Week 2 — DNA Read, Write, & Edit
Part 0: Basics of Gel Electrophoresis
I reviewed this week’s lecture and recitation videos, which provide excellent foundational explanations of gel electrophoresis principles. Having hands-on experience running agarose gels in my research lab, I can confirm that all the technical considerations mentioned are crucial for successful results. One safety tip that particularly resonated with me was handling hot flasks after microwave use. In our lab, we always used heat-resistant gloves, yet we were still advised to minimize contact time with hot containers—a precaution I now understand is essential.
Part 1: Benchling & In-silico Gel Art
I began by locating the Lambda DNA identifier in NCBI’s Genome database. My search strategy involved the keywords “bacteriophage lambda” and the known genome length (48,502 bp). The search returned three results; I selected the first entry because its Assembly status indicated “Viral Project,” which typically signifies a complete genome sequencing effort. Upon verification, the genome size matched the expected 48.5 kb. From this entry, I extracted the GenBank identifier (J02459.1) and directly imported the sequence into Benchling via the “DNA/RNA sequence” import function.
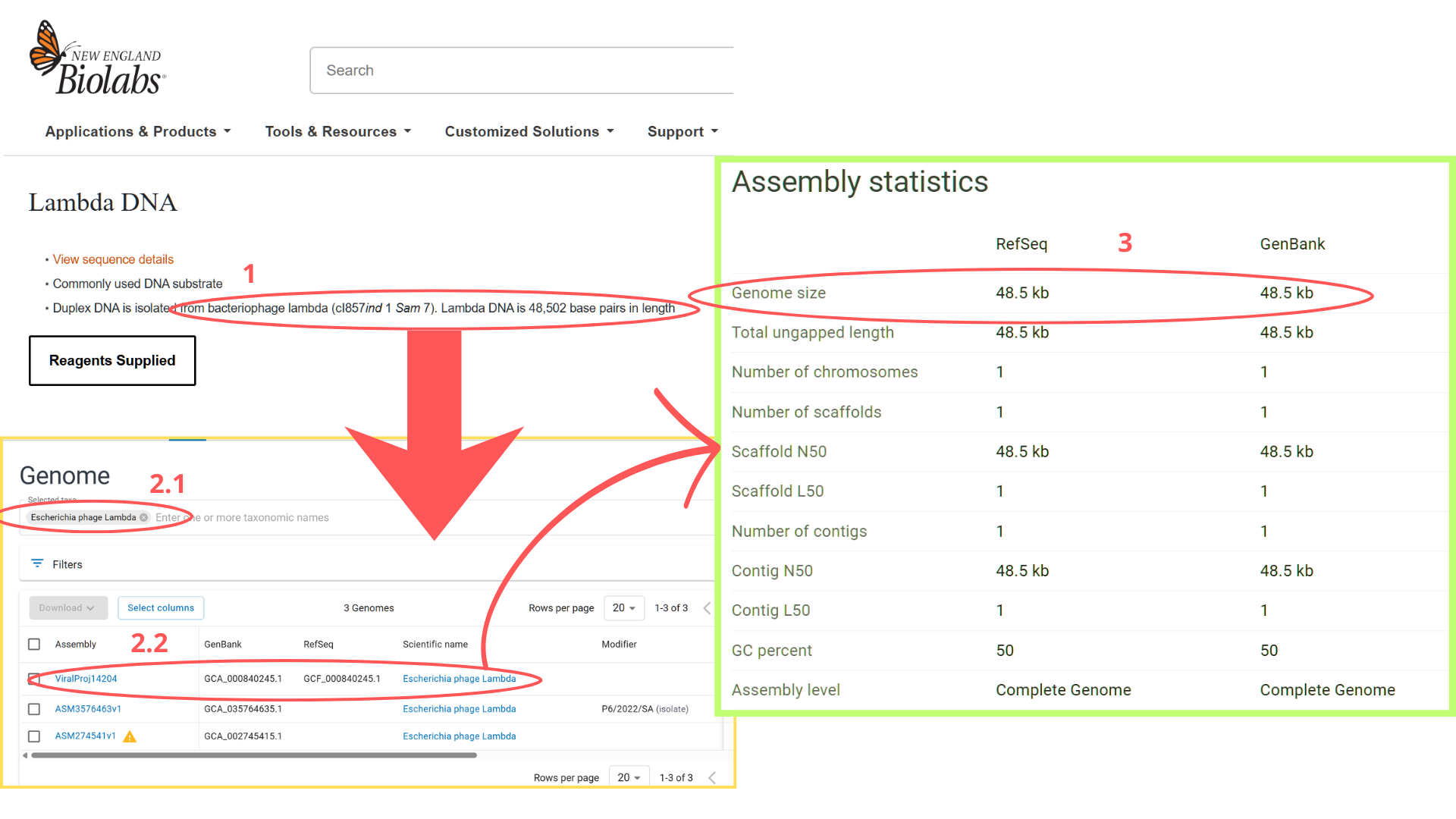

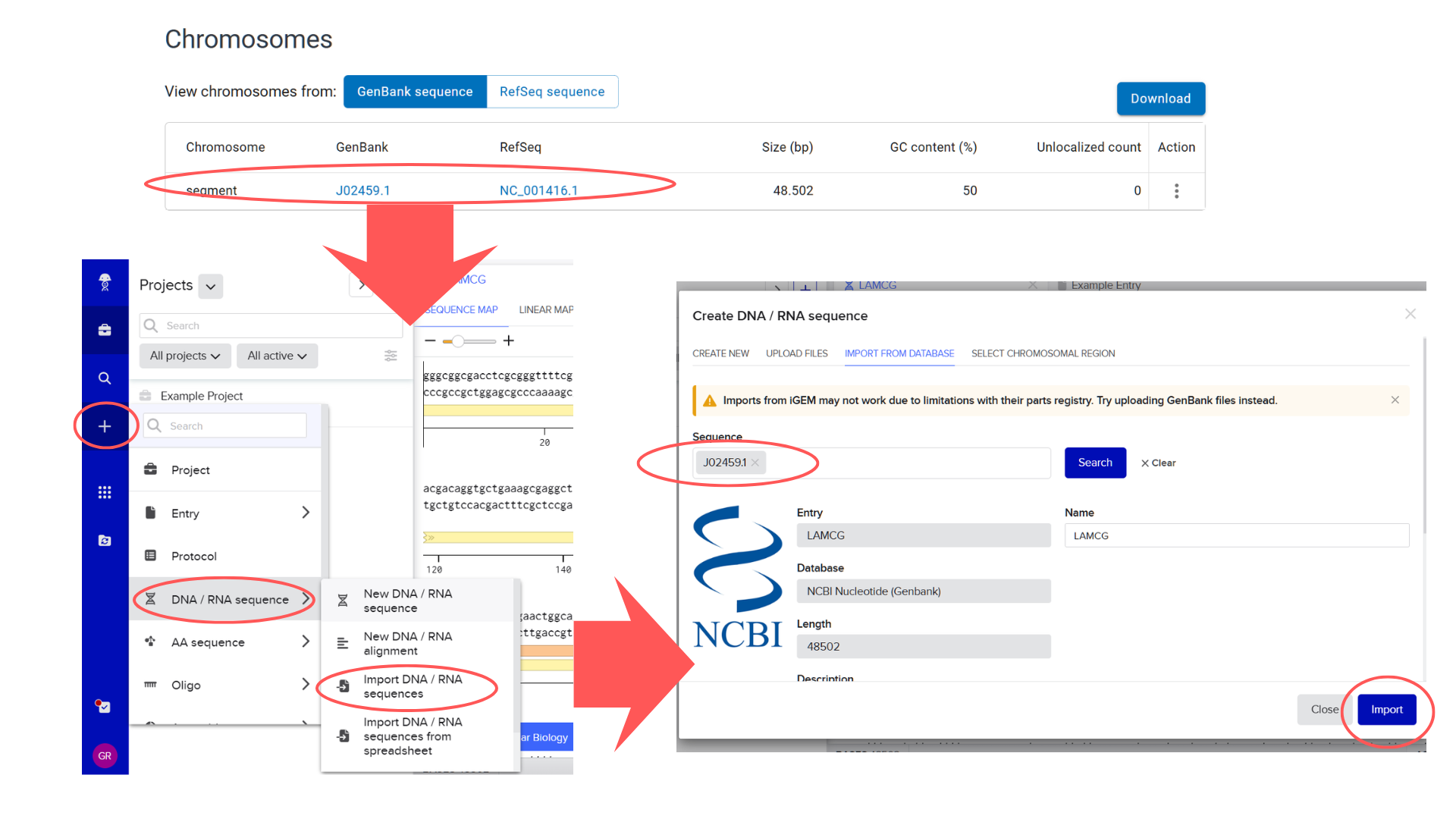

Considering the genome’s 48.5 kb size, I initially planned to use a 50 kb ladder for reference. However, after performing a test restriction digest with EcoRV, HindIII, SacI, KpnI, and SalI, Benchling’s fragment table revealed that all resulting fragments were under 5 kb.
