Week 10 HW: Imaging and Measurement
Part I: Final Project
For your final project:
1. Please identify at least one (ideally many) aspect(s) of your project that you will measure. It could be the mass or sequence of a protein, the presence, absence, or quantity of a biomarker, etc.
- Quantification of pigment concentration through color intensity measurements
- Analysis of pigment degradation (as a proxy for biochemical stability) under environmental conditions
- Material–pigment interaction effects on color retention
2. Please describe all of the elements you would like to measure, and furthermore describe how you will perform these measurements.
Pigment production (via color intensity as proxy) I will assess whether and to what extent pigments are successfully expressed in fibers by quantifying color intensity. Since direct biochemical quantification may not be feasible, color measurements (RGB or CIELAB values) obtained through standardized imaging will serve as a proxy for pigment concentration. This allows comparison between different pigment systems (e.g. anthocyanins vs. betalains) or expression strategies.
Pigment stability (biochemical and environmental stability) To evaluate whether biologically produced pigments are viable for textile use, I will analyze their stability under environmental stressors such as light exposure, washing, and pH variation. Changes in color intensity and hue over time will indicate degradation rates and overall pigment robustness.
Material integration (fiber-specific color retention) A key aspect of the project is whether pigments remain stable and visible within fiber structures. I will therefore evaluate how pigments perform within different fiber types (or simulated substrates), focusing on color retention, uniformity, and resistance to fading. This reflects how effectively pigments are integrated into the material during or after fiber formation.
3. What are the technologies you will use (e.g., gel electrophoresis, DNA sequencing, mass spectrometry, etc.)? Describe in detail.
Bioinformatics and pathway design tools To identify suitable pigment systems, I will use bioinformatics databases and literature-based analysis to select and map biosynthetic pathways (e.g. anthocyanin or betalain pathways). This includes identifying key enzymes and exploring strategies for fiber-specific expression.
