Week 4 Lab & HW
Part A
Why humans eat beef but do not become a cow, eat fish but do not become fish?
Part B: Protein Analysis and Visualization
In this part of the homework, you will be using online resources and 3D visualization software to answer questions about proteins. Pick any protein (from any organism) of your interest that has a 3D structure and answer the following questions.
fast-petase
Briefly describe the protein you selected and why you selected it. I selected fast-petase because im considering using it in my final project. I am curious to learn more about the enzymatic breakdown of PET plastics.
Identify the amino acid sequence of your protein.
MQTNPYARGPNPTAASLEASAGPFTVRSFTVSRPSGYGA GTVYYPTNAGGT VGAIAIVPGYTARQSSIKWWGPRLAS HGFVVITIDTNSTLDQPESRSSQQMAALRQVASLNGTS SSPIY GKVDTARMGVMGWSMGGGGSLISAANNPSLKAA APQAPWHSSTNFSSVTVPTLIFACENDSIAI PVNSSAL PIYDSMSNAKQFLEIKGGSHSCANSG NSNQALIGKKG VAWMKRFMDNDTRYSTFACENPNSTAVSDFRTANCSLE HHHHHH
How long is it? What is the most frequent amino acid? You can use this notebook to count most frequent amino acid - https://colab.research.google.com/drive/1vlAU_Y84lb04e4Nnaf1axU8nQA6_QBP1?usp=sharing
819 bp, Serine, which has 37 occurences
How many protein sequence homologs are there for your protein? Hint: Use the pBLAST tool to search for homologs and ClustalOmega to align and visualize them. Tutorial Here
2,930 PETase homologs