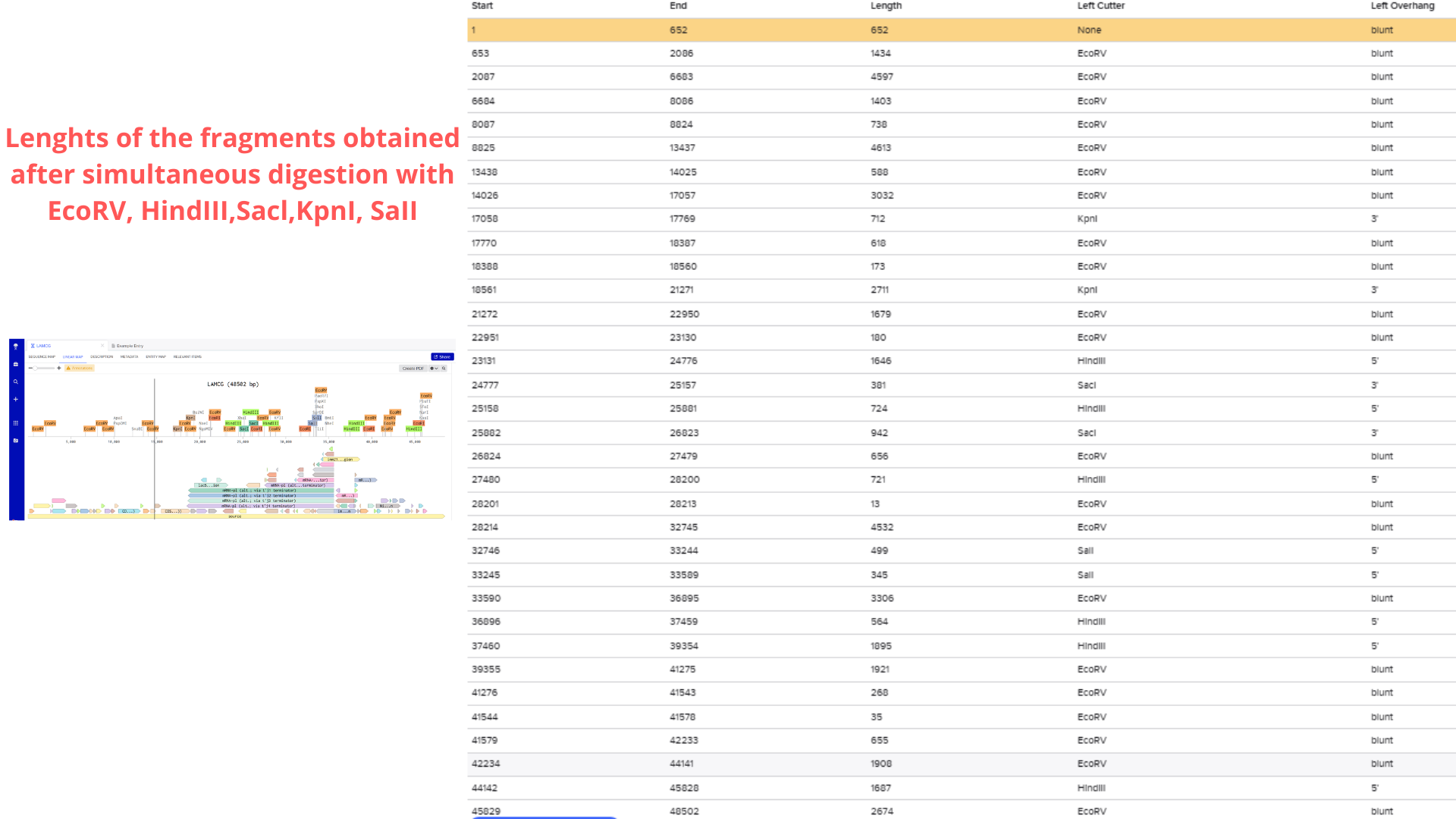

Consequently, I switched to a 5 kb ladder (Geneon’s “DNA Ladder mix - high range”) for my gel art design. The final pattern aims to form a heart shape, as visualized below:
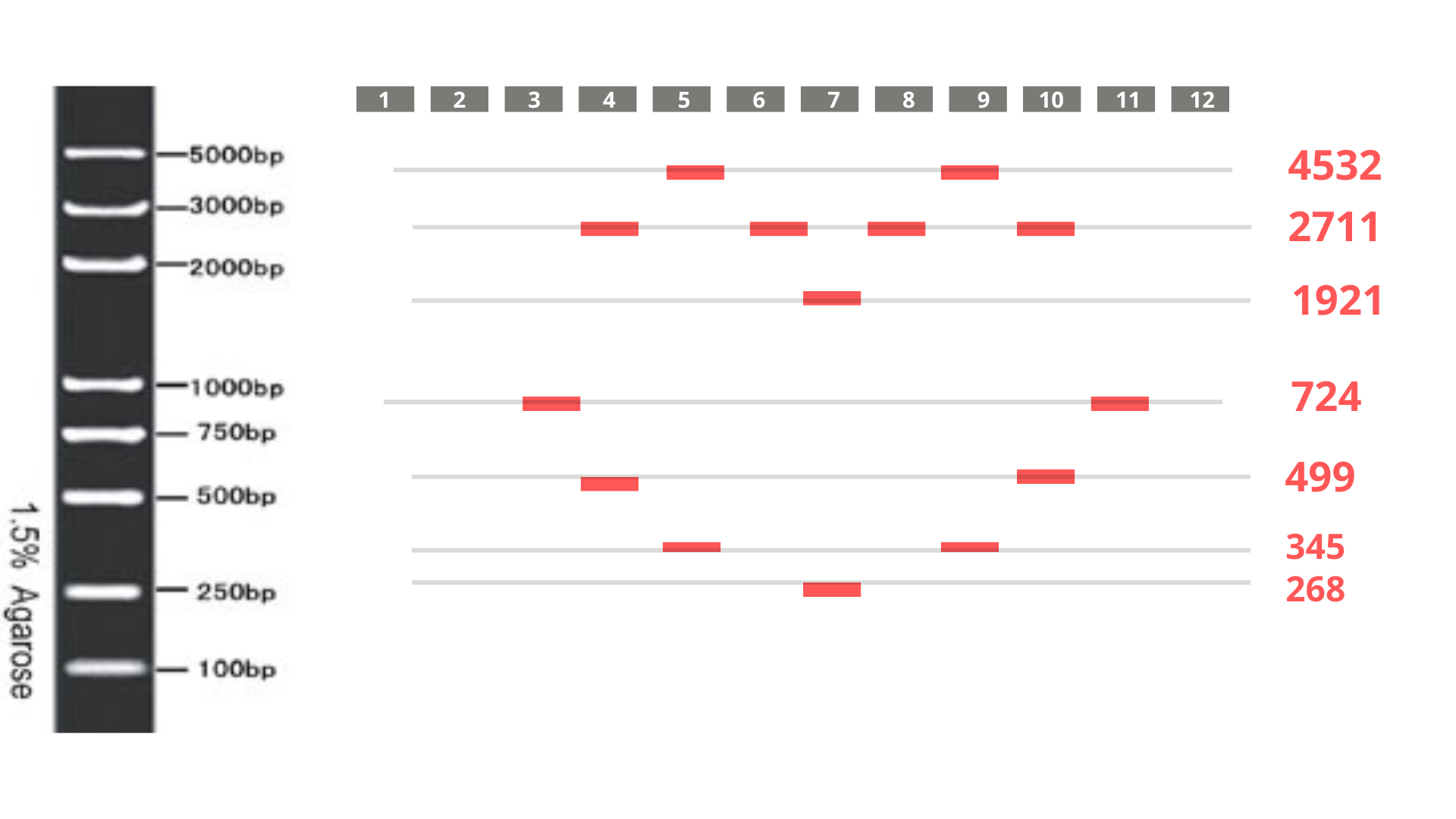

Part 3: DNA Design Challenge
Building on my Week 1 project, I focused on modulating IFN-α levels to reduce poor outcomes in systemic lupus erythematosus (SLE) patients. Current pharmacological approaches for severe SLE also target IFN-α inhibition (Baker et al., 2024). Literature indicates that IFN-α activation occurs through a signaling cascade initiated by Interferon Regulatory Factor 7 (IRF-7). When IRF-7 is no longer needed, the body naturally deactivates it via TRIM28, a SUMO E3 ligase with specific activity against IRF-7 (without affecting the similar IRF-3) (Liang et al., 2011). This mechanism offers a more targeted, less invasive therapeutic approach for SLE.
I retrieved the TRIM28 amino acid sequence from NCBI Protein using the search terms “TRIM28” and “Homo sapiens.” From the results, I selected the third hit (the first two were fragments), obtaining the complete protein