Week 6 HW: Genetic Circuits Part I
Assignment: DNA Assembly
1. What are some components in the Phusion High-Fidelity PCR Master Mix and what is their purpose?
The Phusion High-Fidelity PCR Master Mix contains Phusion DNA polymerase, a enzyme that synthesizes DNA (this one also has proofreading activity). It also includes HF buffer, which provides the optimal pH and salt concentration (MgCl2) required for enzyme activity, as well as dNTPs, which are the nucleotide building blocks used to synthesize the new DNA strands. Glycerol helps stabilize the enzyme, and DMSO may be added to improve amplification of GC-rich regions by reducing secondary structure formation.
2. What are some factors that determine primer annealing temperature during PCR?
Primer annealing temperature is mainly determined by the primer melting temperature (Tm), which depends on primer length, GC content, sequence composition, and buffer salt concentration.
3. There are two methods from this class that create linear fragments of DNA: PCR, and restriction enzyme digests. Compare and contrast these two methods, both in terms of protocol as well as when one may be preferable to use over the other.
PCR generates linear DNA fragments by amplifying a specific region through repeated cycles of denaturation, primer annealing, and extension by DNA polymerase. Restriction enzyme digestion generates linear fragments by cutting existing DNA at specific recognition sites. PCR is preferable when amplifying a specific sequence or adding custom overhangs, while restriction digests are preferable when suitable cut sites are already present in the DNA.
4. How can you ensure that the DNA sequences that you have digested and PCR-ed will be appropriate for Gibson cloning?
Gibson Assembl only works if the sequences you want to clone have homologous overlaps, typically around 20–40 base pairs. So you ensure that it will work by designing PCR primers that include overlap sequences complementary to the neighboring fragment or vector.
5. How does the plasmid DNA enter the E. coli cells during transformation?
