Week 2 HW: DNA read, write, and edit
Week 2 - DNA read, write and edit
We’re in the deep now, time to get the hands dirty.
Part 0 - Basics of Gel electrophoresis -
What did the small DNA fragment say to the large one? “Catch me if you can!”
In gel electophoresis, the positive end should be on the opposite side of the sample well, i.e sample well should be close to the cathode.
DNA is an acid, so it readily gives away a positive ion and becomes negative. DNA is negatively charged due to the phosphates in its sugar-phosphate backbone (PO4 3-).
Top strand is always 5’ to 3’. The other strand is called the template strand.
Video tutorial on Gel electrophoresis - https://www.youtube.com/watch?v=TIZRGt3YAug
Part 1 - Benchling & In-silico Gel Art
- Make benchling account
- import Lambda DNA
- Simulate restriction enzyme digestion with the below enzymes-
- EcoRI
- HindIII
- BamHI
- KpnI
- EcoRV
- SacI
- SalI
- Create a Pattern/image in Paul Vanouse’s Latent Figure Protocol artworks
In benchling - Create new project. In the left menubar, use the “+” button to create new sequence.
Name it, select DNA, then select linear toppology.
Then click create.
Once can also import sequences.
Just copy GenBank code.
Go to import from database in the “+” section.
Paste code and search.
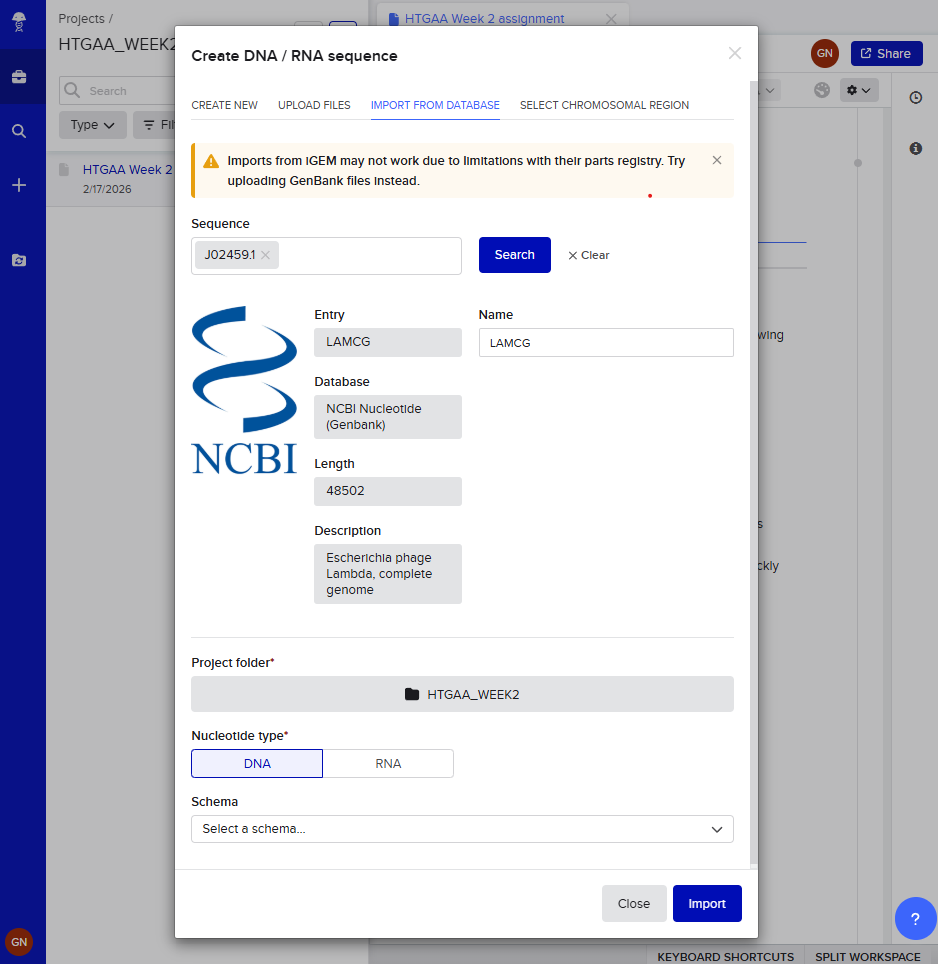

Let’s do a virtual restriction enzyme digestion !
once the DNA is imported and the sequence is visible, select the scissor logo on the right control panel.