Does your protein belong to any protein family?
Cutinase / Dienelactone hydrolase-like
Identify the structure page of your protein in RCSB When was the structure solved? Is it a good quality structure? Good quality structure is the one with good resolution. Smaller the better (Resolution: 2.70 Å) Method: X-RAY DIFFRACTION Resolution: 1.57 Å R-Value Free: 0.140 (Depositor), 0.141 (DCC) R-Value Work: 0.106 (Depositor), 0.107 (DCC) R-Value Observed: 0.108 (Depositor)
Are there any other molecules in the solved structure apart from protein?
The solved crystal structure of FAST-PETase (PDB: 7SH6) contains one Sulfate ion (SO4) molecule besides the protein chain
Does your protein belong to any structure classification family?
Hydrolase
Open the structure of your protein in any 3D molecule visualization software:
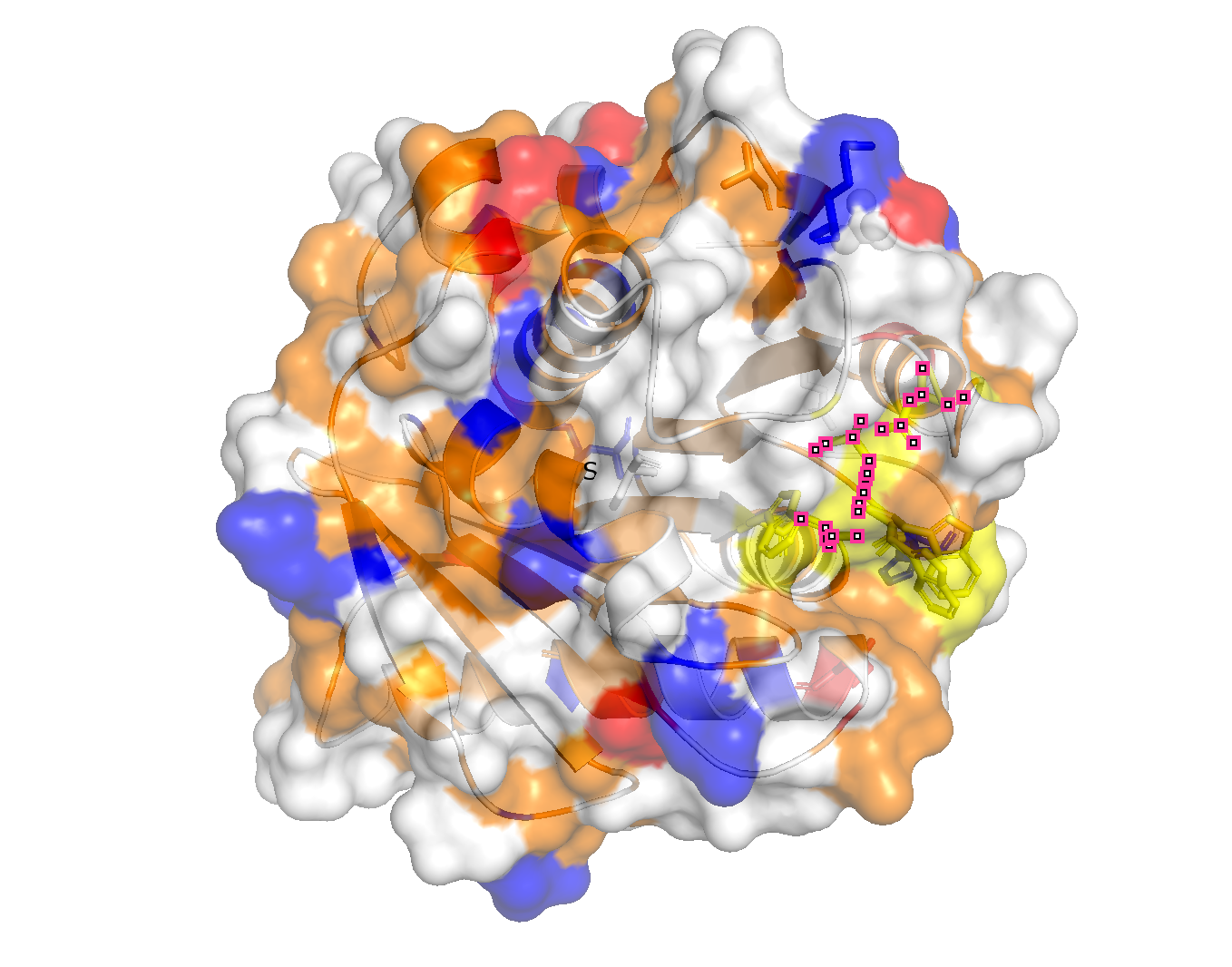
PyMol Tutorial Here (hint: ChatGPT is good at PyMol commands) Visualize the protein as “cartoon”, “ribbon” and “ball and stick”. Color the protein by secondary structure. Does it have more helices or sheets? is organized into 9 β-strands (yellow) and 7 α-helices, so more B-strands. Color the protein by residue type. What can you tell about the distribution of hydrophobic vs hydrophilic residues? The hydrophobic tend towards the center (orange) and the hydrophilic towards the outer surface.
