Week 1 HW: Principles and Practices

 Systemic Lupus Erythematosus (SLE) disproportionately affects women at a 9:1 female/male ratio (McDonald et al., 2015), with higher prevalence in Latin American populations compared to European ancestry groups (Ugarte-Gil et al., 2023). In Ecuador, qualitative studies document significant barriers to SLE treatment and care for women in rural and Andean Indigenous communities, where cultural mismatches with Western medicine delay diagnosis and treatment (Miles, 2011; Bautista-Valarezo et al., 2021). Current immunosuppressive therapies require frequent clinical monitoring unavailable in these settings, leading to uncontrolled disease activity and organ damage.
Systemic Lupus Erythematosus (SLE) disproportionately affects women at a 9:1 female/male ratio (McDonald et al., 2015), with higher prevalence in Latin American populations compared to European ancestry groups (Ugarte-Gil et al., 2023). In Ecuador, qualitative studies document significant barriers to SLE treatment and care for women in rural and Andean Indigenous communities, where cultural mismatches with Western medicine delay diagnosis and treatment (Miles, 2011; Bautista-Valarezo et al., 2021). Current immunosuppressive therapies require frequent clinical monitoring unavailable in these settings, leading to uncontrolled disease activity and organ damage.
I want to propose the design of an in vitro prototype of a synthetic mammalian gene circuit. The idea is that the circuit autonomously senses and regulates type I interferon (IFN-α) levels—a validated biomarker of poor outcomes- elevated in over 70% of systemic lupus erythematosus (SLE) adult patients (Whittall Garcia et al., 2024). When IL-10 is chronically exposed to IFN-α in vivo, it becomes part of a pro-inflammatory signaling, instead of its original anti-inflammatory function (Yuan et al., 2011). So, by detecting and regulating IFN- α levels, it will be possible to restore IL-10 anti-inflammatory properties and improve patients’ outcomes. Communities already distrust the traditional healthcare system, so maybe they would be more receptive to this approach, that for them may seem less invasive and restrictive if conducted correctly.
A primary governance goal necesary for developing the project would be to: Ensure bioethical, development of autonomous therapies for diseases affecting marginalized women.
To reach it, I consider the following subgoals:
- Bioethical guided protocol and trial design: Before human trials, replicability and circuit stability should be guaranteed at least in cellular lines. Bioethics are of utmost importance when designing a biomedical protocol, even more when treating vulnerable groups (e.g. indigenous women). All research decisions are to be made taking into consideration patients’: a) direct personal gains/ benefits from the trial; b) informed consent and willingness to be involved in the trial; and c) rights and personal safety.
- Culturally grounded consent: Develop dynamic consent frameworks respecting communal decision-making models in Indigenous Kichwa communities (Bautista-Valarezo et al., 2021).
- Rural accessibility: Guarantee technology pathways reach women in communities >2 hours from tertiary hospitals.
- Prevent therapeutic abandonment: Ensure patients receive ongoing support if circuit recalibration is needed.