Click on new digest, and add the resitriction enzyme name.
Select enzyme and run digest.
Move onto the virtual digest tab in the right panel to be able to see and compare a virtual gel electrophoresis.
Ensure the right ladder is picked - should be congruent with the machine.
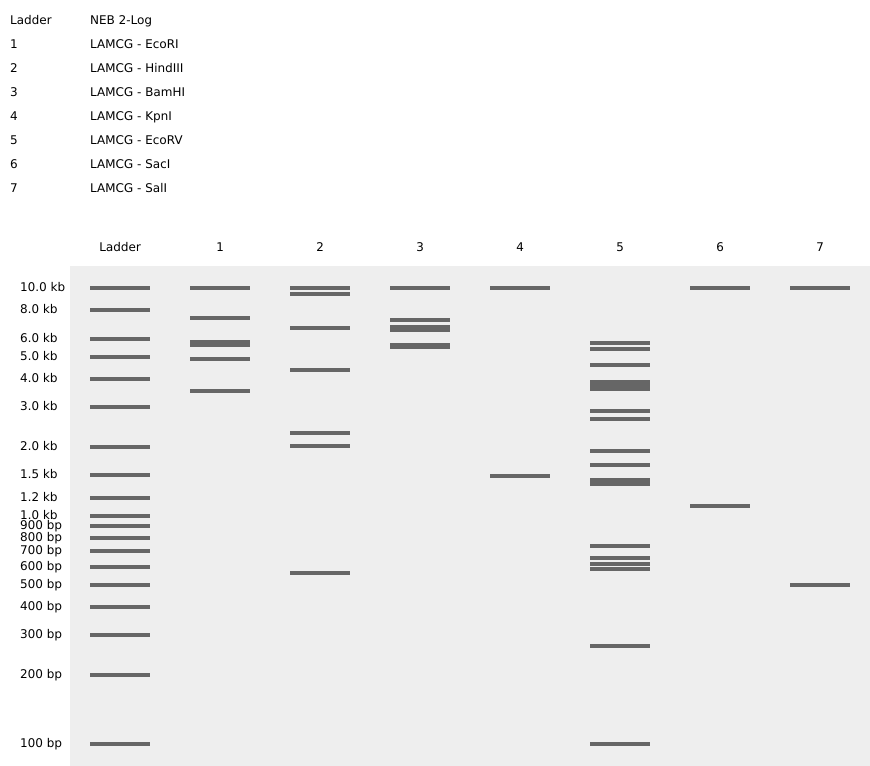

To those who wanna view my benchling work: https://benchling.com/s/seq-3PpRFMxTkg5VowzYd9ya?m=slm-zacrpUMhznCXawpSU6AX
Final part of part 1 - create a virtual art.
I used rcdonovan’s art automation website.
It was quite cumbersome, because I assumed I can draw any shape and it will find it out for me.
Turned out I had to click up and down till I got something I could possibly work with.
Given the proximity to Valentine’s day I decided to go with a heart, with a smiley face in the middle.