Visualize the surface of the protein. Does it have any “holes” (aka binding pockets)? There is one valley in the outer surface as the PET-binding cleft
Part C. Using ML-Based Protein Design Tools
HTGAA ProteinDesign2025 Fold your protein with AlphaFold or ESMFold or Boltz and compare it to the real structure. Comment on: Any predicted vs. experimental differences. It has a 90%+ predicted accuracy for nearly everything Low-confidence regions and why do you think they are low confidence? The main low-confidence areas on the loop regions and the His-tag tail.
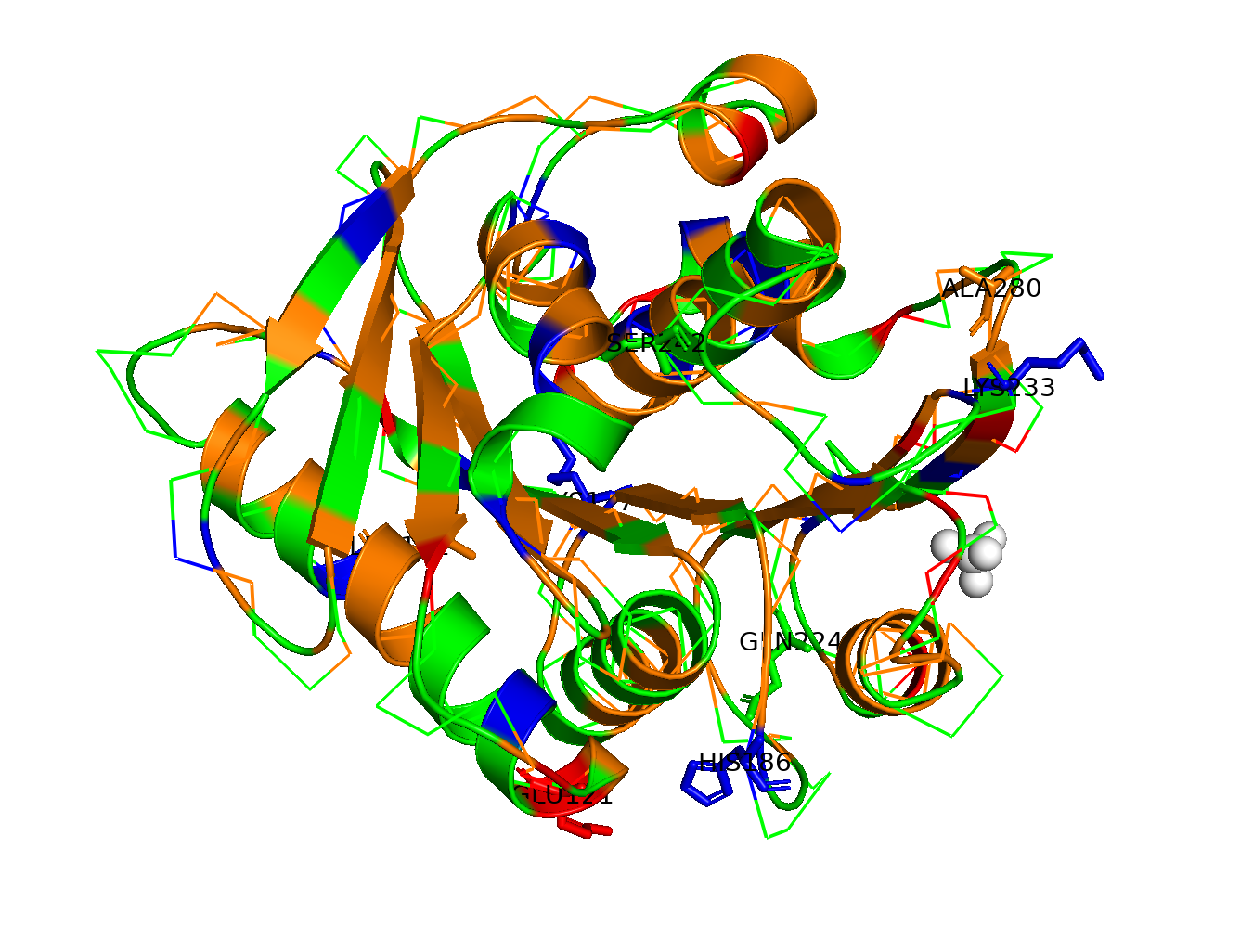
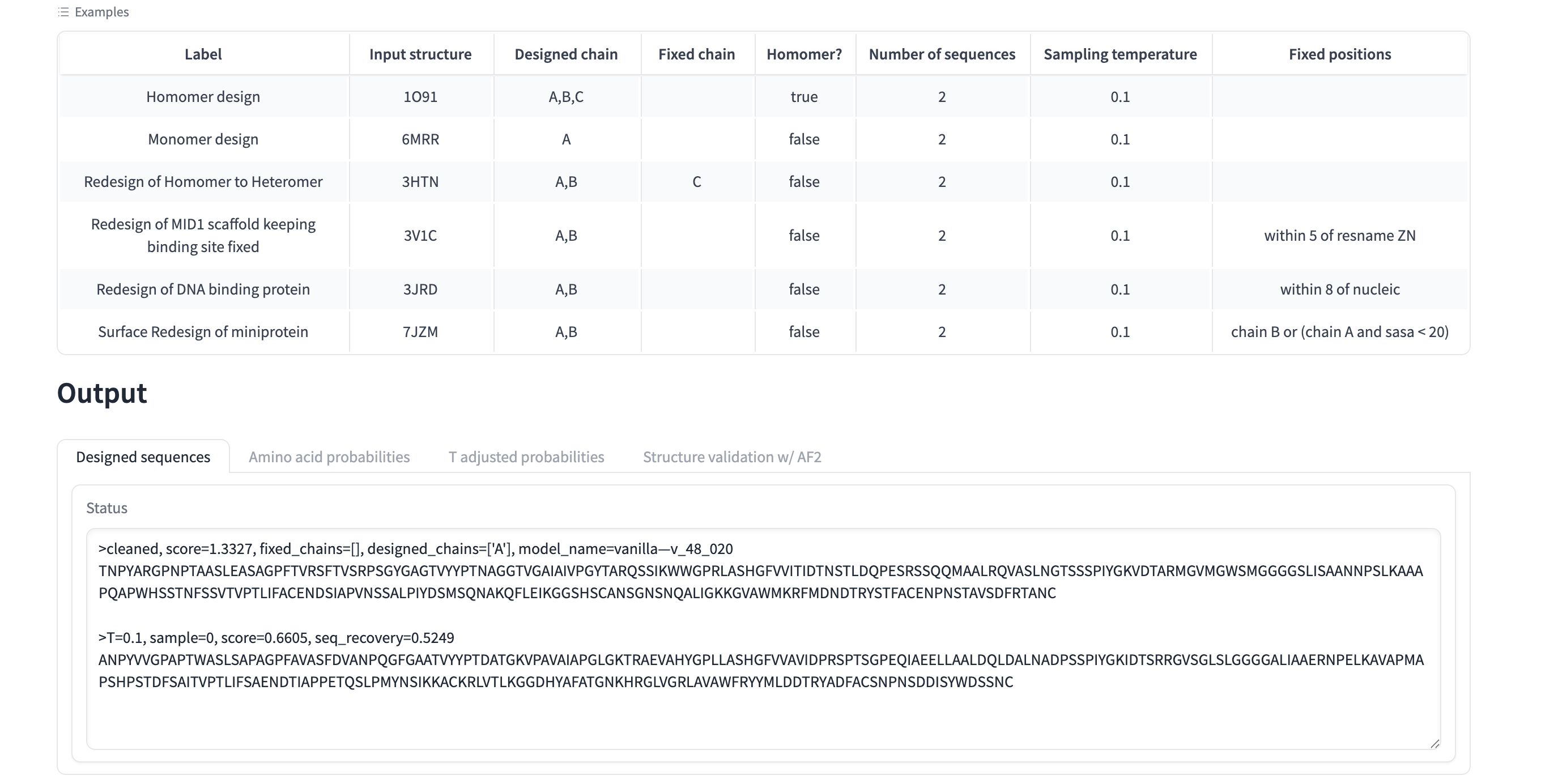
Inverse-fold your structure with ProteinMPNN What sequence do you get?
cleaned, score=1.3327, fixed_chains=[], designed_chains=[‘A’], model_name=vanilla—v_48_020 TNPYARGPNPTAASLEASAGPFTVRSFTVSRPSGYGAGTVYYPTNAGGTVGAIAIVPGYTARQSSIKWWGPRLASHGFVVITIDTNSTLDQPESRSSQQMAALRQVASLNGTSSSPIYGKVDTARMGVMGWSMGGGGSLISAANNPSLKAAAPQAPWHSSTNFSSVTVPTLIFACENDSIAPVNSSALPIYDSMSQNAKQFLEIKGGSHSCANSGNSNQALIGKKGVAWMKRFMDNDTRYSTFACENPNSTAVSDFRTANC
T=0.1, sample=0, score=0.6605, seq_recovery=0.5249 ANPYVVGPAPTWASLSAPAGPFAVASFDVANPQGFGAATVYYPTDATGKVPAVAIAPGLGKTRAEVAHYGPLLASHGFVVAVIDPRSPTSGPEQIAEELLAALDQLDALNADPSSPIYGKIDTSRRGVSGLSLGGGGALIAAERNPELKAVAPMAPSHPSTDFSAITVPTLIFSAENDTIAPPETQSLPMYNSIKKACKRLVTLKGGDHYAFATGNKHRGLVGRLAVAWFRYYMLDDTRYADFACSNPNSDDISYWDSSNC