Analyzing the scope of the project, derivated three ordenance actions that meet real concerns came to mind:
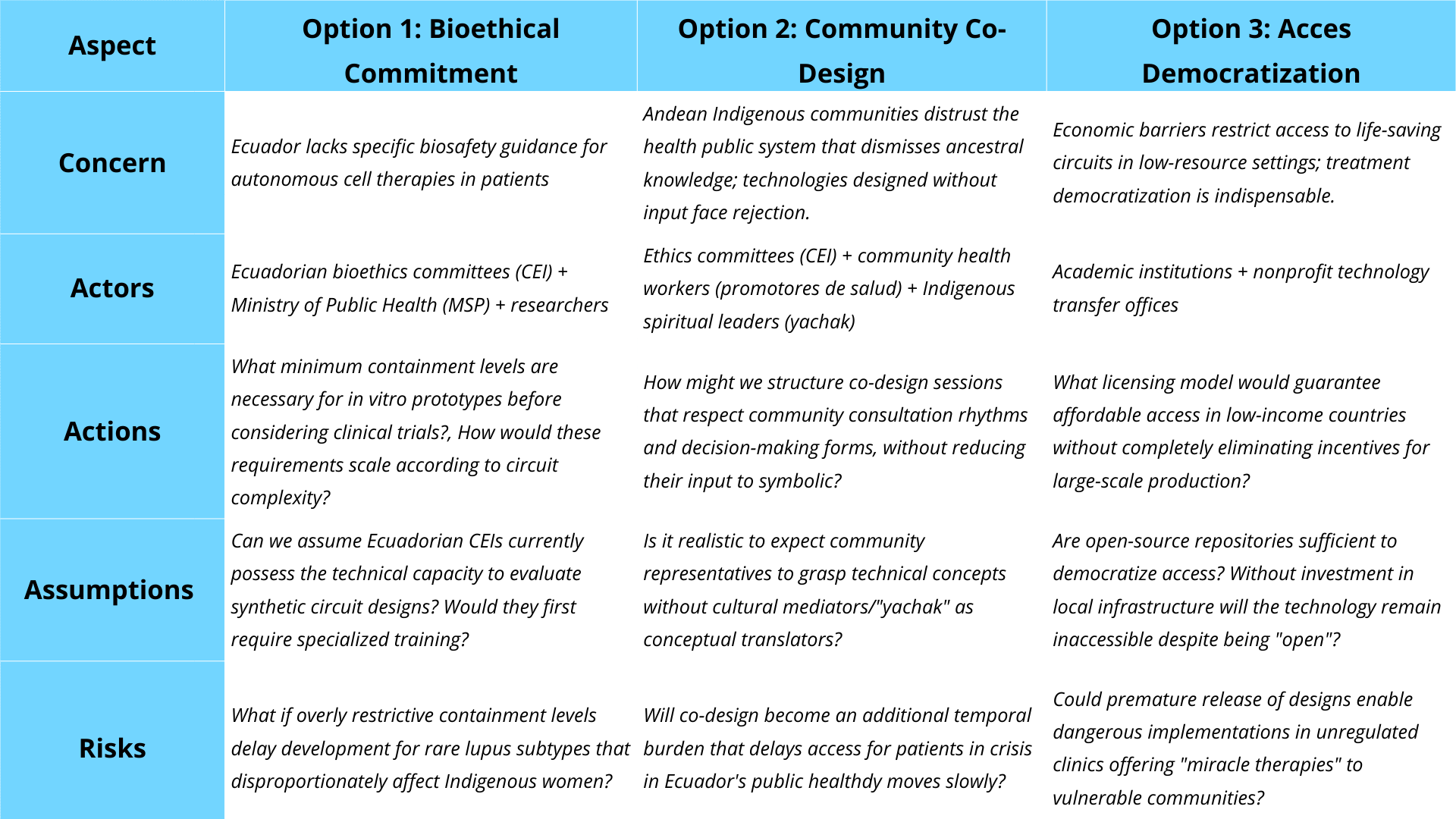

To implement the three ordenance actions, they were evaluated in function of how they follow the proposed Governance’s criterion (from 1 to 3, with 1 as the best):
| Governance Criterion | Option 1: Tiered Containment | Option 2: Co-Design | Option 3: Open-Source |
|---|---|---|---|
| Bioethical trial design | |||
| • Ensures replicability | ●●○ (1) | ○○○ (3) | ●○○ (2) |
| • Guarantees patient safety | ●●○ (1) | ●○○ (2) | ○○○ (3) |
| Culturally-grounded consent | |||
| • Respects communal decisions | ○○○ (3) | ●●○ (1) | ○○○ (3) |
| • Integrates ancestral knowledge | ○○○ (3) | ●●○ (1) | ○○○ (3) |
| Rural accessibility | |||
| • Reaches remote communities | ○○○ (3) | ●○○ (2) | ●●○ (1) |
| • Minimizes economic barriers | ○○○ (3) | ○○○ (3) | ●●○ (1) |
| Prevent therapeutic abandonment | |||
| • Ensures ongoing support | ●○○ (2) | ●○○ (2) | ●●○ (1) |
| • Guarantees long-term monitoring | ●○○ (2) | ○○○ (3) | ○○○ (3) |
| Cross-cutting considerations | |||
| • Feasibility in Ecuador | ●○○ (2) | ●○○ (2) | ○○○ (3) |
| • Does not impede research | ●○○ (2) | ○○○ (3) | ●●○ (1) |
| • Minimizes costs | ○○○ (3) | ●○○ (2) | ●●○ (1) |
Based on the scoring framework above, I would prioritize a sequential implementation strategy: Option 1 (Bioethical Commitment) and Option 2 (Community Co-Design) as mandatory requirements during the research and early clinical phases, with Option 3 (Access Democrtization) introduced only after successful Phase II clinical validation. This sequencing addresses the fundamental trade-off between safety and accessibility—while open-source democratization (Option 3) scores highest for rural accessibility and minimizing economic barriers, its weak performance in patient safety (score 3) makes premature implementation ethically unacceptable for an autonomous therapy affecting vulnerable indigenous women. Containment requirements (Option 1) provide the non-negotiable safety foundation (scores 1,1 for replicability and patient safety), while co-design (Option 2) ensures cultural legitimacy by centering communal decision-making models (scores 1,1 for culturally-grounded consent)—a critical factor given documented Indigenous distrust of medical systems that dismiss ancestral knowledge (Bautista-Valarezo et al., 2021). The primary trade-off I acknowledge is timeline extension: mandatory co-design workshops and dual-redundant kill switch validation will delay deployment compared to a purely technical development path. However, this delay prevents two greater harms: (a) therapeutic abandonment if communities reject culturally inappropriate designs, and (b) catastrophic safety failures from insufficient containment.