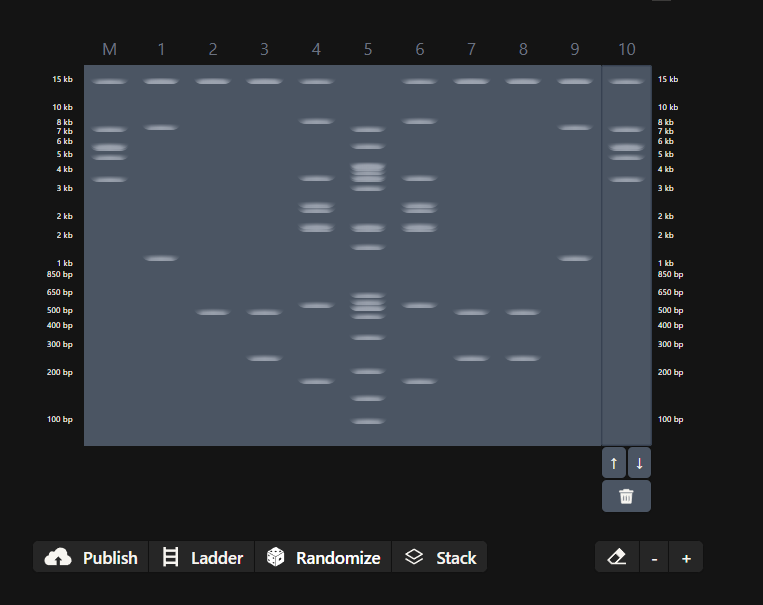

I got the following directions to follow to create the art in benchling
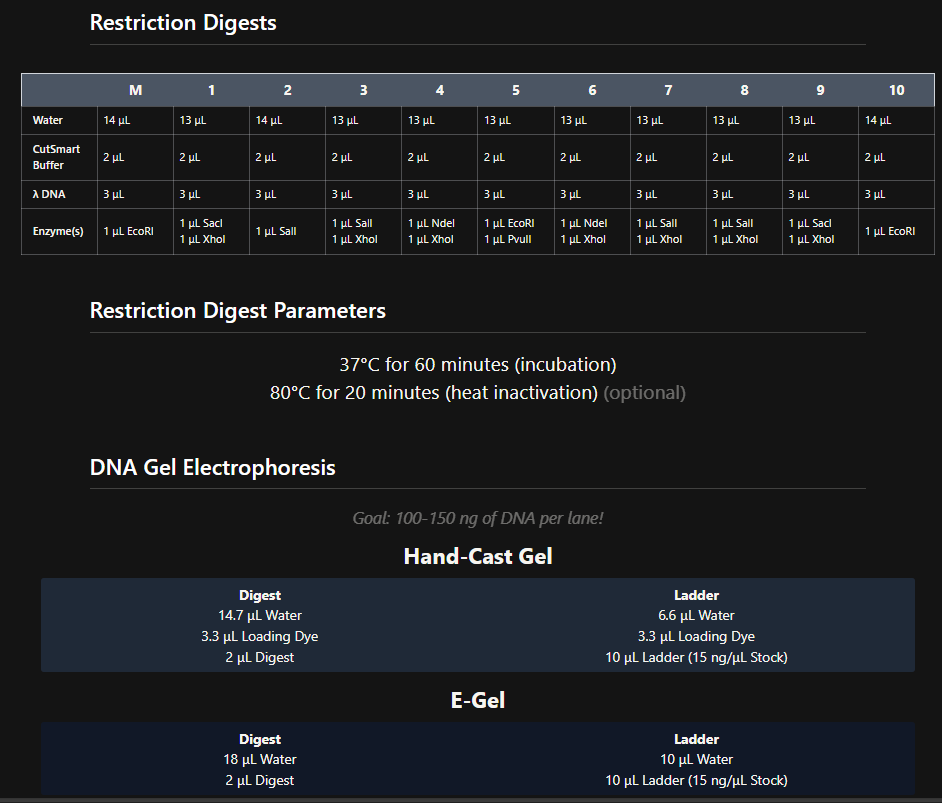

The same done in benchling after doing all the digestions. half the heart is there in gel simualtion, the rest can be made through a mirror reflection.
