Week 3 HW: Lab Automation
Week 3 lab automation
Assignment: Python Script for Opentrons Artwork
- Review Recitation
- Generate an artistic design using the GUI at opentrons-art.rcdonovan.com.
- using the coordinates from the GUI, follow the instructions in the HTGAA26 Opentrons Colab to write your own Python script which draws your design using the Opentrons.
- Submit form
Using AI to generate an image. Image prompt: I am making agar art. I need an image reference. In the image add 3 star flares in the color blue. Lets have an “orbit” around the 3 stars in yellow color, which is an ellipse/spiral that starts thin and gets thicker. In red, lets have a NASA style triangle in the bottom right and top left that is pointing upwards and is small. Give discrete points, not a continous image, similar to bitmap. Use only the colors specified. Give top view.
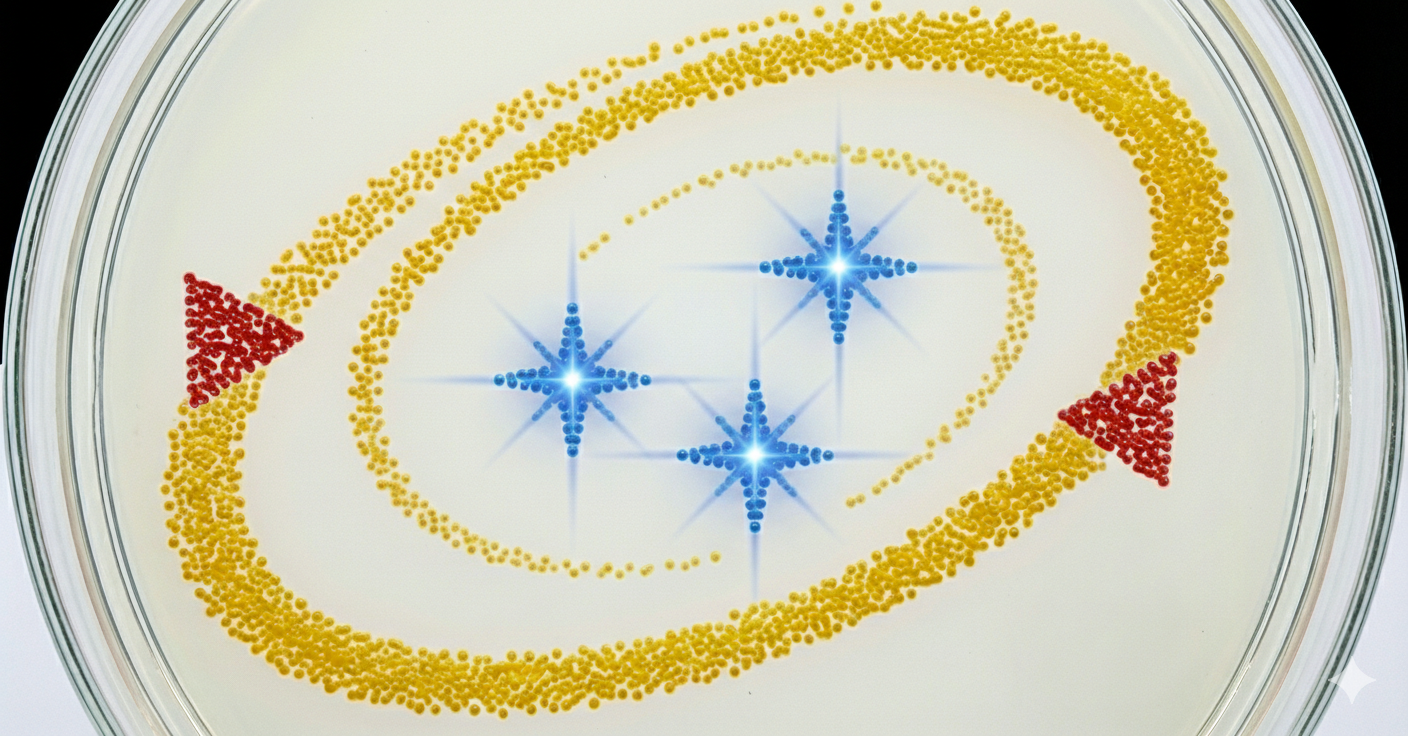


Then used the image to generate art GUI concept on RCdonovan’s opentrons art site
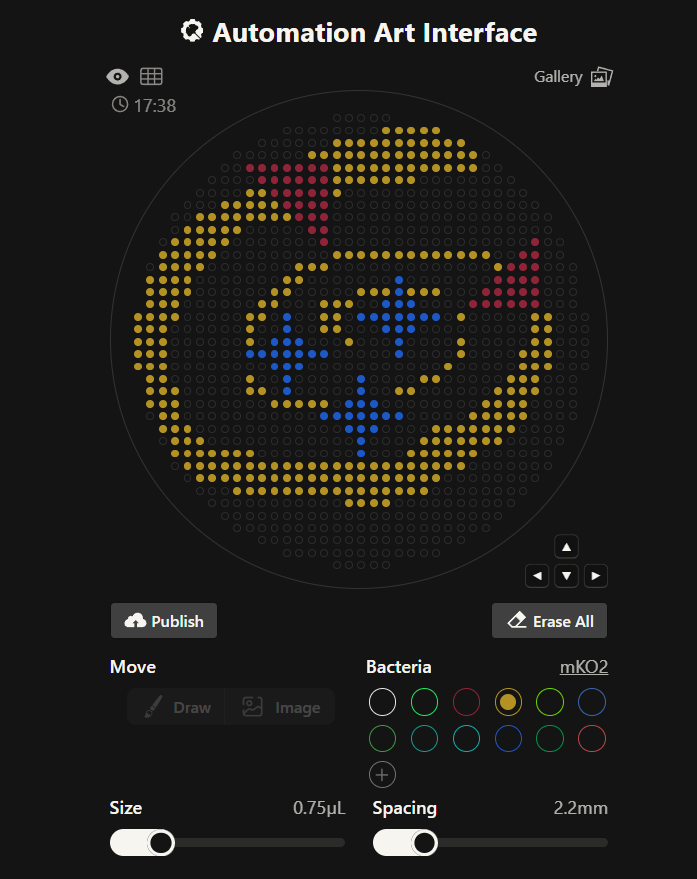

In here are the coordinates I got from the rcdonovan website for red, blue and yellow
mko2_points = [(4.4, 37.4),(6.6, 37.4),(8.8, 37.4),(11, 37.4),(13.2, 37.4),(-4.4, 35.2),(-2.2, 35.2),(0, 35.2),(2.2, 35.2),(4.4, 35.2),(6.6, 35.2),(8.8, 35.2),(11, 35.2),(13.2, 35.2),(15.4, 35.2),(17.6, 35.2),(-8.8, 33),(-6.6, 33),(-4.4, 33),(-2.2, 33),(0, 33),(2.2, 33),(4.4, 33),(6.6, 33),(8.8, 33),(11, 33),(13.2, 33),(15.4, 33),(17.6, 33),(19.8, 33),(-4.4, 30.8),(-2.2, 30.8),(0, 30.8),(2.2, 30.8),(4.4, 30.8),(6.6, 30.8),(8.8, 30.8),(11, 30.8),(13.2, 30.8),(15.4, 30.8),(17.6, 30.8),(19.8, 