Digital color quantification Color intensity will be measured using a standardized photographic setup and analyzed with software such as ImageJ. Extracted RGB or CIELAB values will provide quantitative data for comparing pigment expression and stability.
Spectrophotometric analysis Spectrophotometry will be used to measure absorbance or reflectance of fibers, allowing more precise quantification of pigment presence and optical properties.
Environmental testing protocols To simulate real-world conditions, samples will be exposed to controlled stress factors such as UV/light exposure, washing cycles, and pH variation. These tests will help evaluate pigment durability and material performance.
Waters Part I — Molecular Weight
1. Based on the predicted amino acid sequence of eGFP (see below) and any known modifications, what is the calculated molecular weight? You can use an online calculator like the one at https://web.expasy.org/compute_pi/
Theoretical isoelectric point (pI): 5.90 Theoretical Molecular weight (Mw): 28006.60 Da
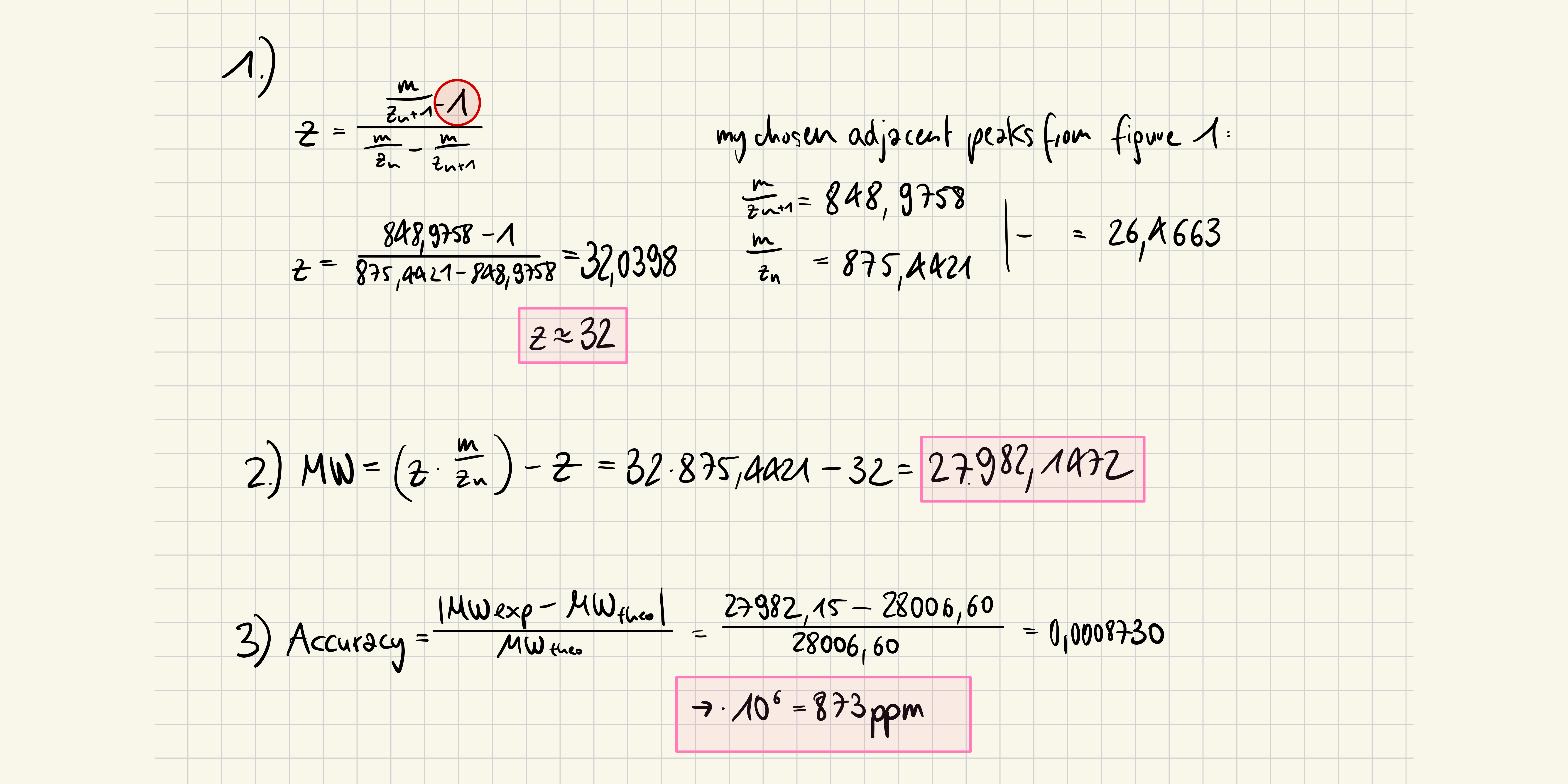
2. Calculate the molecular weight of the eGFP using the adjacent charge state approach described in the recitation. Select two charge states from the intact LC-MS data (Figure 1) and: 2.1 Determine z for each adjacent pair of peaks (n, n+1): 2.2 Determine the MW of the protein: 2.3 Calculate the accuracy of the measurement using the deconvoluted MW from 2.2 and the predicted weight of the protein from 2.1


3. Can you observe the charge state for the zoomed-in peak in the mass spectrum for the intact eGFP? If yes, what is it? If no, why not?
No, since the protein is in a larger charged state than the mass spectrometer can catch. It isn’t visible.
Waters Part II — Secondary/Tertiary structure
not required
Waters Part III — Peptide Mapping - primary structure
1. How many Lysines (K) and Arginines (R) are in eGFP? Please circle or highlight them in the eGFP sequence given in Waters Part I question 1 above. (Note: adding the sequence to Benchling as an amino acid file and clicking biochemical properties tab will show you a count for each amino acid).