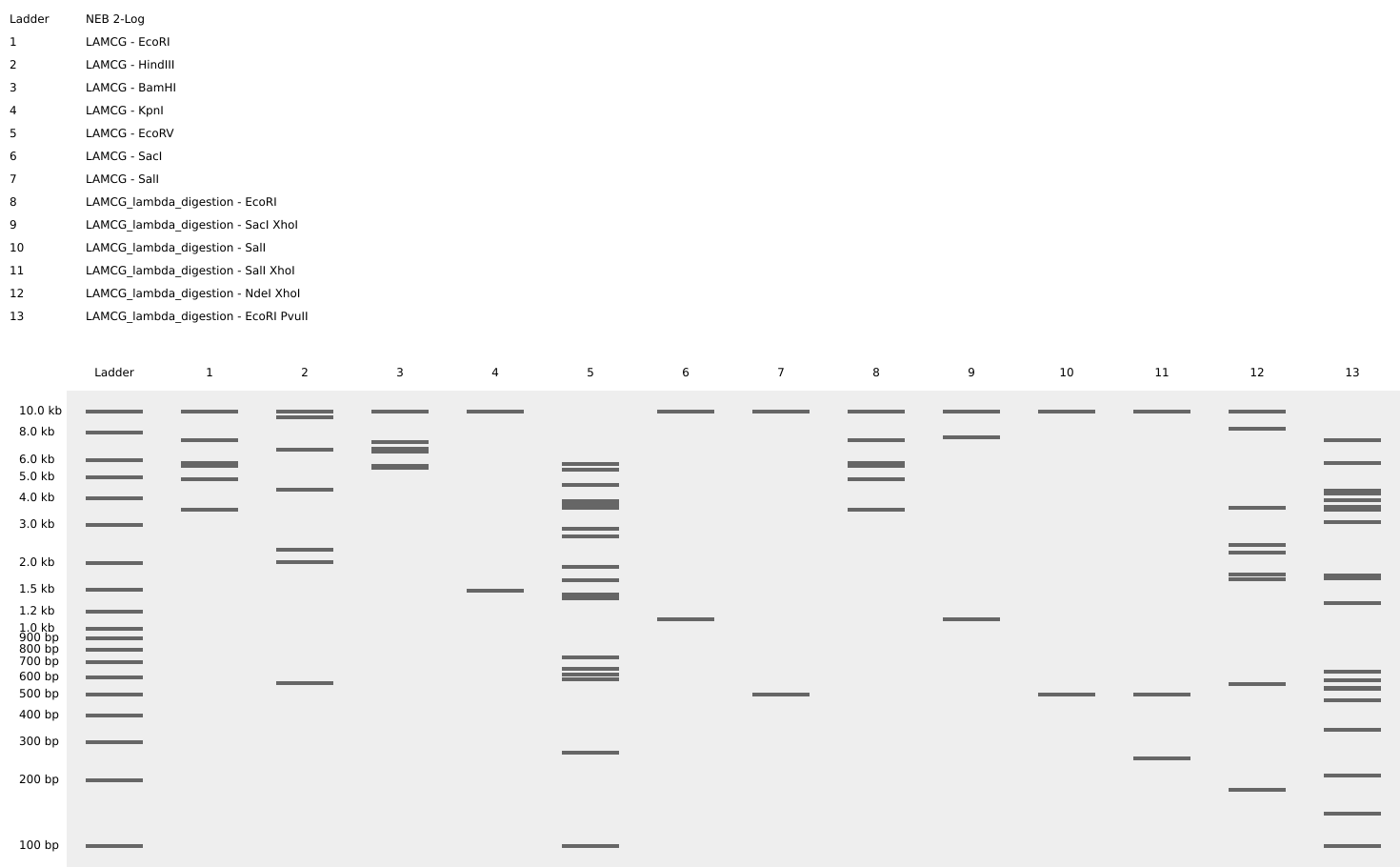

Part 2 - Gel Art - Restriction Digests and Gel electrophoresis
(Unable to do this part since I do not have access to a lab)
Part 3 - DNA Design Challenge
- Choose my Protein
- Reverse translation: Amino acid sequence to DNA nucleotide sequence
- Codon Optimization
- You have a sequence now what?