30.8),(-4.4, 28.6),(-2.2, 28.6),(0, 28.6),(2.2, 28.6),(4.4, 28.6),(6.6, 28.6),(8.8, 28.6),(-19.8, 26.4),(-17.6, 26.4),(-4.4, 26.4),(-24.2, 24.2),(-22, 24.2),(-19.8, 24.2),(-17.6, 24.2),(-15.4, 24.2),(-28.6, 22),(-26.4, 22),(-24.2, 22),(-22, 22),(-19.8, 22),(-17.6, 22),(-15.4, 22),(-13.2, 22),(-30.8, 19.8),(-28.6, 19.8),(-26.4, 19.8),(-24.2, 19.8),(-22, 19.8),(-33, 17.6),(-30.8, 17.6),(-28.6, 17.6),(-26.4, 17.6),(-24.2, 17.6),(-35.2, 15.4),(-33, 15.4),(-30.8, 15.4),(-28.6, 15.4),(-26.4, 15.4),(-4.4, 15.4),(-2.2, 15.4),(0, 15.4),(2.2, 15.4),(4.4, 15.4),(6.6, 15.4),(8.8, 15.4),(11, 15.4),(13.2, 15.4),(15.4, 15.4),(17.6, 15.4),(19.8, 15.4),(22, 15.4),(-35.2, 13.2),(-33, 13.2),(-30.8, 13.2),(-28.6, 13.2),(-11, 13.2),(-8.8, 13.2),(-6.6, 13.2),(24.2, 13.2),(-37.4, 11),(-35.2, 11),(-33, 11),(-30.8, 11),(-13.2, 11),(-11, 11),(-37.4, 8.8),(-35.2, 8.8),(-33, 8.8),(-30.8, 8.8),(-15.4, 8.8),(-13.2, 8.8),(0, 8.8),(2.2, 8.8),(4.4, 8.8),(8.8, 8.8),(11, 8.8),(13.2, 8.8),(-37.4, 6.6),(-35.2, 6.6),(-33, 6.6),(-17.6, 6.6),(-15.4, 6.6),(-6.6, 6.6),(-4.4, 6.6),(-2.2, 6.6),(-39.6, 4.4),(-37.4, 4.4),(-35.2, 4.4),(-33, 4.4),(-19.8, 4.4),(-17.6, 4.4),(-6.6, 4.4),(-4.4, 4.4),(17.6, 4.4),(30.8, 4.4),(33, 4.4),(-39.6, 2.2),(-37.4, 2.2),(-35.2, 2.2),(-19.8, 2.2),(-6.6, 2.2),(-4.4, 2.2),(30.8, 2.2),(33, 2.2),(-39.6, 0),(-37.4, 0),(-35.2, 0),(-19.8, 0),(-2.2, 0),(17.6, 0),(30.8, 0),(33, 