sequence in FASTA format: “>AAH52986.1 TRIM28 protein (Homo sapiens) MAASAAAASAAAASAASGSPGPGEGSAGGEKRSTAPSAAASASASAAASSPAGGGAEALELLEHCGVCRERLRPEREPRLLPCLHSACSACLGPAAPAAANSSGDGGAAGDGTGPAKSRDGERTVYCNVHKHEPLVLFCESCDTLTCRDCQLNAHKDHQYQFLEDAVRNQRKLLASLVKRLGDKHATLQKSTKEVRSSIRQVSDVQKRVQVDVKMAILQIMKELNKRGRVLVNDAQKVTEGQQERLERQHWTMTKIQKHQEHILRFASWALESDNNTALLLSKKLIYFQLHRALKMIVDPVEPHGEMKFQWDLNAWTKSAEAFGKIVAERPGTNSTGPAPMAPPRAPGPLSKQGSGSSQPMEVQEGYGFGSGDDPYSSAEPHVSGVKRSRSGEGEVSGLMRKVPRVSLERLDLDLTADSQPPVFKVFPGSTTEDYNLIVIERGAAAAATGQPGTAPAGTPGAPPLAGMAIVKEEETEAAIGAPPTATEGPETKPVLMALAEGPGAEGPRLASPSGSTSSGLEVVAPEGTSAPGGGPGTLDDSATICRVCQKPGDLVMCNQCEFCFHLDCHLPALQDVPGEEWSCSLCHVLPDLKEEDGSLSLDGADSTGVVAKLSPANQRKCERVLLALFCHEPCRPLHQLATDSTFSLDQPGGTLDLTLIRARLQEKLSPPYSSPQEFAQDVGRMFKQFNKLTEDKADVQSIIGLQRFFETRMNEAFGDTKFSAVLVEPPPMSLPGAGLSSQELSGGPGDGP”
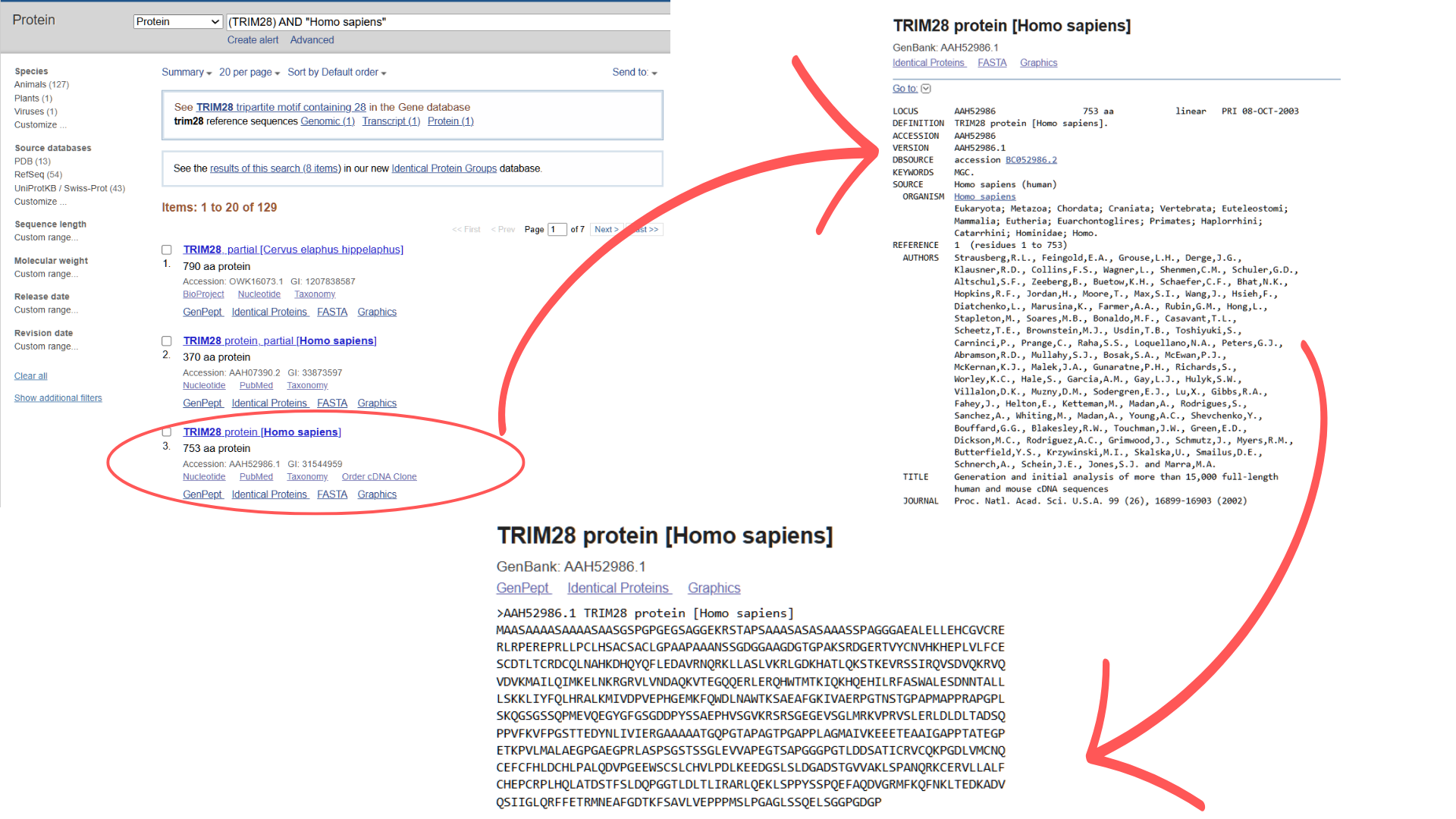

Next, I used NCBI’s “tblastn” BLAST tool to identify the corresponding nucleotide sequence. The top hit (Accession NG_046945.1) showed the highest Query Score (82%) and lowest E-value (3e-164), though the alignment revealed significant gaps between amino acid and nucleotide sequences.