- How does it work in nature/biological systems?
3.1 Choose your protein: In my governance paper, I had talked about the importance of synbio in space. In continuation with that I have chosen to work with LEA (Late Embryogenesis Abundant) proteins. The idea being to produce active pharmaceutical ingredients (API) in space. But dealing with storage and water shortage issues by dehydrating the results of the bioreactor, to reuse the water. But dehydration should not affect the API, hence a protein that is produced along with the API, that protects it will be great. Upon reading this paper, I had an idea of using the LEA3 https://pmc.ncbi.nlm.nih.gov/articles/PMC2292704/.
Protein sequence acquired from: https://www.uniprot.org/uniprotkb/O81843/entry
mRNA Protein Sequence for LEA3
AAN13065.1 unknown protein [Arabidopsis thaliana] MEVPKSSLLMIIFVVASCFLHVKAWHGQTYCGGNATPRCQLRYIDCPEECPTEMFPNSQNKICWVDCFKP LCEAVCRAVKPNCESYGSICLDPRFIGGDGIVFYFHGKSNEHFSIVSDPDFQINARFTGHRPAGRTRDFT WIQALGFLFNSHKFSLETTKVATWDSNLDHLKFTIDGQDLIIPQETLSTWYSSDKDIKIERLTEKNSVIV TIKDKAEIMVNVVPVTKEDDRIHNYKLPVDDCFAHFEVQFKFINLSPKVDGILGRTYRPDFKNPAKPGVV MPVVGGEDSFRTSSLLSHVCKTCLFSEDPAVASGSVKPKSTYALLDCSRGASSGYGLVCRK
Using the following website to do reverse translation: https://www.bioinformatics.org/sms2/rev_trans.html
The site requested a codon table. The codon table of Arabidopsis Thaliana was obtained using: https://www.kazusa.or.jp/codon/cgi-bin/showcodon.cgi?species=3702&aa=1&style=GCG
Reverse translation result:
reverse translation of AAN13065.1 unknown protein [Arabidopsis thaliana] to a 1023 base sequence of most likely codons. atggaagttcctaagtcttctcttcttatgattatttttgttgttgcttcttgttttctt catgttaaggcttggcatggacaaacttattgtggaggaaatgctactcctagatgtcaa cttagatatattgattgtcctgaagaatgtcctactgaaatgtttcctaattctcaaaat aagatttgttgggttgattgttttaagcctctttgtgaagctgtttgtagagctgttaag cctaattgtgaatcttatggatctatttgtcttgatcctagatttattggaggagatgga attgttttttattttcatggaaagtctaatgaacatttttctattgtttctgatcctgat tttcaaattaatgctagatttactggacatagacctgctggaagaactagagattttact tggattcaagctcttggatttctttttaattctcataagttttctcttgaaactactaag gttgctacttgggattctaatcttgatcatcttaagtttactattgatggacaagatctt attattcctcaagaaactctttctacttggtattcttctgataaggatattaagattgaa agacttactgaaaagaattctgttattgttactattaaggataaggctgaaattatggtt aatgttgttcctgttactaaggaagatgatagaattcataattataagcttcctgttgat gattgttttgctcattttgaagttcaatttaagtttattaatctttctcctaaggttgat ggaattcttggaagaacttatagacctgattttaagaatcctgctaagcctggagttgtt atgcctgttgttggaggagaagattcttttagaacttcttctcttctttctcatgtttgt aagacttgtcttttttctgaagatcctgctgttgcttctggatctgttaagcctaagtct acttatgctcttcttgattgttctagaggagcttcttctggatatggacttgtttgtaga aag
