Week 2 HW: DNA r/w/e
Part 1: Benchling & In-silico Gel Art:
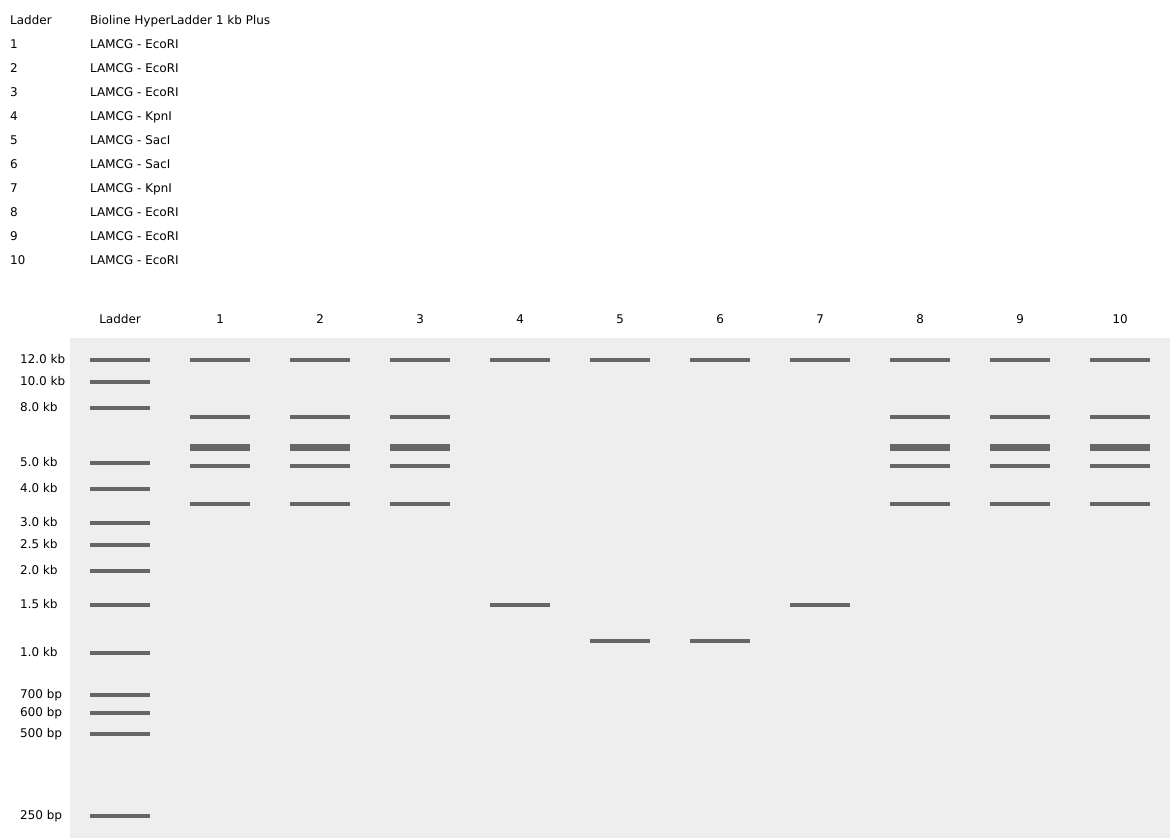
Simulate Restriction Enzyme Digestion with the following Enzymes:
- EcoRI
- HindIII
- BamHI
- KpnI
- EcoRV
- SacI
- SalI
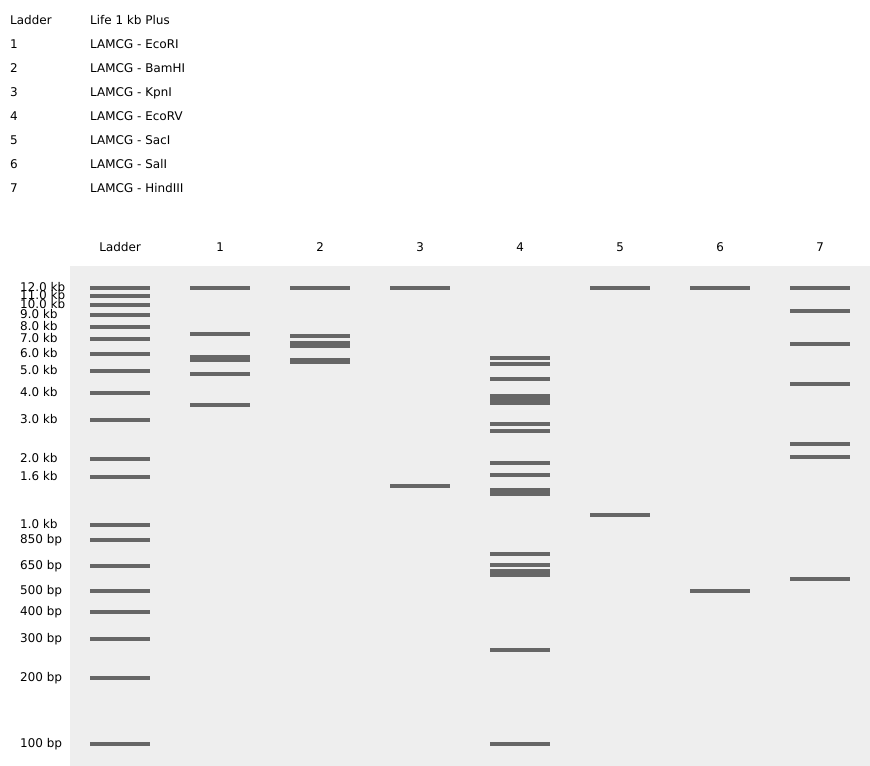


- Create a pattern/image in the style of Paul Vanouse’s Latent Figure Protocol artworks.