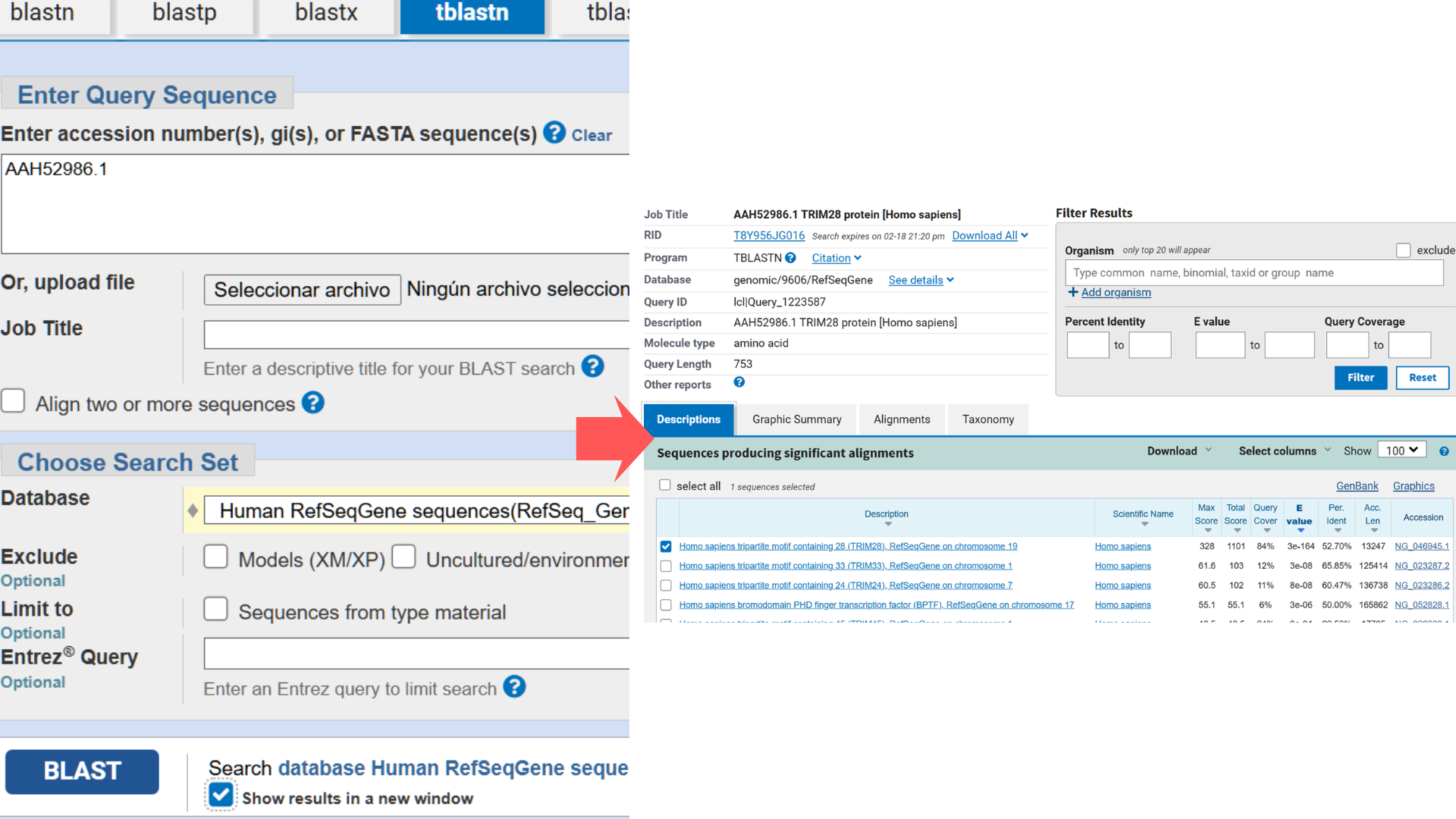

Since multiple codons can encode the same amino acid, nucleotide sequences vary across taxa even when the protein sequence remains identical. Therefore, codon optimization must align with the intended host organism. Translation efficiency depends on tRNA abundance; optimization replaces rare codons with synonymous frequent ones matching the host’s tRNA pool. For my project’s therapeutic goal, I optimized the sequence for human cells.