reverse translation of AAN13065.1 unknown protein [Arabidopsis thaliana] to a 1023 base sequence of consensus codons. atggargtnccnaarwsnwsnytnytnatgathathttygtngtngcnwsntgyttyytn caygtnaargcntggcayggncaracntaytgyggnggnaaygcnacnccnmgntgycar ytnmgntayathgaytgyccngargartgyccnacngaratgttyccnaaywsncaraay aarathtgytgggtngaytgyttyaarccnytntgygargcngtntgymgngcngtnaar ccnaaytgygarwsntayggnwsnathtgyytngayccnmgnttyathggnggngayggn athgtnttytayttycayggnaarwsnaaygarcayttywsnathgtnwsngayccngay ttycarathaaygcnmgnttyacnggncaymgnccngcnggnmgnacnmgngayttyacn tggathcargcnytnggnttyytnttyaaywsncayaarttywsnytngaracnacnaar gtngcnacntgggaywsnaayytngaycayytnaarttyacnathgayggncargayytn athathccncargaracnytnwsnacntggtaywsnwsngayaargayathaarathgar mgnytnacngaraaraaywsngtnathgtnacnathaargayaargcngarathatggtn aaygtngtnccngtnacnaargargaygaymgnathcayaaytayaarytnccngtngay gaytgyttygcncayttygargtncarttyaarttyathaayytnwsnccnaargtngay ggnathytnggnmgnacntaymgnccngayttyaaraayccngcnaarccnggngtngtn atgccngtngtnggnggngargaywsnttymgnacnwsnwsnytnytnwsncaygtntgy aaracntgyytnttywsngargayccngcngtngcnwsnggnwsngtnaarccnaarwsn acntaygcnytnytngaytgywsnmgnggngcnwsnwsnggntayggnytngtntgymgn aar
Using another site to get the codon optimized sequence for Saccharomyces Cerevisiae S288C - from https://en.vectorbuilder.com/tool/codon-optimization.html
ATGGAAGTACCAAAGTCGTCTTTATTAATGATTATATTTGTTGTTGCATCTTGTTTTTTACATGTTAAAGCTTGGCATGGTCAAACTTATTGCGGTGGTAACGCTACACCACGTTGCCAACTTAGATATATCGATTGTCCAGAAGAGTGCCCAACTGAAATGTTTCCAAATTCACAGAACAAAATTTGTTGGGTTGATTGTTTTAAACCACTATGTGAAGCAGTTTGCAGAGCTGTTAAACCAAATTGCGAAAGCTACGGTAGCATATGTTTAGATCCAAGGTTCATTGGAGGAGATGGTATTGTTTTCTACTTCCACGGTAAATCCAATGAACATTTTTCTATTGTATCTGACCCAGATTTTCAAATTAACGCTAGATTTACAGGACATAGACCAGCCGGTAGAACAAGAGATTTCACCTGGATACAAGCTTTAGGATTTCTGTTTAACTCCCATAAGTTCTCTTTAGAAACAACAAAAGTTGCCACCTGGGACTCTAATTTGGATCATTTGAAGTTTACAATTGATGGCCAGGACTTGATAATTCCTCAAGAAACTTTATCAACATGGTACTCCTCTGATAAAGATATTAAAATTGAAAGATTAACCGAAAAGAATTCTGTTATCGTTACAATTAAAGATAAGGCCGAAATTATGGTTAATGTTGTTCCAGTTACAAAAGAAGATGACAGAATACACAATTATAAACTTCCAGTTGATGACTGTTTTGCTCATTTTGAGGTTCAATTCAAATTCATTAACTTGTCTCCTAAAGTTGATGGTATTTTAGGTAGGACTTATAGACCAGATTTCAAGAACCCAGCTAAGCCAGGTGTCGTGATGCCAGTCGTTGGCGGTGAGGATAGCTTTAGAACTTCTTCTTTGTTATCACACGTTTGTAAAACCTGTTTGTTTTCTGAGGACCCTGCCGTTGCTTCTGGTTCTGTTAAACCAAAATCTACTTATGCTCTATTGGATTGTTCAAGAGGTGCCTCCTCCGGTTATGGTTTGGTTTGTAGAAAATGA
What technologies could be used to produce this protein from your DNA? Describe in your words the DNA sequence can be transcribed and translated into your protein. You may describe either cell-dependent or cell-free methods, or both.