MVSKGEELFTG VVPILVELDG DVNGHKFSVS GEGEGDATYG KLTLKFICTT GKLPVPWPTL VTTLTYGVQC FSRYPDHMKQ HDFFKSAMPE GYVQERTIFF KDDGNYKTRA EVKFEGDTLV NRIELKGIDF KEDGNILGHK LEYNYNSHNV YIMADKQKNG IKVNFKIRHN IEDGSVQLAD HYQQNTPIGD GPVLLPDNHY LSTQSALSKD PNEKRDHMVL LEFVTAAGIT LGMDELYKLE HHHHHH
Lysines (K): 19
MVSKGEELFTG VVPILVELDG DVNGHKFSVS GEGEGDATYG KLTLKFICTT GKLPVPWPTL VTTLTYGVQC FSRYPDHMKQ HDFFKSAMPE GYVQERTIFF KDDGNYKTRA EVKFEGDTLV NRIELKGIDF KEDGNILGHK LEYNYNSHNV YIMADKQKNG IKVNFKIRHN IEDGSVQLAD HYQQNTPIGD GPVLLPDNHY LSTQSALSKD PNEKRDHMVL LEFVTAAGIT LGMDELYKLE HHHHHH
Arginines (R): 6
2. How many peptides will be generated from tryptic digestion of eGFP? Navigate to https://web.expasy.org/peptide_mass/ Copy/paste the sequence above into the input box in the PeptideMass tool to generate expected list of peptides. Use Figure 4 below as a guide for the relevant parameters to predict peptides from eGFP. Click “Perform the Cleavage” button in the PeptideMass tool and report the number of peptides generated when using trypsin to perform the digest.
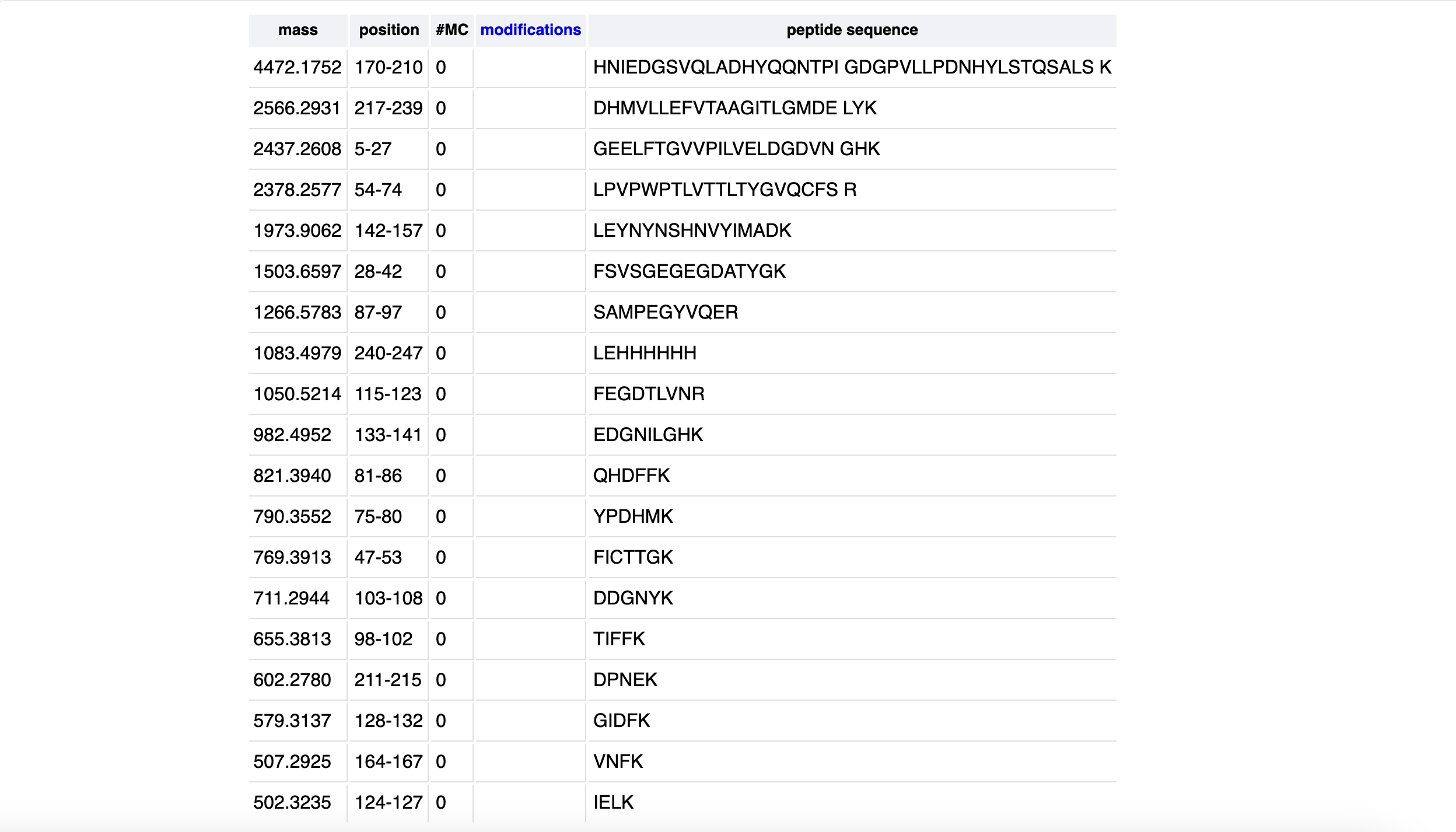

It reports 19 peptides.