My recommendation targets three audiences simultaneously: (1) locally, the Ethics Committees (CEI) of hospitals collaborating with universities’ CEI should adopt co-design requirements immediately; (2) nationally, Ecuador’s Ministry of Public Health (MSP) must develop tiered containment guidelines specific to synthetic biology therapies, preferably within a set time frame; and (3) internationally, the WHO Ethics Advisory Committee should enforce global standards recognizing that “informed consent” for autonomous therapies must accommodate communal decision-making structures in Indigenous contexts. This phased approach honors both the precautionary principle and justice: safety first, then equitable access, never the reverse.
REFLECTING ON WEEK 1,
There was a new ethical concern (for me) that emerged: The tension between the developing circuit regulated therapies for a patient, and the status quo in decision-making embedded in family/community networks that may forbid the patient access to treatment.
In Ecuadorian indigenous communities, health decisions often involve extended family consultation, especially when there is a preexistent prejudice with the local public system. An autonomous circuit designed based on individualism might undermine culturally valued collective decision-making.
As stated before, this requires governance actions that bring together key members first, to reach the whole community. On a more detailed note, circuit designers can reach a middle ground with the indigenous by showcasing how their system interfaces can be linked with existing social decision-making structures. For example: Could the circuit include a “real-time feedback” function to let family members know if an input (e.g. related to their traditional medical care system, or whether to activate a kill switch) could aid the patient? With this I have learned that deployment strategy is as important as technology and funding to effectively reach target communities.
This exercise revealed that what we perceive as autonomy is not a universal value but culturally constructed. Ethical synthetic biology requires not just technical safety, but cultural open mindedness—designing with communities rather than for them. As future builders of biological systems, we must ask not only “Can we build this?” but “Should we build it this way, for these people, in this context?”
Homework Questions from Professor Jacobson:
Nature’s machinery for copying DNA is called polymerase. What is the error rate of polymerase? How does this compare to the length of the human genome. How does biology deal with that discrepancy? The slides (p.8 & p.10) state that polymerase has an error rate of ~1:10⁶ (1 mistake per million base pairs) and that the human genome length has a length of 3.2 Gbp (which could be expressed as 3.2 × 10⁹ bp). Thus, in humans, each replication generates approximately 3,200 mistakes. Biology –not synthetic biology- copes with this by relying on 3’→5’ proofreading, a mechanism where polymerase detects incorrect base pairings, retires the wrong nucleotide and replaces it as needed.