Some technologies that can be used to produce this protein can be:
- Cell based
- Using the S.Cerevisuae as a chassis to make the proteins for us
- Using plasmid based protein expression
- Cell free
- Using PCR
How does DNA get transcribed and translated into protein DNA get converted into RNA by RNA polymerase, in the nucleus. This RNA leaves out into the cytoplasm, and gets translated into amino acids. The RNA strand that leaves the nucleus is the messenger RNA. Which eventually finds its way to a ribosome. In the ribosome, a tRNA combines with codons, and amino acids on the other end, hence making the protein chains.
For the next part using: TPA_inf: Saccharomyces cerevisiae S288C chromosome I, complete sequence https://www.ncbi.nlm.nih.gov/nuccore/BK006935.2/
promoter - pTEF1 Translation Initiation Region (Yeast Kozak Sequence) - AAAAAAATGTCT Start codon - ATG Stop codon - TAA Terminator - tADH1
Found something new that there is y and n and in consensus codons - y is for u/c and n - for any
Part 4 - Prepare a Twist DNA synthesis order
- Create account in twist
- Build your DNA insert sequence - New DNA/RNA sequence in benchling
- Give insert sequence a name with a linear topology
- Add each codon optimized sequence in benchling and color it
- Click on linear map to preview
- Download FASTA file
- Visualize DNA design in SBOL canvas
- On twist, select the Genes option
- Select “Clonal Genes” option
- Import sequence
- Choose your vector
- Choose clonign vectors like pTwist Amp High Copy
- Download construct - GenBank
- Import back into Benchling
Part 5 - DNA read/write/edit
DNA Read
What DNA would you want to sequence and why?
In lecture, a variety of sequencing technologies were mentioned. What tech would you use to perform sequencing on your DNA and why?
Is your method first-, second- or third-generation or other? How so?
What is your input? How do you prepare your input (e.g. fragmentation, adapter ligation, PCR)? List the essential steps.
What are the essential steps of your chosen sequencing technology, how does it decode the bases of your DNA sample (base calling)?
What is the output of your chosen sequencing technology?
Ans. In addition to LEA3, I would sequence DNA related to microbial environmental monitoring and crew health, specifically microbial genomes present in the spacecraft habitat and bioreactor systems. Monitoring microbial populations is important in long-duration missions because microbial evolution, contamination, or biofilm formation can affect both human health and biological manufacturing systems. Sequencing allows detection of pathogenic organisms, identification of mutations in engineered production strains, and verification that biological systems remain genetically stable over time. For sequencing, I would use Oxford Nanopore sequencing, which is a third-generation sequencing technology, because it allows long-read sequencing, minimal sample preparation, compact instrumentation, and real-time analysis, making it suitable for autonomous space environments. The input is purified DNA extracted from environmental or biological samples, which is prepared by DNA extraction, optional fragmentation depending on desired read length, adapter ligation, and sometimes PCR amplification if DNA quantities are low. During sequencing, individual DNA molecules pass through a nanopore embedded in a membrane, and changes in ionic current are measured as different nucleotide sequences move through the pore; these electrical signal changes are decoded computationally into base sequences through base-calling algorithms. The output of the sequencing process is a digital file containing nucleotide sequences (reads), along with quality scores, which can then be assembled, compared to reference genomes, or analyzed for mutations and species identification.