3. Based on the LC-MS data for the Peptide Map data generated in lab (please use Figure 5a as a reference) how many chromatographic peaks do you see in the eGFP peptide map between 0.5 and 6 minutes? You may count all peaks that are >10% relative abundance.
According to Figure 5a, there are 22 peaks: 0.43, 0.61, 0.79, 1.20, 1.43, 1.80, 1.85, 1.93, 2.17, 2.26, 2.54, 2.78, 3.27, 3.53, 3.59, 3.70, 4.30, 4.48, 4.64, 4.87, 5.06, 5.43
4. Assuming all the peaks are peptides, does the number of peaks match the number of peptides predicted from question 2 above? Are there more peaks in the chromatogram or fewer?
There are more peaks in the chromatogram. -> 3
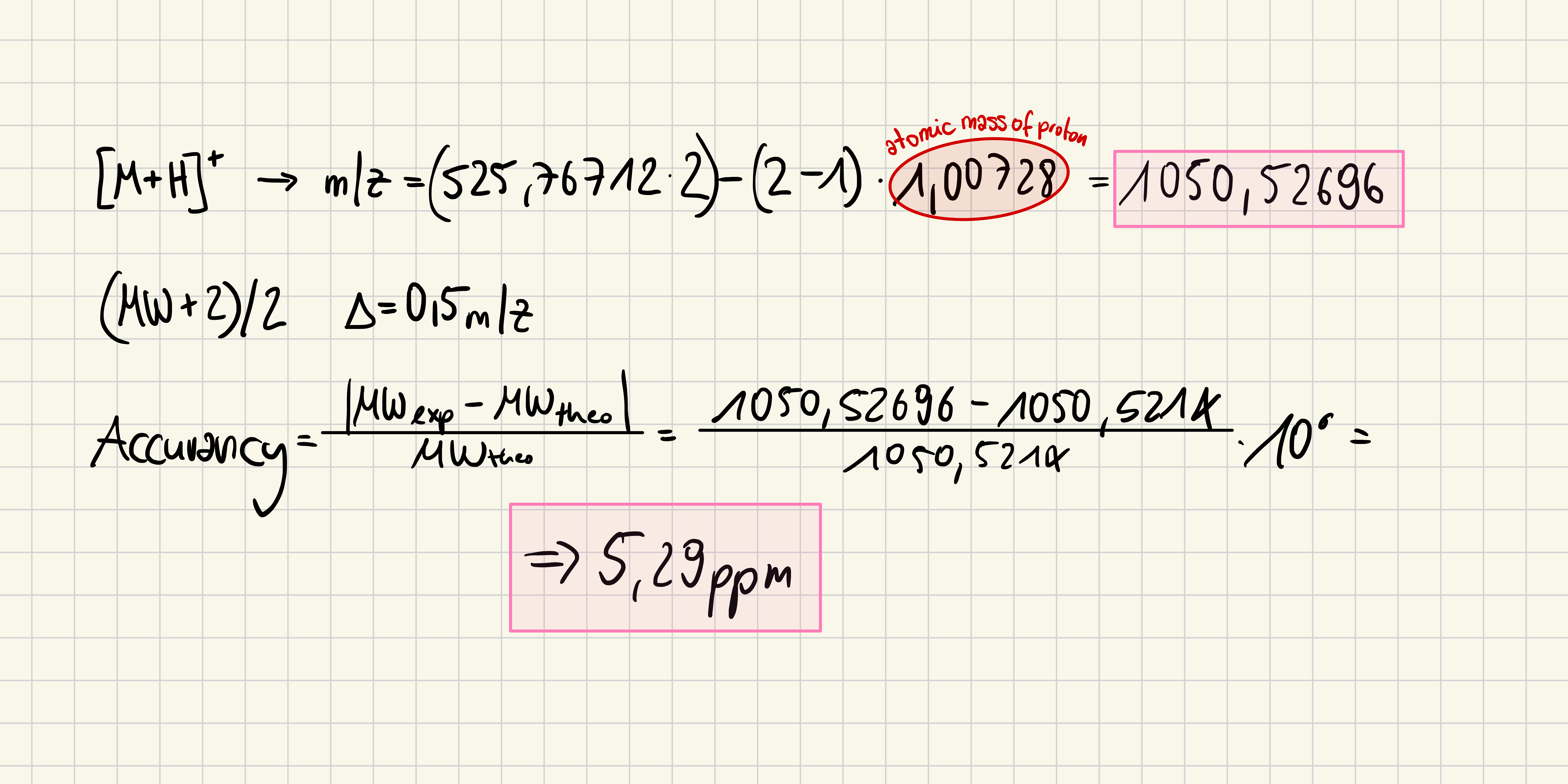
5. Identify the mass-to-charge of the peptide shown in Figure 5b. What is the charge () of the most abundant charge state of the peptide (use the separation of the isotopes to determine the charge state). Calculate the mass of the singly charged form of the peptide
526,25918 - 525,76712 = 0.49206 ~ 0.5
6. Identify the peptide based on comparison to expected masses in the PeptideMass tool. What is mass accuracy of measurement? Please calculate the error in ppm.
FEGDTLVNR