There are two famous methods for transformation (the process in which the plasmid enters the cell):
- Chemical Transformation (Heat Shock): where cells are treated with cold divalent cations and then briefly exposed to high temperatures (42°C) to create transient pores in the membrane.
- Electroporation: electrical pulse are used to create temporary pores in the membrane, allowing DNA entry.
6.Describe another assembly method in detail (such as Golden Gate Assembly)
- Explain the other method in 5 - 7 sentences plus diagrams (either handmade or online).
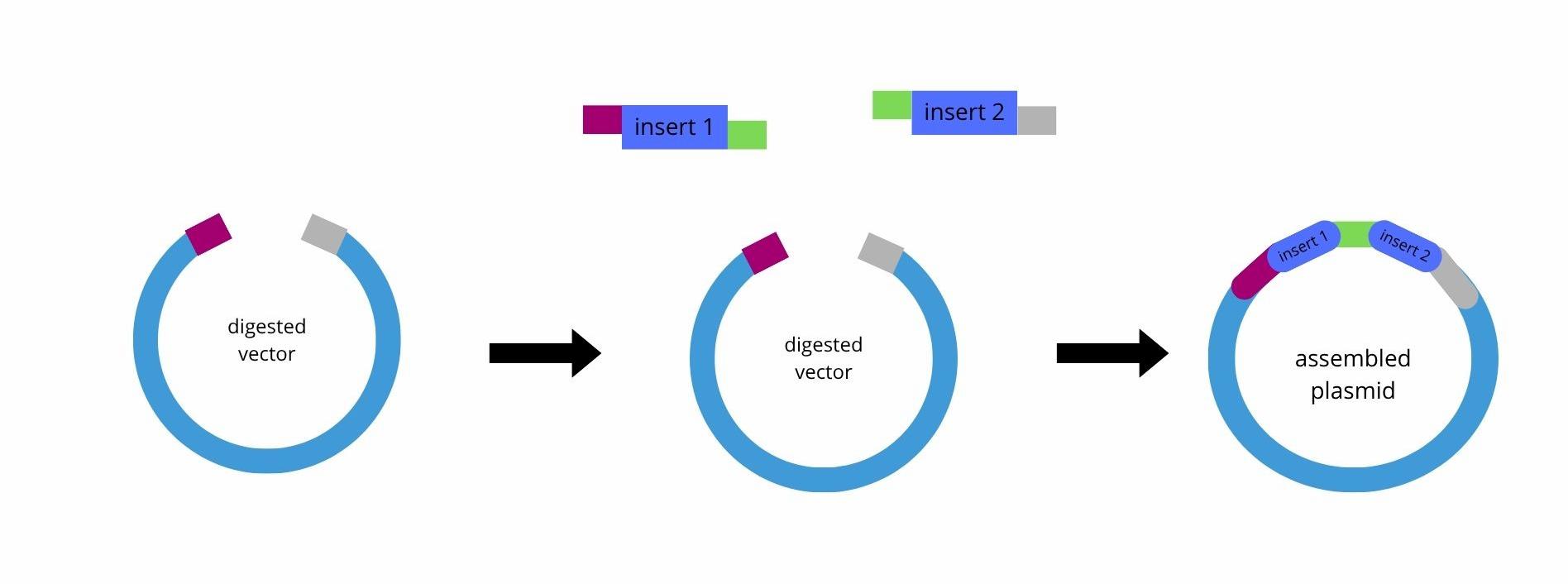
Golden Gate Assembly is a DNA assembly method that uses Type IIS restriction enzymes, such as BsaI or BsmBI, together with DNA ligase to join multiple DNA fragments in a defined order. Unlike standard restriction enzymes, the Type IIS enzymes cut outside of their recognition sequence, which possibilizes the creation of custom single-stranded overhangs. These overhangs can be designed so that each fragment only ligates to the correct neighboring fragment, and this is important if you want an assembly more directional (where the order matters, for example).