DNA Write
- What DNA would you want to synthesize (e.g., write) and why? These could be individual genes, clusters of genes or genetic circuits, whole genomes, and beyond. As described in class thus far, applications could range from therapeutics and drug discovery (e.g., mRNA vaccines and therapies) to novel biomaterials (e.g. structural proteins), to sensors (e.g., genetic circuits for sensing and responding to inflammation, environmental stimuli, etc.), to art (DNA origamis). If possible, include the specific genetic sequence(s) of what you would like to synthesize! You will have the opportunity to actually have Twist synthesize these DNA constructs! :)
I would synthesize a genetic circuit for sensing oxidative stress and activating a protective response in Saccharomyces cerevisiae. In a space environment, radiation and altered metabolism can increase reactive oxygen species (ROS), which can damage both cells and pharmaceutical production pathways. A ROS-responsive genetic circuit could act as a biological sensor that activates protective genes or temporarily pauses production when stress levels become high, improving reliability and safety of autonomous biomanufacturing systems.
The construct would include a stress-responsive promoter, a reporter or regulatory protein, and a downstream response gene. For example, a simplified design could include a yeast oxidative stress promoter (such as a promoter derived from oxidative stress response genes), followed by a coding sequence for a fluorescent reporter or regulatory protein, and a transcription terminator.
- What Technology would you use to perform this DNA synthesis and why *. What are the essential steps in the chosen sequencing methods? *. What are the limitations of your sequencing methods in terms of speed, accuracy and scalability?
I would use chemical DNA synthesis using phosphoramidite chemistry, which is the most common and reliable method used by commercial DNA synthesis companies. This method builds DNA one nucleotide at a time on a solid support, allowing precise control over the final sequence and making it suitable for synthesizing gene-length constructs.
The essential steps include chemically adding nucleotides sequentially to build short DNA fragments, removing protective groups after each step, assembling short fragments into a full gene if needed, and verifying the final sequence through DNA sequencing.
The main limitations are that longer DNA sequences are harder to synthesize directly because small errors accumulate during each chemical step. This limits the length of DNA that can be made in one piece and requires assembly from shorter fragments. While accuracy is high after verification, synthesis speed depends on chemical cycle time, and very large constructs become slower and more expensive to produce at scale.
source - Chatgpt 5.2
DNA edit
- What DNA would you want to edit and why?
I would edit the DNA of Saccharomyces cerevisiae to modify stress-response and metabolic regulation genes so that the organism produces pharmaceutical compounds efficiently while limiting uncontrolled growth. In a space environment, it is useful for production organisms to remain stable, tolerate radiation and oxidative stress, and enter low-activity states when production is not required. Editing genes involved in stress tolerance or metabolic regulation could allow the yeast to maintain productivity without excessive biomass accumulation, improving safety and reducing resource consumption in a closed system.
- What Techhnology or Technologies would be used to perform these DNA edits and why?
- How does your technology of choice edit DNA? What are the essential steps?
I would use CRISPR–Cas9 genome editing, because it allows precise, targeted modification of DNA and is widely used in yeast engineering. CRISPR-based editing is efficient, relatively simple to design, and supports both gene knockouts and insertion of new genetic elements at specific locations in the genome.
- What preparation do you need to do (e.g. design steps) and what is the input (e.g. DNA template, enzymes, plasmids, primers, guides, cells) for the editing?
CRISPR–Cas9 uses a guide RNA to direct the Cas9 enzyme to a specific DNA sequence in the genome. Cas9 creates a double-stranded break at that location. The cell then repairs the break using its natural repair mechanisms. If a repair template is provided, the cell incorporates the new DNA sequence during repair, allowing precise edits such as insertions, deletions, or replacements.
What are the limitations of your editing methods (if any) in terms of efficiency or precision?
The preparation involves designing a guide RNA that matches the target DNA sequence and designing a repair template containing the desired genetic change. The inputs typically include a plasmid encoding Cas9, the guide RNA sequence, the repair DNA template, and yeast cells to be edited. After transformation, edited cells are selected and verified using sequencing to confirm that the intended modification was successfully introduced.