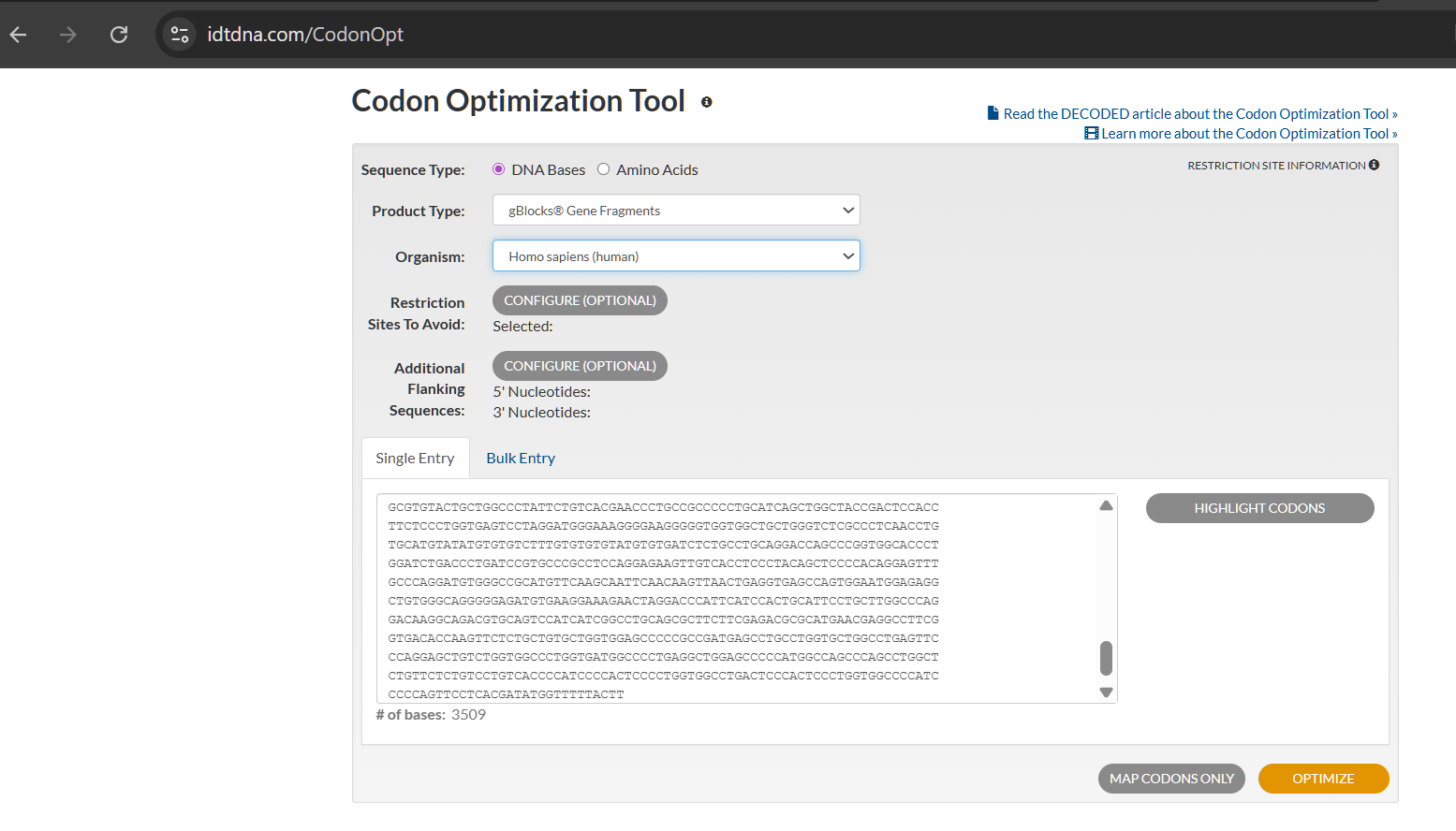

(Note: The optimization tool flagged this sequence as “high complexity,” indicating potential synthesis challenges with commercial methods.)
With the human-optimized sequence ready, the next step is transforming this digital information into a functional protein capable of immune modulation. Two feasible routes exist: cell-dependent or cell-free systems. I could use a cell-free system for initial circuit prototyping (better cost-benefit ratio) and transition to live-cell systems for actual production.
Part 4: Prepare a Twist DNA Synthesis Order
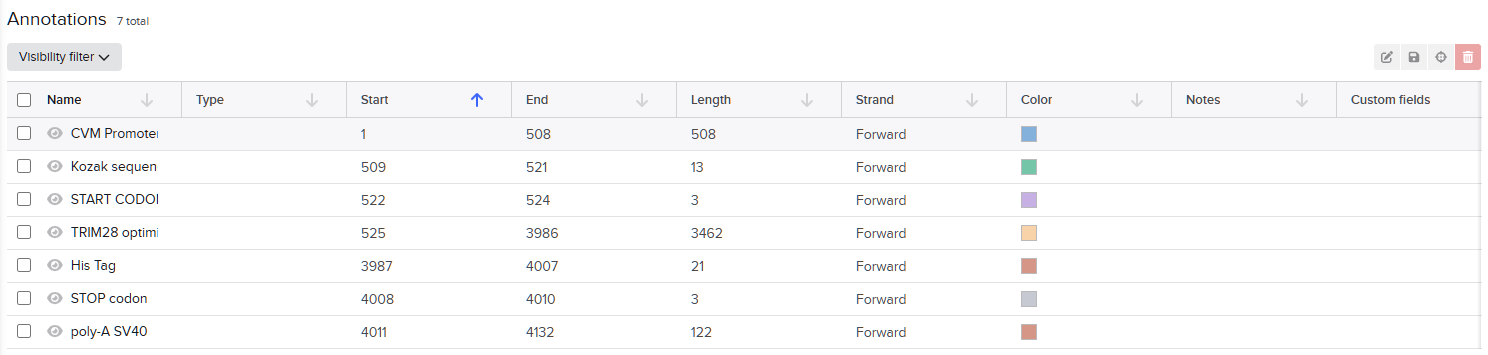
For this exercise, I designed a vector for constitutive TRIM28 expression. In the final project version, I plan to add an IFN-α sensor that inhibits TRIM28 production when concentration exceeds a specific threshold. For now, expressing TRIM28 alone in human cells requires these regulatory elements (McCarty et al., 2023): CMV promoter, Kozak sequence (5’ UTR), Start codon, Optimized TRIM28 coding sequence, His Tag, Stop codon and SV40 polyadenylation signal (3’ UTR).These elements ensure transcriptional and translational efficiency and accuracy.