0),(-39.6, -2.2),(-37.4, -2.2),(-35.2, -2.2),(17.6, -2.2),(28.6, -2.2),(30.8, -2.2),(33, -2.2),(-39.6, -4.4),(-37.4, -4.4),(-35.2, -4.4),(15.4, -4.4),(28.6, -4.4),(30.8, -4.4),(33, -4.4),(-37.4, -6.6),(-35.2, -6.6),(-19.8, -6.6),(11, -6.6),(13.2, -6.6),(26.4, -6.6),(28.6, -6.6),(30.8, -6.6),(-37.4, -8.8),(-35.2, -8.8),(-33, -8.8),(-19.8, -8.8),(-17.6, -8.8),(6.6, -8.8),(8.8, -8.8),(24.2, -8.8),(26.4, -8.8),(28.6, -8.8),(30.8, -8.8),(-37.4, -11),(-35.2, -11),(-33, -11),(-15.4, -11),(-13.2, -11),(-11, -11),(-8.8, -11),(-6.6, -11),(22, -11),(24.2, -11),(26.4, -11),(28.6, -11),(-35.2, -13.2),(-33, -13.2),(19.8, -13.2),(22, -13.2),(24.2, -13.2),(26.4, -13.2),(28.6, -13.2),(-35.2, -15.4),(-33, -15.4),(-30.8, -15.4),(13.2, -15.4),(15.4, -15.4),(17.6, -15.4),(19.8, -15.4),(22, -15.4),(24.2, -15.4),(26.4, -15.4),(-33, -17.6),(-30.8, -17.6),(-28.6, -17.6),(11, -17.6),(13.2, -17.6),(15.4, -17.6),(17.6, -17.6),(19.8, -17.6),(22, -17.6),(24.2, -17.6),(-33, -19.8),(-30.8, -19.8),(-28.6, -19.8),(-26.4, -19.8),(-24.2, -19.8),(-22, -19.8),(-19.8, -19.8),(8.8, -19.8),(11, -19.8),(13.2, -19.8),(15.4, -19.8),(17.6, -19.8),(19.8, -19.8),(-30.8, -22),(-28.6, -22),(-26.4, -22),(-24.2, -22),(-22, -22),(-19.8, -22),(-17.6, -22),(-15.4, -22),(-13.2, -22),(-11, -22),(-8.8, -22),(-6.6, -22),(-4.4, -22),(-2.2, -22),(0, -22),(2.2, -22),(4.4, -22),(6.6, -22),(8.8, -22),(11, -22),(13.2, -22),(15.4, -22),(17.6, -22),(-28.6, -24.2),(-26.4, -24.2),(-24.2, -24.2),(-22, -24.2),(-19.8, -24.2),(-17.6, -24.2),(-15.4, -24.2),(-13.2, -24.2),(-11, -24.2),(-8.8, -24.2),(-6.6, -24.2),(-4.4, -24.2),(-2.2, -24.2),(0, -24.2),(2.2, -24.2),(4.4, -24.2),(6.6, -24.2),(8.8, -24.2),(11, -24.2),(13.2, -24.2),(-22, -26.4),(-19.8, -26.4),(-17.6, -26.4),(-15.4, -26.4),(-13.2, -26.4),(-11, -26.4),(-8.8, -26.4),(-6.6, -26.4),(-4.4, -26.4),(-2.2, -26.4),(0, -26.4),(2.2, -26.4),(4.4, -26.4),(6.6, -26.4),(8.8, -26.4),(11, -26.4),(-2.2, -28.6),(0, -28.6),(2.2, -28.6),(4.4, -28.6)] electra2_points = [(6.6, 11),(6.6, 8.8),(4.4, 6.6),(6.6, 6.6),(8.8, 6.6),(-13.2, 4.4),(0, 4.4),(2.2, 4.4),(4.4, 4.4),(6.6, 4.4),(8.8, 4.4),(11, 4.4),(13.2, 4.4),(-13.2, 2.2),(4.4, 2.2),(6.6, 2.2),(8.8, 2.2),(-15.4, 0),(-13.2, 0),(-11, 0),(6.6, 0),(-19.8, -2.2),(-17.6, -2.2),(-15.4, -2.2),(-13.2, -2.2),(-11, -2.2),(-8.8, -2.2),(-6.6, -2.2),(6.6, -2.2),(-15.4, -4.4),(-13.2, -4.4),(-11, -4.4),(-13.2, -6.6),(0, -6.6),(-13.2, -8.8),(0, -8.8),(-2.2, -11),(0, -11),(2.2, -11),(-6.6, -13.2),(-4.4, -13.2),(-2.2, -13.2),(0, -13.2),(2.2, -13.2),(4.4, -13.2),(6.6, -13.2),(-2.2, -15.4),(0, -15.4),(2.2, -15.4),(0, -17.6),(0, -19.8)] mrfp1_points = [(-19.8, 30.8),(-17.6, 30.8),(-15.4, 30.8),(-13.2, 30.8),(-11, 30.8),(-8.8, 30.8),(-6.6, 30.8),(-17.6, 28.6),(-15.4, 28.6),(-13.2, 28.6),(-11, 28.6),(-8.8, 28.6),(-6.6, 28.6),(-15.4, 26.4),(-13.2, 26.4),(-11, 26.4),(-8.8, 26.4),(-6.6, 26.4),(-13.2, 24.2),(-11, 24.2),(-8.8, 24.2),(-6.6, 24.2),(-11, 22),(-8.8, 22),(-6.6, 22),(-8.8, 19.8),(-6.6, 19.8),(-6.6, 17.6),(30.8, 17.6),(28.6, 15.4),(30.8, 15.4),(26.4, 13.2),(28.6, 13.2),(30.8, 13.2),(24.2, 11),(26.4, 11),(28.6, 11),(30.8, 11),(22, 8.8),(24.2, 8.8),(26.4, 8.8),(28.6, 8.8),(30.8, 8.8),(19.8, 6.6),(22, 6.6),(24.2, 6.6),(26.4, 6.6),(28.6, 6.6),(30.8, 6.6)]
The copied google colab link is here: https://colab.research.google.com/drive/18vdbBdfkDA-w4Ehbb-0FUUs5g1yCzt-Q#scrollTo=pczDLwsq64mk&line=7&uniqifier=1 Please reach out to me if for some reason the file is not shared or viewable.