It’s a smiley face!! :)

Part 3: DNA Design Challenge:
3.1 Chose your protein
Alkaline phosphatase. I chose this protein because it plays because it is important for nutrient sensing, as it is naturally activated under conditions of phosphate limitation. In Escherichia coli, alkaline phosphatase (PhoA) hydrolyzes organic phosphate compounds to release inorganic phosphate, directly linking environmental signals to nutrient availability. Additionally, it is a well-characterized enzyme with a resolved structure and extensively documented sequence information, making it an ideal model protein for computational and experimental analysis.
Sequence: (P00634)MKQSTIALALLPLLFTPVTKARTPEMPVLENRAAQGDITAPGGARRLTGDQTAALRDSLSDKPAKNIILLIGDGMGDSEITAARNYAEGAGGFFKGIDALPLTGQYTHYALNKKTGKPDYVTDSAASATAWSTGVKTYNGALGVDIHEKDHPTILEMAKAAGLATGNVSTAELQDATPAALVAHVTSRKCYGPSATSEKCPGNALEKGGKGSITEQLLNARADVTLGGGAKTFAETATAGEWQGKTLREQAQARGYQLVSDAASLNSVTEANQQKPLLGLFADGNMPVRWLGPKATYHGNIDKPAVTCTPNPQRNDSVPTLAQMTDKAIELLSKNEKGFFLQVEGASIDKQDHAANPCGQIGETVDLDEAVQRALEFAKKEGNTLVIVTADHAHASQIVAPDTKAPGLTQALNTKDGAVMVMSYGNSEEDSQEHTGSQLRIAAYGPHAANVVGLTDQTDLFYTMKAALGLK
3.2 Reverse Translate:atgaaacagagcaccattgcgctggcgctgctgccgctgctgtttaccccggtgaccaaa gcgcgcaccccggaaatgccggtgctggaaaaccgcgcggcgcagggcgatattaccgcg ccgggcggcgcgcgccgcctgaccggcgatcagaccgcggcgctgcgcgatagcctgagc gataaaccggcgaaaaacattattctgctgattggcgatggcatgggcgatagcgaaatt accgcggcgcgcaactatgcggaaggcgcgggcggcttttttaaaggcattgatgcgctg ccgctgaccggccagtatacccattatgcgctgaacaaaaaaaccggcaaaccggattat gtgaccgatagcgcggcgagcgcgaccgcgtggagcaccggcgtgaaaacctataacggc gcgctgggcgtggatattcatgaaaaagatcatccgaccattctggaaatggcgaaagcg gcgggcctggcgaccggcaacgtgagcaccgcggaactgcaggatgcgaccccggcggcg ctggtggcgcatgtgaccagccgcaaatgctatggcccgagcgcgaccagcgaaaaatgc ccgggcaacgcgctggaaaaaggcggcaaaggcagcattaccgaacagctgctgaacgcg cgcgcggatgtgaccctgggcggcggcgcgaaaacctttgcggaaaccgcgaccgcgggc gaatggcagggcaaaaccctgcgcgaacaggcgcaggcgcgcggctatcagctggtgagc gatgcggcgagcctgaacagcgtgaccgaagcgaaccagcagaaaccgctgctgggcctg tttgcggatggcaacatgccggtgcgctggctgggcccgaaagcgacctatcatggcaac attgataaaccggcggtgacctgcaccccgaacccgcagcgcaacgatagcgtgccgacc ctggcgcagatgaccgataaagcgattgaactgctgagcaaaaacgaaaaaggctttttt ctgcaggtggaaggcgcgagcattgataaacaggatcatgcggcgaacccgtgcggccag attggcgaaaccgtggatctggatgaagcggtgcagcgcgcgctggaatttgcgaaaaaa gaaggcaacaccctggtgattgtgaccgcggatcatgcgcatgcgagccagattgtggcg ccggataccaaagcgccgggcctgacccaggcgctgaacaccaaagatggcgcggtgatg gtgatgagctatggcaacagcgaagaagatagccaggaacataccggcagccagctgcgc attgcggcgtatggcccgcatgcggcgaacgtggtgggcctgaccgatcagaccgatctg ttttataccatgaaagcggcgctgggcctgaaa
3.3 Codon optimization
Codon optimization is important because organisms differ on the way the use certain codons. We know the genetic code is universal and degenerate, meaning that one aminoacid can be encoded by multiple codons, but different organisms do not use these synonymous codons with the same frequency. This phenomenon, known as codon usage bias, affects how efficiently a gene is translated. If a DNA sequence contains codons that are rare in the host organism, the corresponding tRNAs may be scarce, which can slow down translation, reduce protein yield, or even affect proper folding. Therefore codon optimization adjust the sequence to match the codons preferred by the organism, therefore improving the yield and overall expression.