During the reaction, the DNA fragments are first digested by the restriction enzyme, producing complementary sticky ends. DNA ligase then joins these compatible ends. Because the recognition sites are removed after correct ligation, the final assembled product is no longer cut in subsequent cycles, which increases assembly efficiency. The reaction is typically performed in thermocycling steps alternating between digestion (for example, 37°C) and ligation (16°C).

Assignment: Asimov Kernel
Repressilator
I ran the repressilator using a 24-hour duration because it generated plots such as the one published by Elowitz in the year 2000.
This is my design (copied from the demo):
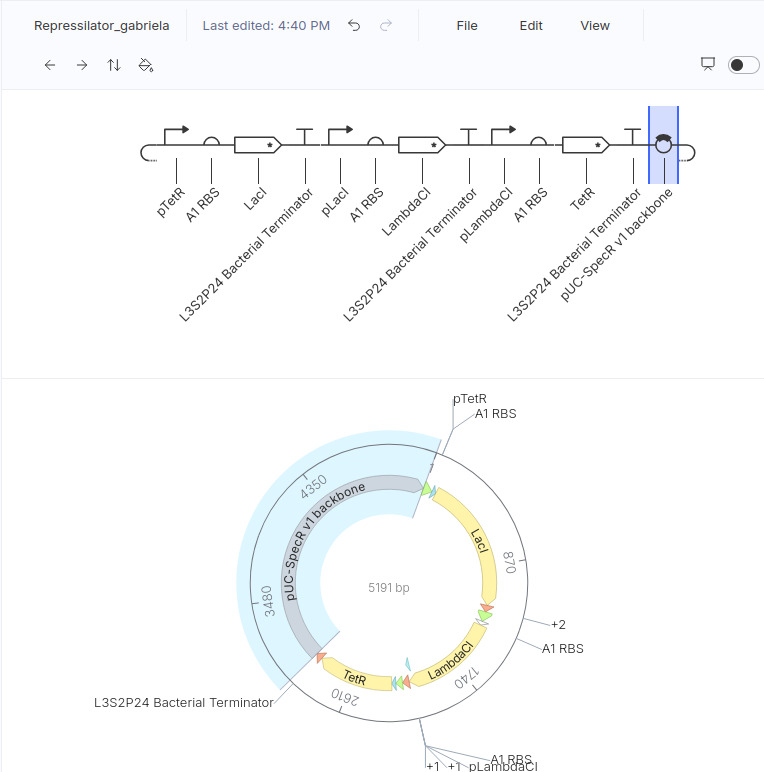
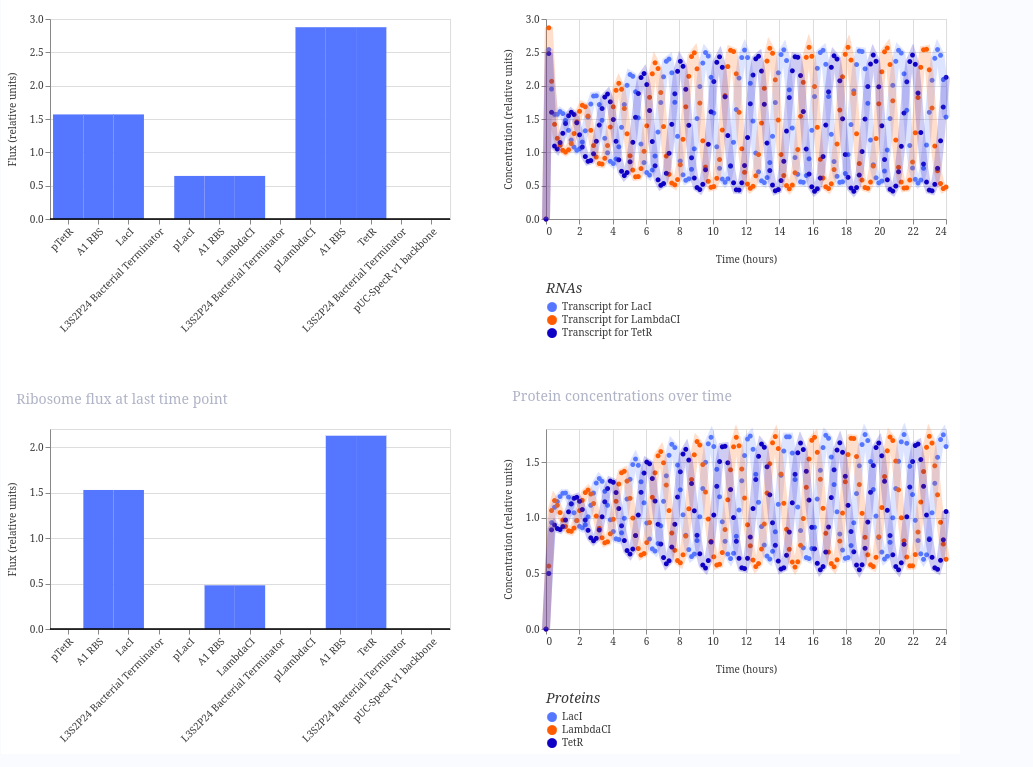
 These are the plots generated:
These are the plots generated:
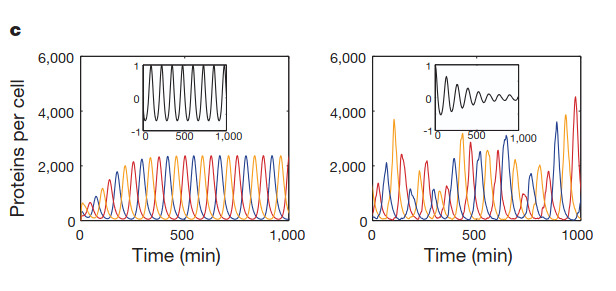
 And this is a figure taken from the Elowitz’s article
