Week 5 HW: Protein Design Part II
Part A: SOD1 Binder Peptide Design (From Pranam)
Part 1: Generate Binders with PepMLM
I first generated the mutated sequence: SOD1 A4VMATK**V**VCVLKGDGPVQGIINFEQKESNGPVKVWGSIKGLTEGLHGFHVHEFGDNTAGCTSAGPHFNPLSRKHGGPKDEERHVGDLGNVTADKDGVADVSIEDSVISLSGDHCIIGRTLVVHEKADDLGKGGNEESTKTGNAGSRLACGVIGIAQ
I then used the PepMLM-650M notebook to generate peptide binders conditioned on the mutant SOD1 sequence. I set the peptide length to 12 amino acids and generated candidate binders from the model output. The resulting peptides and their pseudo-perplexity values are shown below.
| Binder | Pseudo Perplexity |
|---|---|
| WRYYAVALALKE | 11.254131 |
| WLVYVAAARLKE | 11.480680 |
| KRYYAAGAAWWE | 14.88393 |
| WHYPAVVARHWE | 12.81336 |
| FLYRWLPSRRGG |
As the article mentions it, pseudo perplexity (PPL) is used to score binders.
Part 2: Evaluate Binders with AlphaFold3
I evaluated the predicted complexes between mutant SOD1 (A4V) and each generated peptide using AlphaFold3. Here is the table with the results:
| Peptide | ipTM | Binding location | |
|---|---|---|---|
| WRYYAVALALKE | 0.25 | outer surface of the β-barrel | |
| WLVYVAAARLKE | 0.24 | outer surface of the β-barrel | |
| KRYYAAGAAWWE | 0.45 | outer surface of the β-barrel, but a bit more interesting because it has a higher ipTM shows secondary structure | |
| WHYPAVVARHWE | 0.29 | outer surface of the β-barrel, also shows a bit of secondary structure | |
| FLYRWLPSRRGG | 0.34 | outer surface of the β-barrel |
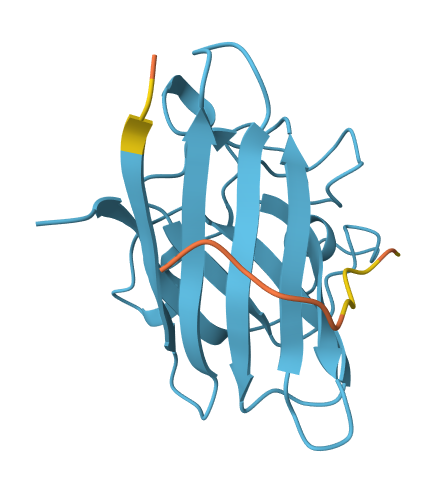
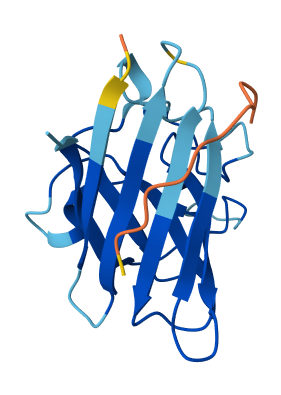
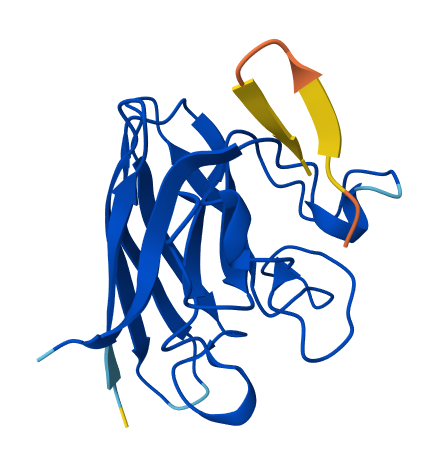
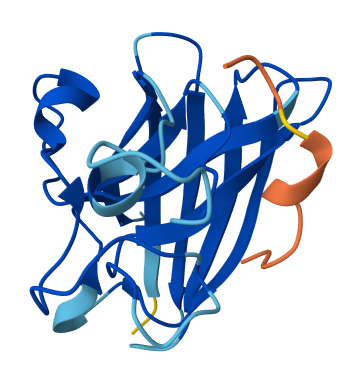
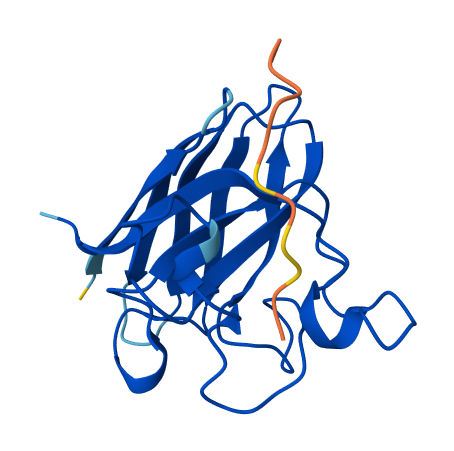
All of the peptides appear to contact basically only the outer surface of the β-barrel, which I guess is not ideal. Here are some pictures:
Across all candidates, the peptides consistently localized to the outer surface of the SOD1 β-barrel, without clear engagement of the N-terminal region where the A4V mutation is located, nor obvious interaction with the dimer interface. In all cases, the peptides appeared largely surface-bound rather than deeply buried, suggesting weak or nonspecific interactions.