For this project, I chose to optimize the codon sequence for Escherichia coli because it is one of the most widely used host organisms for recombinant protein expression. It grows rapidly, is inexpensive to culture, has well-characterized genetics, and many expression vectors are available. Since alkaline phosphatase (PhoA) is naturally found in E. coli, optimizing for this host ensures efficient production while maintaining compatibility with standard molecular biology workflows.
I used *VectorBuilder’s Codon Optimization Tool.
Result: Pasted Sequence: GC=61.29%, CAI=1.00 // Improved DNA[1]: GC=59.38%, CAI=0.93
Sequence: ATGAAACAGAGCACCATTGCGCTGGCGCTGCTGCCGCTGCTGTTTACCCCGGTCACGAAAGCGCGCACCCCGGAAATGCCGGTGCTGGAAAACCGCGCAGCACAGGGTGATATCACCGCGCCGGGCGGTGCGCGCCGCCTGACCGGCGATCAGACCGCGGCCCTGCGCGATAGCCTGAGCGATAAACCGGCGAAAAATATTATTCTGCTGATTGGCGACGGTATGGGCGATAGCGAAATCACCGCCGCGCGCAATTATGCGGAAGGCGCCGGTGGCTTTTTTAAAGGTATCGATGCGCTGCCGCTGACCGGCCAGTACACCCACTACGCGCTGAATAAAAAAACTGGTAAACCGGATTATGTCACCGATAGTGCGGCCAGCGCAACCGCGTGGAGCACCGGCGTGAAAACCTACAATGGCGCGCTGGGCGTGGATATTCATGAAAAAGATCACCCGACGATTCTGGAAATGGCGAAAGCGGCCGGCCTGGCGACCGGCAATGTGAGCACCGCGGAACTGCAGGATGCCACCCCGGCGGCGCTGGTGGCCCATGTGACCAGCCGTAAATGCTATGGTCCGAGCGCGACCAGCGAAAAATGTCCGGGCAACGCGCTGGAAAAAGGTGGCAAAGGCAGCATTACCGAACAGCTGCTGAACGCGCGTGCCGATGTGACCCTGGGCGGAGGCGCAAAAACCTTTGCCGAAACCGCGACCGCGGGCGAATGGCAGGGCAAAACCCTGCGCGAACAGGCGCAGGCCCGCGGTTATCAGCTGGTTAGCGATGCGGCCAGCCTGAATAGCGTGACCGAAGCGAACCAGCAGAAACCGCTGCTGGGCCTGTTTGCGGATGGTAATATGCCGGTGCGCTGGCTGGGTCCGAAAGCGACCTATCACGGTAACATTGATAAACCGGCGGTGACCTGCACCCCGAACCCGCAGCGCAACGATAGCGTGCCGACCCTGGCACAGATGACCGATAAAGCCATTGAACTGCTGAGCAAAAATGAAAAAGGCTTTTTCCTGCAGGTGGAAGGCGCGTCCATTGATAAACAGGATCACGCAGCCAACCCGTGTGGCCAGATTGGCGAAACCGTGGATCTGGATGAAGCGGTGCAGCGTGCCCTGGAATTTGCGAAAAAAGAAGGTAACACCCTGGTGATTGTGACCGCAGACCATGCGCATGCGAGCCAGATCGTGGCGCCGGATACCAAAGCGCCGGGTCTGACCCAAGCGTTGAATACCAAAGATGGTGCGGTGATGGTGATGAGCTATGGCAACAGCGAGGAAGATAGCCAGGAACACACCGGCAGTCAGCTGCGTATTGCCGCATACGGCCCGCATGCGGCGAACGTGGTGGGCCTGACCGATCAGACCGACCTGTTTTACACCATGAAAGCGGCACTGGGCCTGAAA
3.4. You have a sequence! Now what?
Once the optimized DNA sequence is obtained, it can be cloned into an expression plasmid containing a promoter, ribosome binding site, and selection marker, and then introduced into a host such as Escherichia coli through transformation. Inside the cell, the promoter drives transcription of the DNA into mRNA by RNA polymerase, and ribosomes bind to the mRNA to translate its codons into the corresponding amino acid sequence, producing the alkaline phosphatase protein, which then folds into its functional structure. This is the cell-dependent method.