In Benchling, starting from an empty file:

- I sequentially assembled these components to form the cassette:
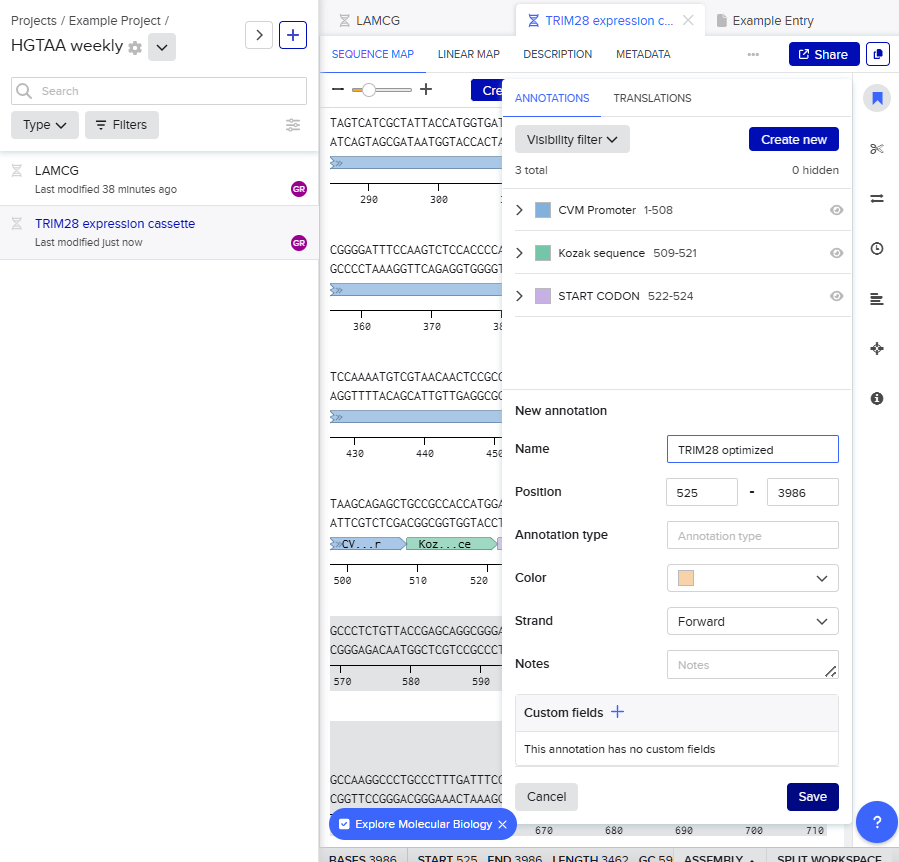

- CMV Promoter:
CGTTACATAACTTACGGTAAATGGCCCGCCTGGCTGACCGCCCAACGACCCCCGCCCATTGACGTCAATAATGACGTATGTTCCCATAGTAACGCCAATAGGGACTTTCCATTGACGTCAATGGGTGGAGTATTTACGGTAAACTGCCCACTTGGCAGTACATCAAGTGTATCATATGCCAAGTACGCCCCCTATTGACGTCAATGACGGTAAATGGCCCGCCTGGCATTATGCCCAGTACATGACCTTATGGGACTTTCCTACTTGGCAGTACATCTACGTATTAGTCATCGCTATTACCATGGTGATGCGGTTTTGGCAGTACATCAATGGGCGTGGATAGCGGTTTGACTCACGGGGATTTCCAAGTCTCCACCCCATTGACGTCAATGGGAGTTTGTTTTGGCACCAAAATCAACGGGACTTTCCAAAATGTCGTAACAACTCCGCCCCATTGACGCAAATGGGCGGTAGGCGTGTACGGTGGGAGGTCTATATAAGCAGAGCT
- Kozak Sequence: GCCGCCACCAUGG
- Start Codon: ATG
- Human-optimized TRIM28 sequence:
TGT GTC CTG CCA TGG CCC CAC CCT CTG CAC TTG CCT AAC AAG TGG CCC TCT GTT ACC GAG CAG GCG GGA AAC GAG GCC CCC ACT CAT TCT GCT AGT TGA GTC GGG GGC GGC GAG GGC CAA GGC CCT GCC CTT TGA TTT CCA CTC CCT AGA GCT TCT CAG GTT TCT GGC TGG TAG ACC TAT TGT CTT CTG CAA AGA ACA CAA GCA TAG ACA CCT TGC GCC GTG CTC TAA GAG CTT TAA TAT AGC CAC CTG CCC AGG CTC CCT GCA CAG TGT CCC CAG GGG CCT CCA GTC TCT CCC AAG GCC TAA TGG CTG GGC GGC TGG GTC CCA CCT CTG CCC TAA CTC AGC GCT CAG GAG CAC ATA TGC CTG CTG AGC GGA ACT TCT AGC TGA AGG ATG CAA TGA GGC ACC AGC GCA TCC TCT TGG CCC CAC TGG TAG TCA GCG CTG GGG ACG AAC ATG CAG CAC TGT CGA CGC GCA CCC AGG CGC TTT GCA GCA CGC AAG TGT GGA TTC TGG GGA TGC GGG GGG GGT CCC GGT CAG CAA GAT CCG ACG TGA CCT GAC CTG CTG TGC CCC CTG GAA AGT GCA AGA TGC CTG ACA TAT AGA TCT GTA TGC AAG TGG ATG TCA CGG TGG CCG TCC TGT CGG TCC TAA CGG TCC TAG ATC AGC GGT GCC GTG TGC TGG TCC ATG ATG CCA CGC TAA GCA TTG TGC AGA TAG GAG GGG TCC CTG TCC CCC CTG TGT TAA CTG ATG ATG CTC AGT GGA GTG CGA AGC GAC CCA AGC ATC CTG CTT CCA AGC CTG CTC ACT TCA TAT ACT TTT TAC CTG CAG AAA GTC ACC GAG GGA CAA CAG GAG CGC CTG GAA AGA CAA CAC TGG ACA ATG ACG AAA ATT CAG AAG CAT CAG GAA CAC ATT CTC CGC TTC GCG TCT TGG GCG CTG GAA AGC GAT AAC AAC ACG GCA CTG TTG CTG AGC AAA AAA CTG GTC TGC ACT GGA GGC TTG CTG GCA GGC GGC TTC CAG GCT GGG GGC AGC CAG TAC CTT AAA AGT CTG TTT GTC GTC GGG GTC CTC ATC TTG TCA TCT CTG AGC CTG CCC ACG AGG TCC ACT AGT AGC TGT ACC GGC CCT AGC CGC TGA CTG TGG ATA CCA TGG AGC CAC ATG GCA AGG TGA AGT TTT TCC GGA ACT TCC ATG CCT GGC CCG AGG GTG CCT AGA CCG CTG GTG