How many different ways are there to code (DNA nucleotide code) for an average human protein? In practice what are some of the reasons that all of these different codes don’t work to code for the protein of interest? According to slide 6, the average human protein length is 1036 bp. A codon (a base pair triplets) codes for one protein. Assuming that the given protein length on the slide excluded non-coding base pairs of the gene, we have: 1036/ 3= ~345 amino acids. The protein length is fixedly of ~345 amino acids, however due to codon degeneracy, more than one base pair triplet combination/codon can code for the same amino acid (e.g. both AGA and AGG code for Arg). So, with 3 codons per amino acid and 345 amino acid “slots” an average human protein can have: 3³⁴⁵ possible coding ways. In nature, a huge percentage of these coding ways are not viable, as there exists a codon usage bias, set by natural selection, to improve efficiency and precision in translation. The slides (p.39- p. 43) show that regions with a high content of GC (and AT) pairs form hairpin-like structures. These interfere with transcription/ translation as they can act as signaling regions, inducing splicing or stopping transcription/translation. Thus, maybe codon usage bias intents to keep G-C % to a minimum.
Homework Questions from Dr. LeProust:
What’s the most commonly used method for oligo synthesis currently? Why is it difficult to make oligos longer than 200nt via direct synthesis? Why can’t you make a 2000bp gene via direct oligo synthesis? In the slides (p.11) the first automated method of oligo synthesis was, proposed by ABI (Applied Biosystems), with subsequent optimizations. However, it was mainly based on the CPG method, which in turn uses the phosphoramidite method, both designed by Caruthers (Matteucci & Caruthers, 1981). So, at its core, the most used is the phosphoramidite method. The cycle consists of phosphoramidite-based synthesis consists of four steps repeated per nucleotide addition: -Coupling: Phosphoramidite addition -Capping: Blocking unreacted sites -Oxidation: Stabilizing the bond -Deblock: Removing the protecting group for the next cycle The yield per cycle is determined by the coupling efficiency and the length (in bases) of the oligonucleotide by the formula:
So, even if coupling efficiency always reaches a 90.0%-99.5% range, the longer the length, the poorer the yield (Merck, 2026). This limitation is shown when applying the formula to an oligo with a length of 200 bases at 99,5%: Yield= 0.995200 ^ 200= 0.367= 36.7% The formula works as it explains that in each cycle, while 99% of the oligonucleotide chains are synthetized, the remaining 1% is truncated and starts to accumulate. Only 0.995 ^ (n cycles) of the original chains reach complete length.
Based on that logic, if a direct oligo synthesis was performed for an oligo with a 2000 bp length, the yield would not be significative (<0.001%). On the other hand, by following the multiplexed gene fragment solution proposed on the slides, the overall yield increases, making synthesis more accurate as a whole.
Homework Question from George Church:
Given slides #2 & 4 (AA:NA and NA:NA codes), what code would you suggest for AA:AA interactions? Nucleic acid/Nucleic acid (NA:NA) code refers to the nucleotides forming determined/ fixed base pairs that will always bond with one another: “A” bonds with “T” (“U” in RNA), and “C” bonds with “G”. Based on this empirical rule, this type of code becomes a programable basepair code, allowing us to design a sequence with certainty. In the case of Amino acid/ Nucleic acid (AA:NA) code, the rule is that the ribosome can translate one stranded sequence - by reading codons, base triplets- into amino acids. Each codon codes for a specific amino acid, allowing the synthesis of bigger proteins. However, in translation, there is more than one codon that can be translated into the same amino acid (codon degeneracy). In that line, the amino acid/ amino acid (NA: NA) code should be the set, deterministic rule that describes all amino acid interactions. Protein primary structures result from a covalent bond between the N terminus of one to the c terminus of another. Nevertheless, there is not such a thing as a generalized fixed way of interaction among amino acids, due to the presence of their variable side chains. The functional groups make the side chains unique, directly giving amino acid their properties: polarity, charge/ attraction (covalent bonds) and the ability to form hydrogen bridges. When interacting, those properties will make interactions-that cause secondary, tertiary and quaternary structures- no deterministic, but statistical and contextual. So, the difficulty of assigning a code to amino acid interactions lies mainly in the variability introduced by the side chain. And that same variability what makes them form functional proteins. Bertozzi biorthogonal reactions occur independently without affecting the natural biochemistry/natural reactions of molecules in the cell. In them a “clic” reaction occurs between an azide and an alkyne cycle, resulting in in a new compound that is a union of both compounds, like “connecting to parts of a seatbelt” (Agard et al., 2005). Now this technology is widely used for fluorescent marking and observation of cellular processes. But maybe we could go an step further, and directly explore how modifying side chains to have azide and alkyne cycles as functional groups impact in improving predictability of secondary, tertiary and quaternary structures.
