Week 3 HW: Lab Automation
Part I: Python Script for Opentrons Artwork
Your task this week is to Create a Python file to run on an Opentrons liquid handling robot.
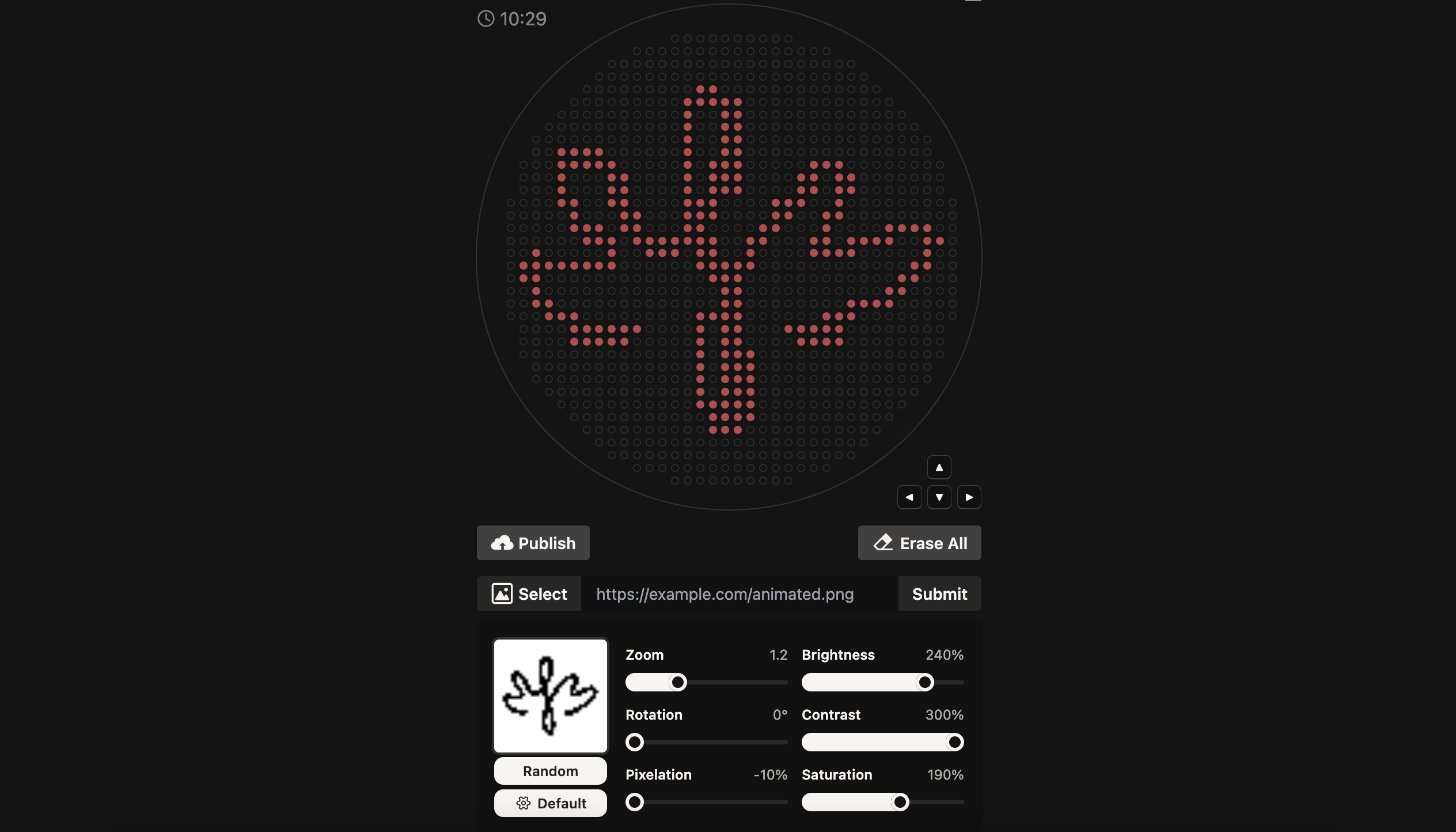

Firstly, I used Ronan’s Automation Art Interface to translate my logo into a pixelated biological artwork. The software converted the image into a set of coordinate outputs, where each tuple (x, y) represents the precise millimeter offset from the calibrated center of the agar plate. Each of these coordinate pairs defines the placement of a single 1 µL droplet, allowing the robot to reconstruct the digital logo physically on the plate.