The ipTM values for all complexes were relatively low (0.24–0.45), indicating limited confidence in the predicted protein–peptide interfaces. Among the candidates, KRYYAAGAAWWE showed the highest ipTM (0.45) and displayed some degree of secondary structure, which may suggest a slightly more organized interaction compared to the other peptides. However, even this value remains below the range typically associated with strong or reliable binding predictions.
Overall, the PepMLM-generated peptides did not show strong or well-localized binding to functionally relevant regions of SOD1. Compared to the known binder, none of the generated peptides clearly exceeded its predicted interaction quality. In fact, both the generated peptides and the known binder appeared to produce relatively weak or diffuse binding modes in AlphaFold, suggesting that this system may be challenging to model accurately using this approach. Also, all binders were generally colored yellow to orange, indicating lower confidence on the prediction of the peptide’s structure itself.
Part 3: Evaluate Properties of Generated Peptides in the PeptiVerse
| Peptide | Binding Affinity (pKd/pKi) | Solubility | Hemolysis Prob. | Net Charge (pH 7) | Molecular Weight (Da) | GRAVY |
|---|---|---|---|---|---|---|
| WRYYAVALALKE | 6.958 | Soluble | 0.082 | 0.77 | 1482.7 | 0.15 |
| WLVYVAAARLKE | 6.231 | Soluble | 0.065 | 0.77 | 1418.7 | 0.61 |
| KRYYAAGAAWWE | 6.706 | Soluble | 0.052 | 0.77 | 1471.6 | -0.79 |
| WHYPAVVARHWE | 5.873 | Soluble | 0.036 | -0.06 | 1550.7 | -0.59 |
| FLYRWLPSRRGG | 5.968 | Soluble | 0.047 | 2.76 | 1507.7 | -0.71 |
Across all candidates, PeptiVerse predicted high solubility (probability = 1.000) and low hemolysis probability (~0.03–0.05), suggesting that none of the peptides present immediate toxicity or formulation concerns. However, all peptides were predicted to have weak binding affinity (pKd/pKi ≈ 5.8–6.7), which is at least (😟) consistent with the AlphaFold3 results, where all complexes showed relatively low ipTM values (0.24–0.45) and surface-bound interactions on the β-barrel.
From a developability perspective, KRYYAAGAAWWE and WHYPAVVARHWE appear particularly favorable due to their moderate net charge (~0), high solubility, and low hemolysis probability. In contrast, FLYRWLPSRRGG has a significantly higher positive charge (+2.76), which may increase nonspecific interactions despite acceptable solubility and low hemolysis prediction.
No clear correlation was observed between structural confidence and predicted binding affinity. For example, KRYYAAGAAWWE, which had the highest ipTM in AlphaFold3 (0.45), showed relatively weak predicted affinity (6.706), while WHYPAVVARHWE displayed the strongest predicted affinity (5.873) despite a lower ipTM (0.29). This suggests that structural confidence alone is not sufficient to identify the strongest binders.
Overall, the results indicate that the generated peptides tend to form weak, nonspecific interactions, both structurally and energetically. If selecting one peptide to advance, I would choose KRYYAAGAAWWE, as it provides the best balance between structural confidence (highest ipTM) and favorable therapeutic properties, even though its predicted binding affinity remains modest. This makes it the most reasonable starting point for further optimization.