Google form post-submission link (for TAs only): https://docs.google.com/forms/d/e/1FAIpQLSddW_4xl32vZHADiUUx4llXO-fwLYnOpT-YNbJB1FozGhT7jQ/alreadyresponded
My code (ignoring the boiler plate):
et voila, the image generated by the opentrons simulation
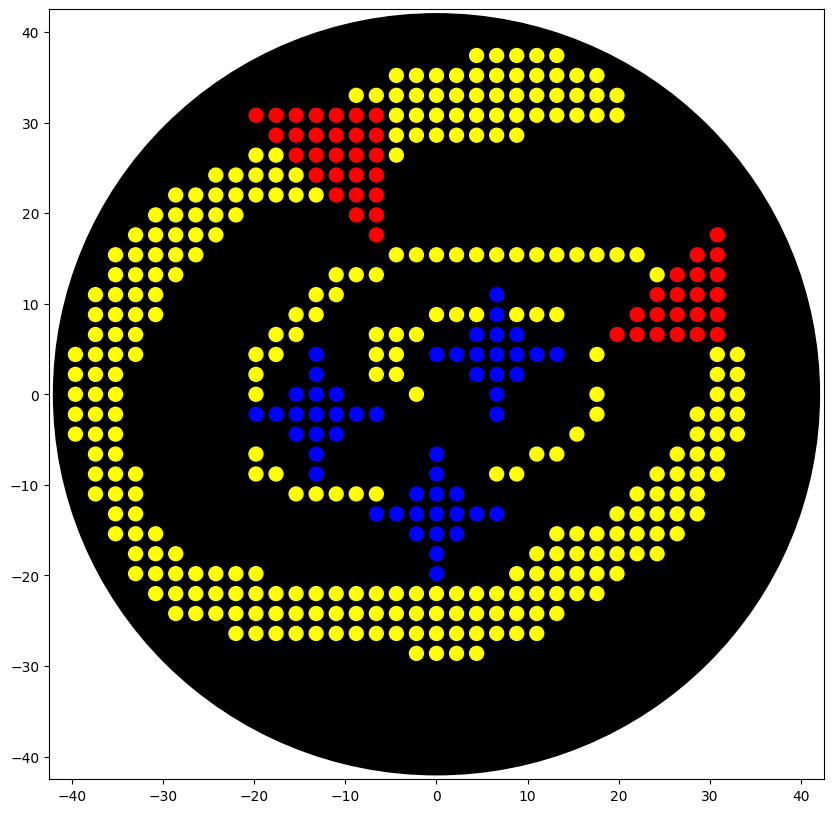

Post-Lab Questions
- Find and describe a published paper that utilizes the Opentrons or an automation tool to achieve novel biological applications.
- Write a description about what you intend to do with automation tools for your final project
Published paper that uses automation: There are various providers of lab automation. Be it opentrons, trilobio, tetsuwan scientific or ginkgo bioworks.
