Essential steps of DNA synthesis
Digital sequence design of the gene or construct. Chemical synthesis of short DNA oligonucleotides. Assembly of oligos into longer DNA fragments using enzymatic methods. Error correction and amplification of assembled fragments. Cloning into plasmids and propagation in bacteria. Sequence verification to confirm correctness before delivery.
Limitations of this method
- Speed: Gene synthesis can take days to weeks depending on sequence length and complexity.
- Accuracy: Errors can occur during synthesis or assembly, especially in repetitive or GC-rich sequences, requiring verification and correction.
- Scalability: Although modern platforms are highly scalable, very long DNA constructs or entire genomes remain costly and technically challenging.
- Sequence constraints: Certain sequences (e.g., strong repeats or toxic genes) can be difficult to synthesize or maintain in host organisms.
5. What DNA would you want to edit and why? In class, George shared a variety of ways to edit the genes and genomes of humans and other organisms. Such DNA editing technologies have profound implications for human health, development, and even human longevity and human augmentation. DNA editing is also already commonly leveraged for flora and fauna, for example in nature conservation efforts, (animal/plant restoration, de-extinction), or in agriculture (e.g. plant breeding, nitrogen fixation). What kinds of edits might you want to make to DNA (e.g., human genomes and beyond) and why?
Once again I would want to edit DNA related to honeybee health, specifically genes that contribute to resistance against Varroa mite infestation. Potential edits could focus on genes involved in immune response, grooming behavior, or parasite detection, enhancing bees’ natural ability to remove mites or better tolerate infections transmitted by them. Instead of introducing entirely new traits, the goal would be to support or amplify naturally occurring resistance traits, similar to selective breeding but with greater precision. More broadly, responsible DNA editing could also be applied in agriculture and conservation to help organisms adapt to climate change, reduce pesticide use, and improve resilience in vulnerable ecosystems.
6. What technology or technologies would you use to perform these DNA edits and why? How does your technology of choice edit DNA? What are the essential steps? What preparation do you need to do (e.g. design steps) and what is the input (e.g. DNA template, enzymes, plasmids, primers, guides, cells) for the editing? What are the limitations of your editing methods (if any) in terms of efficiency or precision?
I would use CRISPR–Cas9, which allows precise modification of genes within an organism’s genome. This technology is widely used because it is relatively simple, efficient and adaptable to many organisms.
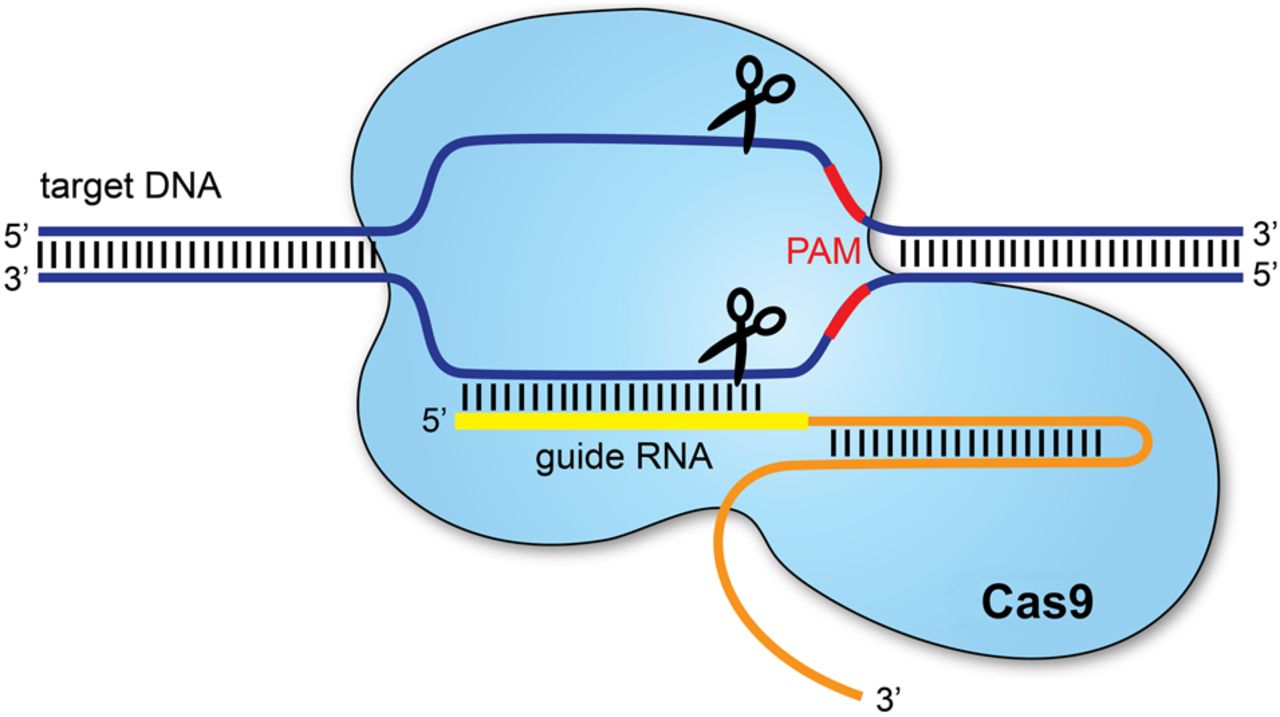

CRISPR–Cas9 edits DNA by using a guide RNA (gRNA) that directs the Cas9 enzyme to a specific DNA sequence. Cas9 then creates a cut at the targeted location. The cell’s natural DNA repair mechanisms repair this cut, and during repair, scientists can either disable a gene or insert a modified DNA sequence.
- Essential editing steps include:
- Designing a guide RNA targeting the gene of interest.
- Delivering the guide RNA and Cas9 enzyme into cells.
- Cas9 cutting the DNA at the chosen site.
- Cellular repair mechanisms introducing deletions, modifications, or inserting new DNA.
- Screening cells or organisms to confirm successful edits.
Preparation and required inputs
Before editing, we must design the genetic modification and ensure the target gene is well characterized. Required inputs typically include:
- Guide RNA sequences targeting the gene
- Cas9 enzyme or Cas9-encoding plasmid
- A donor DNA template if inserting new sequences
- Delivery system (e.g., plasmids, viral vectors, or microinjection)
- Target cells or embryos to be edited
Limitations
The main limitations of CRISPR/Cas9 relate to delivery, accuracy and ethical concerns. A major challenge is safely delivering the editing system into the correct cells in living organisms, as current delivery vectors have size or efficiency limitations. Another concern is off-target effects, where unintended parts of the genome may be edited, potentially causing harmful consequences such as cancer. Editing efficiency can also vary, meaning not all cells receive the desired modification. Additionally, editing germline cells or embryos raises significant ethical and long-term safety concerns, since changes would be inherited by future generations and their consequences are uncertain. 6
6: Redman, M., King, A., Watson, C., King, D. (2016) What is CRISPR/Cas9?, Archives of Diseases in Childhood, 101, 213–215. doi:10.1136/archdischild-2016-310459