I then transferred these coordinates into Google Colab, where I programmed the Opentrons protocol. Before beginning the patterning process, the robot picks up a single 20 µL pipette tip. Since the entire artwork is executed in one color, only one tip is required for the whole procedure, minimizing material use.
The main challenge was preventing the pipette from exceeding its 20 µL capacity. Because each droplet dispenses 1 µL and the artwork consists of many coordinate points, the system must repeatedly refill the pipette. However, simply aspirating 20 µL at fixed intervals can lead to overfilling if residual liquid remains inside the tip.
To solve this, I implemented a volume-tracking mechanism in the code with support of AI (ChatGPT). A variable continuously monitors the remaining liquid in the pipette. The robot only aspirates when the remaining volume reaches zero, and it calculates the exact amount needed—either the full 20 µL capacity or just the remaining volume required to complete the artwork. After each 1 µL dispense, the tracked volume is reduced accordingly. This ensures that the pipette never exceeds its capacity while allowing the artwork to be executed seamlessly and efficiently.
Post-Lab Questions
Find and describe a published paper that utilizes the Opentrons or an automation tool to achieve novel biological applications.
A suitable published paper that utilizes an automation tool for a novel biological application is:
DeRoo, J. B., Jones, A. A., Caroline K. Slaughter, C. K., Ahr, T. W., Stroup, S. M., Thompson, G. B., Snow, C. D. (2025) Automation of protein crystallization scaleup via Opentrons-2 liquid handling. SLAS Technology, 32, https://doi.org/10.1016/j.slast.2025.100268
This study employs the Opentrons OT-2 automated liquid handling robot to optimize protein crystallization workflows. Protein crystallization is a critical step in structural biology, particularly for determining protein structures via X-ray crystallography. Traditionally, crystallization screening requires extensive manual pipetting, which is time-consuming, error-prone, and difficult to standardize.
The novelty of the paper lies in:
Low-cost automation of crystallization screening The researchers adapted the open-source OT-2 robot to perform precise nanoliter- to microliter-scale liquid handling for setting up crystallization trials. This democratizes access to automation, as most conventional crystallization robots are prohibitively expensive.
Workflow optimization and reproducibility The study demonstrates how automated pipetting improves reproducibility and throughput compared to manual methods. It allows systematic variation of crystallization conditions (e.g., precipitant concentration, pH, additives) in a controlled and programmable manner.
Open-source customization A key contribution is the development of customizable protocols that other laboratories can replicate and modify. The paper highlights how open-source hardware and software can accelerate biological research innovation without reliance on proprietary systems.
Biological Impact
By automating crystallization optimization:
- Screening becomes more efficient and scalable.
- Experimental variability is reduced.
- Smaller laboratories gain access to structural biology techniques that were previously limited to well-funded institutions.
Write a description about what you intend to do with automation tools for your final project. You may include example pseudocode, Python scripts, 3D printed holders, a plan for how to use Ginkgo Nebula, and more. You may reference this week’s recitation slide deck for lab automation details.
Rather than relying on manual trial-and-error biology, I aim to build programmable experimental pipelines using robotic liquid handling, custom labware, and computational control for my final project. I intend to use modular lab automation tools to develop reproducible, scalable biological systems.
How automation would be used
High-Throughput Construct Assembly
Using a modular liquid handler such as the Opentrons OT-2:
- Automated Golden Gate or Gibson assemblies
- Parallel plasmid construction
- Controlled transformation setups
- Replicate culture inoculations
Example pseudocode:
for construct in plasmid_library: pipette.transfer(2, promoter_plate[construct], assembly_well) pipette.transfer(2, gene_plate[construct], assembly_well) pipette.transfer(6, master_mix, assembly_well)
This would allow rapid screening of different antimicrobial peptide constructs, dsRNA delivery systems, or immune-modulating pathways.
Custom 3D-Printed Bee Microbiome Holders
I would design:
- 3D-printed micro-incubation chambers
- Gut-simulating microfluidic inserts
- Controlled feeding modules
Ginkgo Nebula Integration
Through Ginkgo Bioworks’s Nebula platform, I could:
- Analyze microbiome sequencing data
- Identify candidate symbionts
- Screen gene clusters linked to antimicrobial production
- Compare engineered vs natural strain performance
To put it in a nutshell, lab automation transforms these projects from speculative ideas into experimentally rigorous, iterative engineering systems.
Part III: Final Project Ideas
As explained in this week’s recitation, add 1-3 slides in your Node’s section of this slide deck with 3 ideas you have for an Individual Final Project. Be sure to put your name, city, and country on your slide!