- This latest paper describes using GPT 5 and ginkgo bioworks autonomous lab to do cell-free protein synthesis: Using a GPT-5-driven autonomous lab to optimize the cost and titer of cell-free protein synthesis | bioRxiv
- In this paper, the team was able to achieve a 27% increase in protein titer, and a 40% reduction in cost of cell free protein synthesis (CFPS) in $/gram. Protein of choice was sfGFP - super green fluorescent protein
- Pydantic was used to validate that the AI generated protocols were valid before sending it to ginkgo bioworks.
LLM generated:
- programmatic specification of multi-instrument biological workflows by Ginkgo’s Catalyst software
- Use YAML/XML output from ginkgo to redesign and optimize the experiment further.
Description of what I intend to do: I want to use GPT5 and Ginkgo’s RAC to optimize protein-MOF symbiosis in ginkgo’s RAC automatic lab.
I uploaded the paper into ChatGPT, asked it to program itself to design an experiment for the above.
Output:
Ginkgo catalyst orchestration system JSON workflow graph:
High-level protein workflow pseudocode:
Final Project Ideas
- Update committed listener project page under tokyo bioclub My ideas all revolve around synbio for space inspired by this paper: https://www.nature.com/articles/s41467-023-37910-1
- Make in earth for space - Send things we couldn’t send before to space in new ways.
- Make in space for earth - Protein crystals and APIs that can only be made in microgravity
- Make in space for space - Hardware to enable space synbio slides yet to be updated
few more ideas I have been exploring
- exploring how MOFs and proteins go hand in hand
- dealing with composites that have in built bio-sensors.
- bio-electric connections
Committed listener page has been updated.
Final ideas submitted:
- Bio-hybrid Reef builder Robot
- Protein@MOFs cell-free system for extreme habitat stability
- Cell free On-chip cartridge framework for modular exobiology research instrumentation
Personal slides link (could have more than 3 ideas, but in shared slide, only 3 are shared): https://docs.google.com/presentation/d/14bcZB3e4qWsHIfbFh_S83iBtNpFeC-ODNHlv9NLe4sI/edit?usp=sharing
Sources
Using a GPT-5-driven autonomous lab to optimize the cost and titer of cell-free protein synthesis
Alexus A. Smith, Edmund L. Wong, Ronan C. Donovan, Brad A. Chapman, Ryan Harry, Pooyan Tirandazi, Paulina Kanigowska, Elizabeth A. Gendreau, Robert H. Dahl, Michal Jastrzebski, Jose E. Cortez, Christopher J. Bremner, José C. Morales Hemuda, James Dooner, Ian Graves, Rahul Karandikar, Christopher Lionetti, Kevin Christopher, Andrew L. Consiglio, Alyssa Tran, William McCusker, Duy X. Nguyen, Isis Botelho Nunes da Silva, Alvaro R. Bautista-Ayala, Monica P. McNerney, Sean Atkins, Michael McDuffie, Will Serber, Bradley P. Barber, Trinh Thanongsinh, Andrew Nesson, Bibek Lama, Brandon Nichols, Cameron LaFrance, Tenzing Nyima, Alicia Byrn, Rashard Thornhill, Bryan Cai, Lizvette Ayala-Valdez, Alycia Wong, Austin J. Che, Walter Thavarajah, Daniel Smith, Thomas F. Knight Jr., David W. Borhani, Jerry Tworek, Mostafa Rohaninejad, Ahmed El-Kishky, Nathan C. Tedford, Tejal Patwardhan, Yunxin Joy Jiao, Reshma P. Shetty
bioRxiv 2026.02.05.703998; doi: https://doi.org/10.64898/2026.02.05.703998
My chatGPT chat asking it to design an experiment: https://chatgpt.com/share/69ad5492-7f1c-8003-87aa-b1758c36f74b