GGC CCT CAG CTC TAT TTG ACT CTG TTG CTG CCA CAT GTT CTG CAT CTG CTG ACC TTT CTG TCT AGT TTC TGC CGC CAG GAT TGT GGC CGC GCT AGT TGG CAC TGA CTG AAC AGA CCC TGT ACC CAT GGC CCT TCT AAG AGC CCC AGA GCT CCG GAA CAG GCC GGG CTC TGG CAG CAA CCC GGC GAG CAG GAG CGG GGC CCA CGC AAA GGG TGG GCG GGC AAT GGA GTT CAA TAG CAG GAG AGG GGA CCA CGC CAG GGA GGC TGG GCA GGT AAC GGC GTA CAG TAG GGA AGT TGT CCT ACA GAG GCT GAG GGG AGC GCA CTG TTC ACC GCA CAC GGC GGA GCA GGG AGA CTC TGG CTT TGG GTG CGC TAA GTA GGC CTT CCC TCA GGT TGG GGT AGA GCC CCC GTG TTG TTC CTG CCA CAG CCC GCC AGC CTG CTT TCC CCA TTC TCA AAA GAG ATG ATC CCG ACC CAG GTA CAG TCT CCC ATG TGT CAG GTG TAG AAC GGG AAG TAC GGC ACG TCC CCA GGC GGG GAG GTC GAT GGC GGT TGG GGT TGT ACC CAT GCA AGA CTC CTT CAT TTG CAG GTT CCC CTT CGC TGA GGG CGG GGA GAA CGC CCG TAT GCC CAA GGC GCA ACT TCC GAG CCA TGA ACG CCA GGC CCT GGC CCC CAC TCT TAA CAG CCC GCA ACA AGG CTG CAG GGT CTC CCA AGG CAG TAC CAC TGA GGC CTG CAG CCC TAC TGC TAC TAG ACG TGG CGA TGC CGA TGC TCC TAC CGA CCG GCC CGA GAT TGT GCT TGT AGG AAT CCA TGG TGC CCT ACT CCA GGG TGG CAT GGC CAC TGC CAG GGT AAG CCA GTG CCT CGC AAT TAC AGC TGT CGA ATG AAA CCT GTG GTG CAG GTG TGG ACC TTG CTC AAT ACG CCC CAT CAC CAC CTG GCT GGC GGA GGC GAC GGG GGC TGT CAC TGG TCT CCC TCC TAC TGC CAT TGA GGG CCT TAA GAC CAG ACT TGT GCC TAC GGG AGC TGC GGC GGC TCA TGG TGC TAA GGG TCC CCC CCT GGA CTG ACA TGA TGG CAG CAC CAG CTG AGG GCC GGG GGC GGA GGG TCT TAG GGG TAT CTG AGC CCG CGA TGG TGG CCA GGT AAC CCT GGG TGA CAG TGC CAT CAC CTG CCC TGC CTG CCC GAA GCT CGA CGA TCT GGC TAC GTC CAG CCA GTA TAA GTG CTG TTC CCT CCT GGG CTC TCT CCC GCC GGG CCT GCC GGA TGT ACA CGG GTC AGT GTG CGC TTG GTG GGG GTG AAG AGC GGG TGT TGG GCC GTC TGG ACA GGC TCC TGC GCA GAT CCT TAT TTT CTG CAC CCA AGG GAA GAG TGG TCT TGT AGC CTC TGT CAC GTT TTG CCC GAC CTG AAA GAA GAA GAT GGC TCA CTC TCC CTC GAC GGG GCA GAC TCT ACT GGA GTT GTG GCA AAG CTG TCT CCC GCT AAC CAA AGG GTC CGC GCT GGG GTC ACC TGA GTG GGT CTC CCT CGG GAG GCC CTT TAG GTG CTG CCT CGG GCG GTG ACG AGT TTG ACA ATG TTC GTG GAA ATG TAG GCC TGC ACG GCT GGT CCT ATC TTG TCC CGG ACG CTG CCT CCT CCC GCC TCT GCA GGA TAT AGA CTG CAC CTT TTG CCC GGC GAG AGT TAG GAC GGA AAA GGC GAA GGC GGC GGC GGC TGT TGG GTG TCT CCC AGC ACT TGC GCC TGT ATA TGT GTA TCA CTG TGC GTG TAC GTA TGA AGC CTG CCA GCT GGA CCT GCC CGG TGG CAC CCC GGC TCT