Is it the same as the original sequence you folded? No, only 52% of the sequence is the same Why yes or no? This is possible because many different amino acid combinations can produce the same protein structure, its the structure that matters more than the combinations themselves.
Part D. Group Brainstorm on Bacteriophage Engineering
Find a group of ~3–4 students Review the Bacteriophage Final Project Goals: Increased stability (easiest) Higher titers (medium) Higher toxicity of lysis protein (hard) Brainstorm Session Choose one or two main goals from the list that you think you can address computationally (e.g., “We’ll try to stabilize the lysis protein,” or “We’ll attempt to disrupt its interaction with E. coli DnaJ.”). Write a 1-page proposal (bullet points or short paragraphs) describing: Which tools/approaches from recitation you propose using (e.g., “Use Protein Language Models to do in silico mutagenesis, then AlphaFold-Multimer to check complexes.”). Why you think those tools might help solve your chosen sub-problem. One or two potential pitfalls (e.g., “We lack enough training data on phage–bacteria interactions.”). Include a schematic of your pipeline This resource may be useful: HTGAA Protein Engineering Tools Individually put your plan on your website page Each group’s short plan for engineering a bacteriophage Schedule time ( HTGAA Protein Engineering Feedback) to get feedback/discuss your ideas, and put the feedback on your website