And this is a figure taken from the Elowitz’s article

My constructs
Simple GFP expression construct
This construct is a simple inducible reporter plasmid where the pTac promoter controls expression of GFP (BBa_E0040), allowing fluorescence to increase when the promoter is activated.
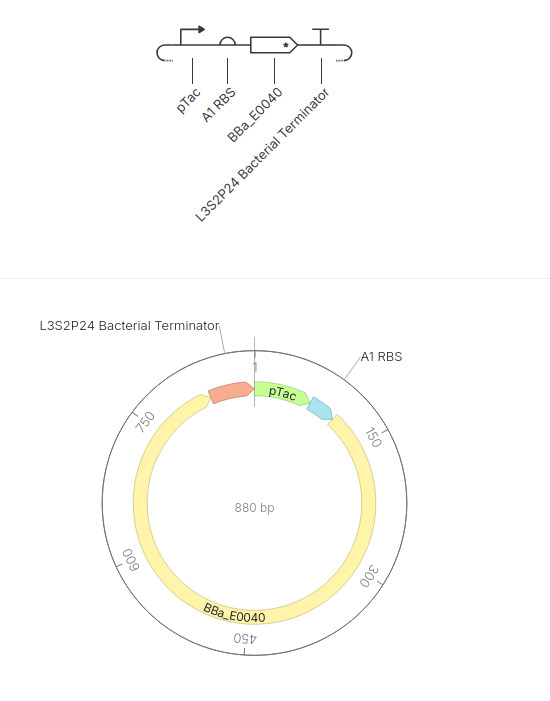
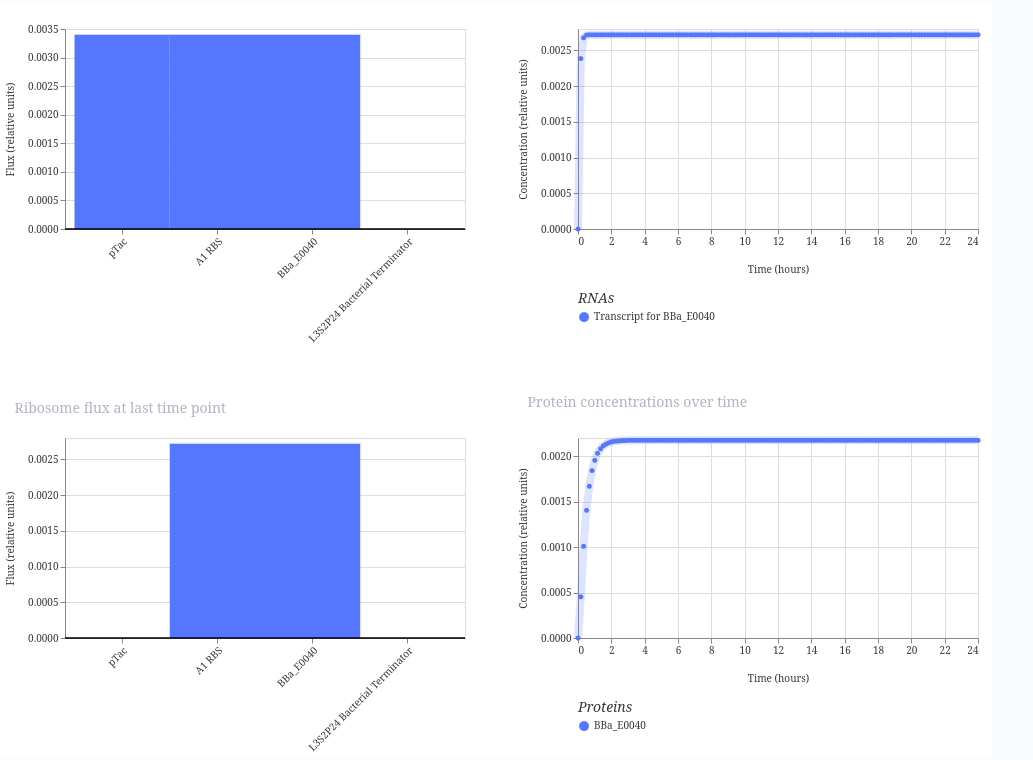
 The result is fluorescence to increase over time when the promoter is active.
The result is fluorescence to increase over time when the promoter is active.
Here is the result:

XOR gate
BasicallY: Output = ON only if Input A OR Input B are ON (if both are ON , then the output is OFF ) Therefore, the truth table can be written as
| Input A | Input B | Output |
|---|---|---|
| 0 | 0 | 0 |
| 0 | 1 | 1 |
| 1 | 0 | 1 |
| 1 | 1 | 0 |
Here is my design:
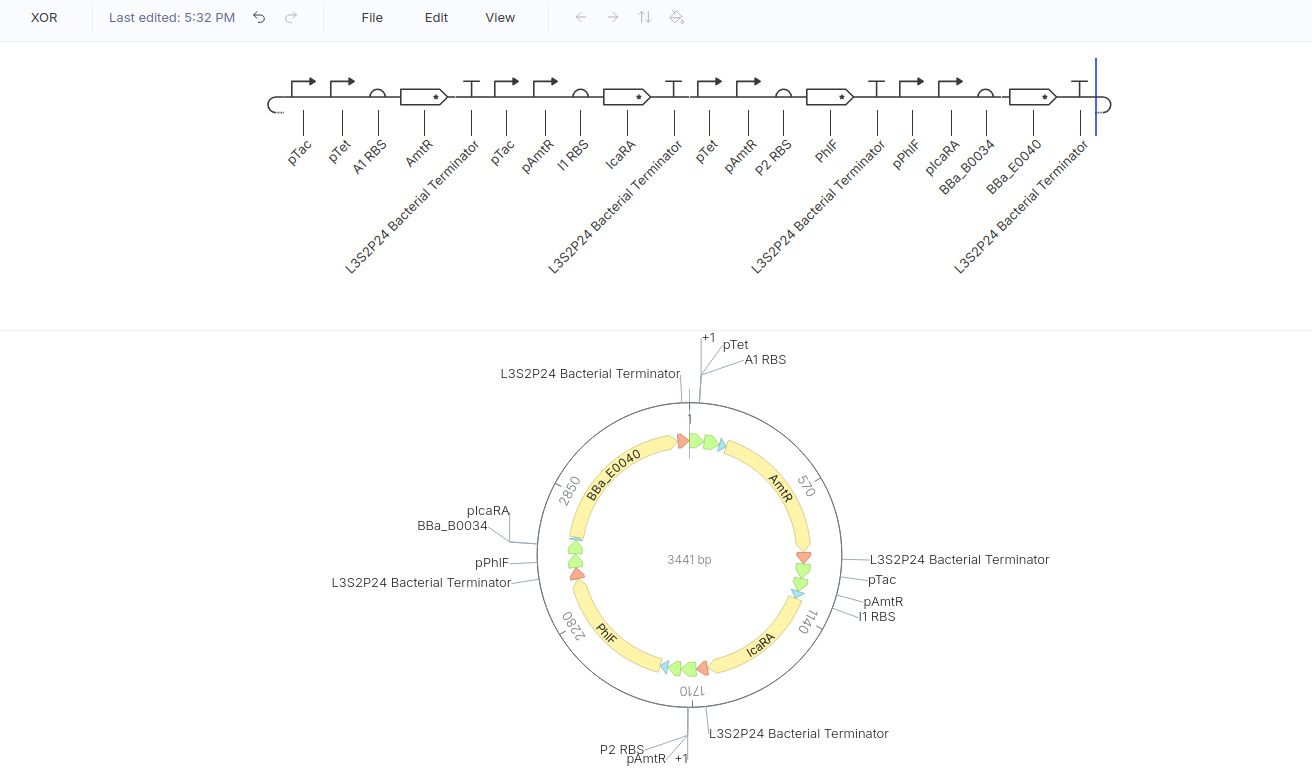
 Here is the result:
Here is the result:
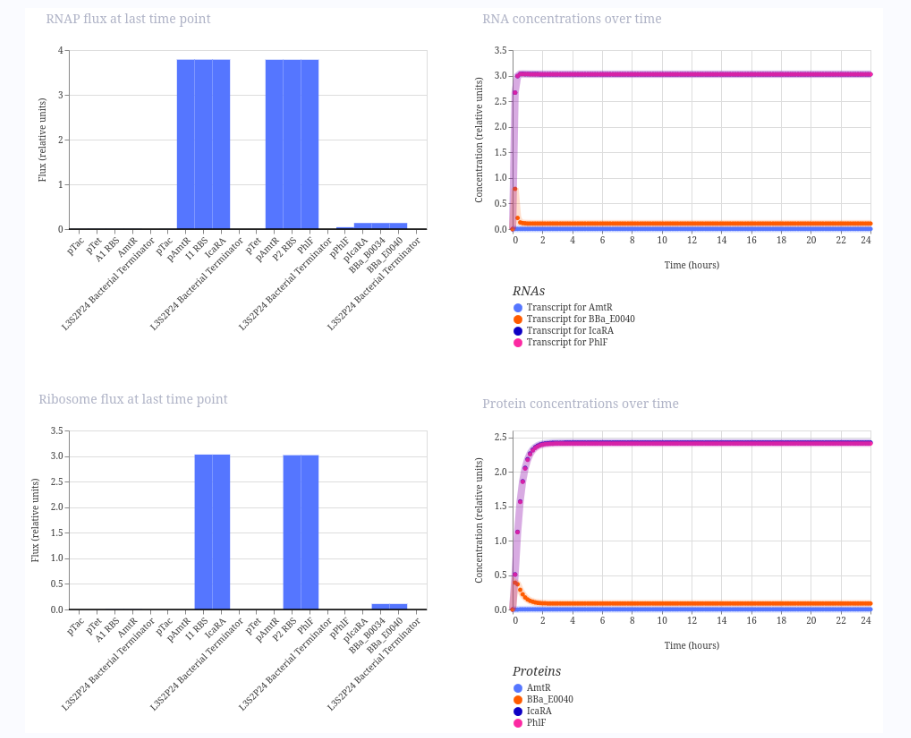

This is not exactly what I expected as it only shows the 1 1 0 case, but I’m not sure how to modulate it.