Part 4: Prepare a Twist DNA Synthesis Order:
I attempted to place the Twist DNA synthesis order, but the platform did not function properly. Although I was able to log in, a message stating “Contact your Distributor” repeatedly appeared, and the system automatically logged me out, preventing me from proceeding with the order. I am aware that other students in my region (Brazil/Latin America) experienced similar difficulties accessing the platform.
Part 5: DNA Read/Write/Edit:
DNA Read
(i) What DNA would you want to sequence (e.g., read) and why?
I would sequence soil metagenomic DNA from the sugarcane rhizosphere (and bulk soil as a control) to understand which microbial communities and genes are present that control nutrient cycling. In particular, I’d look for taxa and functional genes involved in phosphorus solubilization/mineralization (e.g., phosphatases), nitrogen transformations (nitrification/denitrification), and other pathways that determine whether nutrients are retained, lost, or made bioavailable. This would let me link nutrient status (low/high P or N) to shifts in microbial composition and metabolic potential.
(ii) In lecture, a variety of sequencing technologies were mentioned. What technology or technologies would you use to perform sequencing on your DNA and why? I would use Illumina shotgun metagenomic sequencing, which is a second-generation technology because it performs massively parallel sequencing of millions of short DNA fragments using sequencing-by-synthesis chemistry. The input would be total DNA extracted from soil samples, which would be fragmented, end-repaired, ligated to sequencing adapters (with barcodes if multiplexing), PCR-amplified if necessary, and loaded onto a flow cell to generate clusters. During sequencing, fluorescently labeled nucleotides are incorporated one at a time, and imaging after each cycle detects the emitted fluorescence to determine the identity of each base (base calling). The output consists of millions of short reads in FASTQ format, containing both nucleotide sequences and quality scores, which can then be analyzed to identify microbial taxa and nutrient-cycling genes.
DNA Write
(i) What DNA would you want to synthesize (e.g., write) and why?
I would like to synthesize a nutrient-responsive genetic circuit for soil bacteria that turns nutrient mobilization “on” only when the environment is nutrient-limited—for example, a phosphate-starvation–responsive promoter (Pho regulon) driving a codon-optimized alkaline phosphatase (PhoA) module to help mineralize organic phosphate into bioavailable phosphate. Concretely, the construct would look like [P_Pho]–RBS–phoA–terminator, where the PhoA protein sequence can be sourced from UniProt P00634 and the Pho regulon is sourced from the well-characterized PhoB/PhoR phosphate-sensing regulatory system in bacteria.
(ii) What technology or technologies would you use to perform this DNA synthesis and why?
I would use commercial gene synthesis based On sdlid-phase phosphoramidite oligonucleotide synthesis followed by DNA assembly (like Gibson Assembly,as I think we are going to learn in HTGAA) because this is the most reliable way to “write” a custom genetic circuit with a good sequence control. In practice, short oligos are chemically synthesized cycle-by-cycle, then cleaved/deprotected and purified, and multiple fragments are assembled into the full construct (they are oftten sequneced afterward, to check if everything is ok). The main limitations are that error rates and cost increase with sequence length, as long constructs require multi-fragment assembly and verification.
DNA Edit
(i) What DNA would you want to edit and why?
I would want to edit the genomes of plant-associated soil bacteria to enhance their ability to sense nutrient limitation and respond in a controlled, beneficial way. Specifically, I would modify regulatory regions of genes involved in phosphate metabolismto fine-tune their sensitivity and dynamic range.
(ii) What technology or technologies would you use to perform these DNA edits and why?
I would use CRISPR-Cas9 genome editing because it is the standard technique and it allows precise modification of specific DNA sequences in bacteria. CRISPR edits DNA by using a designed guide RNA (gRNA) that directs the Cas9 nuclease to a complementary target sequence, where Cas9 introduces a double-strand break; the cell then repairs this break either through non-homologous end joining (introducing small insertions or deletions) or through homology-directed repair if a donor DNA template is provided, enabling precise edits. Preparation involves designing the guide RNA sequence, constructing or obtaining a plasmid encoding Cas9 and the gRNA, optionally designing a repair template with homology arms, and delivering these components into the target bacterial cells (e.g., via transformation or electroporation). Limitations include variable editing efficiency depending on the organism, potential off-target effects if guide design is imperfect, lower efficiency of homology-directed repair in some bacteria, and challenges in delivering editing machinery into non-model environmental strains.