REFERENCES
- Agard, N. J., Prescher, J. A., & Bertozzi, C. R. (2005). A strain-promoted [3 + 2] Azide−Alkyne cycloaddition for covalent modification of biomolecules in living systems [J. am. Chem. Soc. 2004, 126, 15046−15047]. Journal of the American Chemical Society, 127(31), 11196–11196. https://doi.org/10.1021/ja059912x
- Bautista-Valarezo, E., Duque, V., Verhoeven, V., Mejia Chicaiza, J., Hendrickx, K., Maldonado-Rengel, R., & Michels, N. R. M. (2021). Perceptions of Ecuadorian indigenous healers on their relationship with the formal health care system: barriers and opportunities. BMC Complementary Medicine and Therapies, 21(1), 65. https://doi.org/10.1186/s12906-021-03234-0
- Matteucci, M. D., & Caruthers, M. H. (1981). Synthesis of deoxyoligonucleotides on a polymer support. Journal of the American Chemical Society, 103(11), 3185–3191. https://doi.org/10.1021/ja00401a041
- McDonald, G., Cabal, N., Vannier, A., Umiker, B., Yin, R. H., Orjalo, A. V., Jr, Johansson, H. E., Han, J.-H., & Imanishi-Kari, T. (2015). Female bias in systemic lupus erythematosus is associated with the differential expression of X-linked toll-like receptor 8. Frontiers in Immunology, 6, 457. https://doi.org/10.3389/fimmu.2015.00457
- Merck. (2026). Polymerase Chain Reaction ApplicationsDNA Oligonucleotide Synthesis DNA Oligonucleotide Synthesis. Sigmaaldrich.com. https://www.sigmaaldrich.com/EC/en/technical-documents/technical-article/genomics/pcr/dna-oligonucleotide-synthesis
- Miles, A. (2011). Emerging chronic illness: women and lupus in Ecuador. Health Care for Women International, 32(8), 651–668. https://doi.org/10.1080/07399332.2010.529214
- Ugarte-Gil, M. F., Fuentes-Silva, Y., Pimentel-Quiroz, V. R., Pons-Estel, G. J., Quintana, R., Pons-Estel, B. A., & Alarcón, G. S. (2022). Global excellence in rheumatology in Latin America: The case of systemic lupus erythematosus. Frontiers in Medicine, 9, 988191. https://doi.org/10.3389/fmed.2022.988191
- Whittall Garcia, L. P., Gladman, D. D., Urowitz, M., Bonilla, D., Schneider, R., Touma, Z., & Wither, J. (2024). Interferon-α as a biomarker to predict renal outcomes in lupus nephritis. Lupus Science & Medicine, 11(2). https://doi.org/10.1136/lupus-2024-001347
- Yuan, W., DiMartino, S. J., Redecha, P. B., Ivashkiv, L. B., & Salmon, J. E. (2011). Systemic lupus erythematosus monocytes are less responsive to interleukin-10 in the presence of immune complexes. Arthritis and Rheumatism, 63(1), 212–218. https://doi.org/10.1002/art.30083