7. What is the percentage of the sequence that is confirmed by peptide mapping? (see Figure 6)
90.7% of sequence covered
Waters Part IV — Oligomers
1. We will determine Keyhole Limpet Hemocyanin (KLH)’s oligomeric states using charge detection mass spectrometry (CDMS). CDMS single-particle measurements of KLH allow us to make direct mass measurements to determine what oligomeric states (that is, how many protein subunits combine) are present in solution. Using the known masses of the polypeptide subunits (Table 1) for KLH, identify where the following oligomeric species are on the spectrum shown below from the CDMS (Figure 7):
7FU Decamer → 10 x 340 kDa = 3,400 kDa = 3.4 MDa
8FU Didecamer → 20 × 400 kDa = 8,000 kDa = 8.0 MDa
8FU 3-Decamer → 30 × 400 kDa = 12,000 kDa = 12 MDa
8FU 4-Decamer → 40 × 400 kDa = 16,000 kDa = 16 MDa
Regarding the peaks in Figure 7, I see peaks at 3.4 MDa, 8.33 MDa, 12.67 MDa and a smaller one close to 16-17 MDa.
Waters Part V — Did I make GFP?
1. Please fill out this table with the data you acquired from the lab work done at the Waters Immerse Lab in Cambridge, or else the data screenshots in this document if you were unable to have lab work done at Waters.