GAT CCT GAC CCA TGT CCT CCA CCG GGC GAA GTG GTA ACC TCC CTT CAG CTG CCA ACT GGC GTT TGT CCA GGT TGT GGC CCA CAT GTG CAA GCA ATT CAA CAG GTG AAC TGA GGC GAA CCC GTG GAG TGG AGG GGA TGC GGC CAG GGA GAA ATG TAA AGG AAA GAA TTG GGG CCC ATC CAT CCA CTG CAT AGC TGC CTG GCC CAG GAC AAA GCT GAT GTG CAG AGC ATC ATT GGA CTG CAG CGC TTC TTC GAG ACT AGA ATG AAT GAG GCT TTC GGT GAC ACG AAG TTC TCT GCC GTG CTG GTG GAG CCG CCC CCT ATG TCT TTG CCA GGA GCG GGA CTT TCA AGT CAG GAA CTC TCC GGC GGT CCT GGG GAT GGC CCA TAA GGC TGG AGT CCA CAT GGC CAG CCG TCA CTC GCC CTT TTC TCT GTT CTG AGC CCA CAT CCC CAC AGC CCC GGC GGC TTG ACG CCT ACG CCT TGG TGG CCA CAT CCT CCT GTT CCT CAC GAC ATG GTT TTC ACT TTT
- His Tag: CATCACCATCACCATCATCAC
- Stop Codon: TGA
- SV40 Poly-A Signal:
TAAGATACATTGATGAGTTTGGACAAACCACAACTAGAATGCAGTGAAAAAAATGCTTTATTTGTGAAATTTGTGATGCTATTGCTTTATTTGTAACCATTATAAGCTGCAATAAACAAGTT
I exported the cassette as a FASTA file and uploaded it to Twist.
When deciding between Clonal Genes and Gene Fragments, the determining factors were: my cassette’s small size (even accounting for future sensor addition) and the practicality of working with a plasmid vector. For the backbone, I selected pTwist CMV OriP, which ensures high-level transient expression in mammalian cells, provides ampicillin resistance, and facilitates elevated protein expression.
Upon uploading, the platform flagged multiple repetitive sequence regions—consistent with the “high complexity” warning from the optimization tool. TRIM28 is an enzyme with functional domains; repetitive regions are expected. However, I’m concerned that my edits to bypass synthesis constraints may have compromised these functional domains.
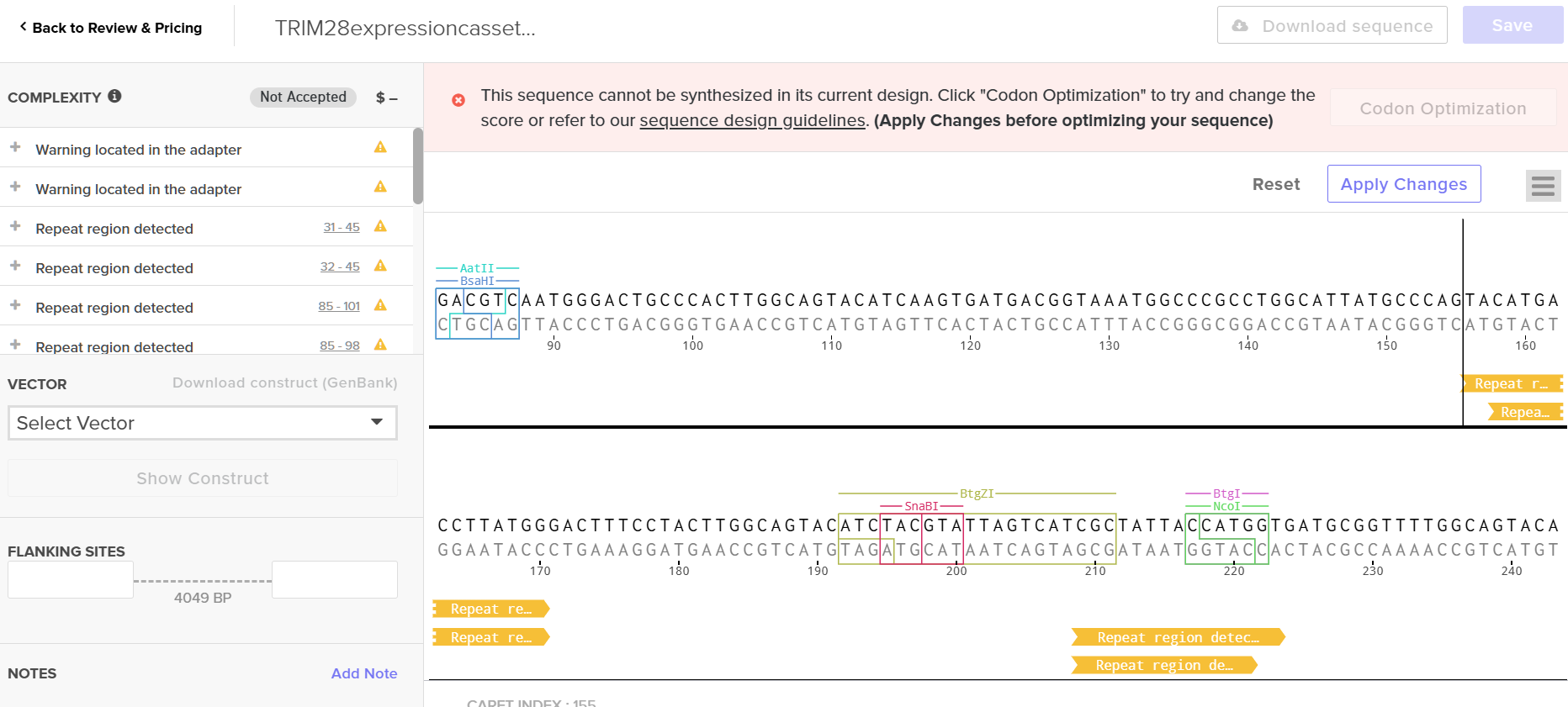

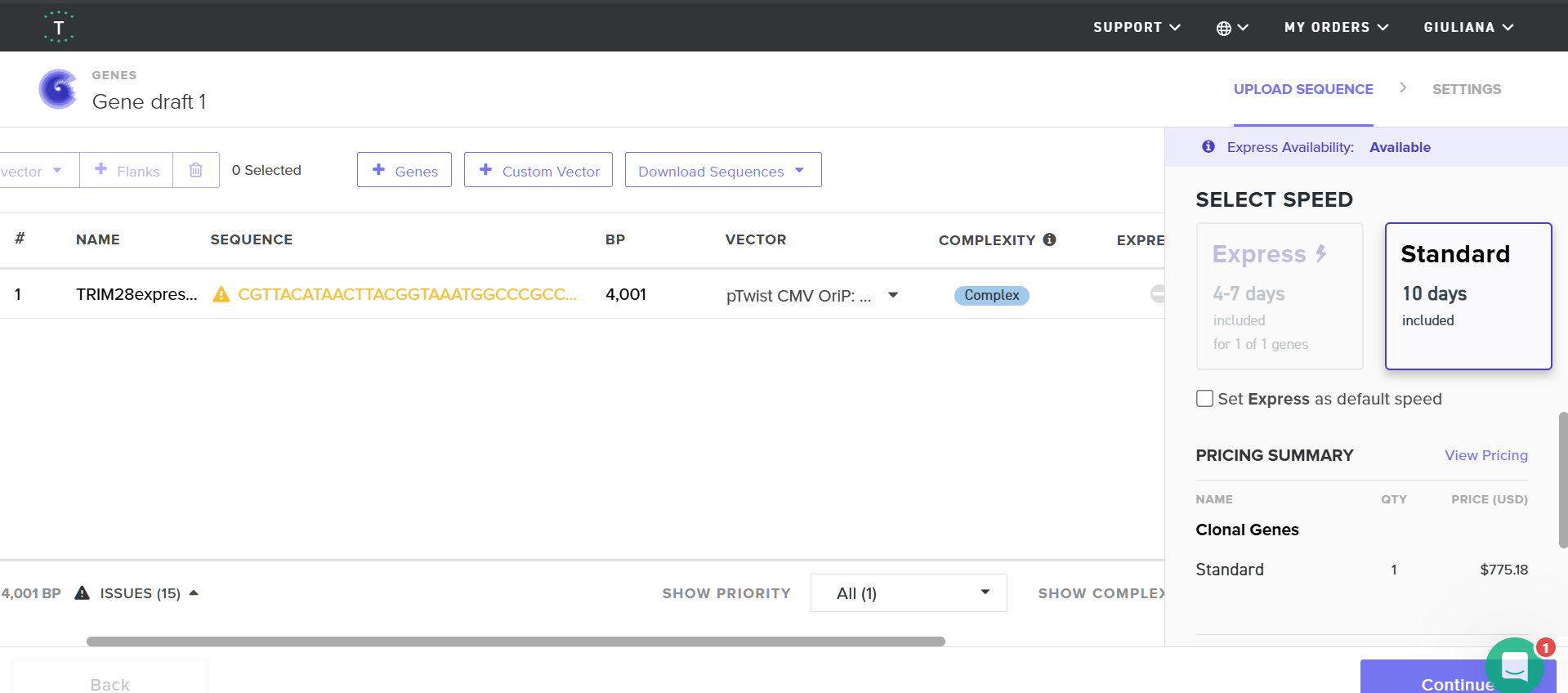

Anyway, I downloaded the resulting GenBank file and re-imported it into Benchling.

- The final plasmid/vector is:
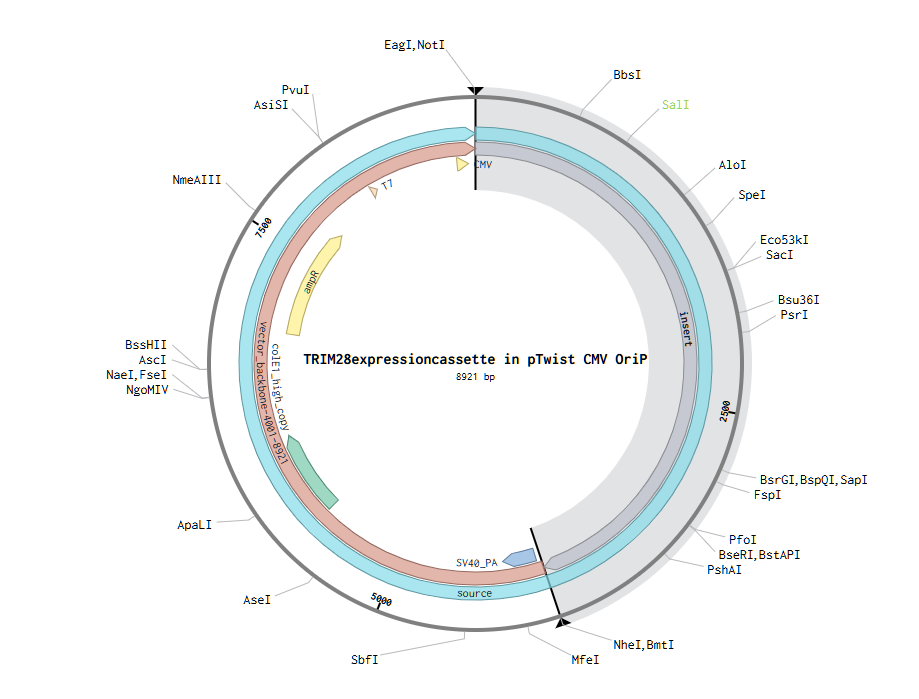

Part 5: DNA Read/Write/Edit
- 5.1 DNA Read: (i) What DNA would you want to sequence and why? In my proposed lupus autoregulatory circuit project, I would sequence cell-free DNA (cfDNA) from patient plasma to detect interferon-stimulated gene (ISG) signatures that predict disease flares. cfDNA sequencing offers a minimally invasive way to aquire information aboud patients’ in-real-time situation. Elevated IFN-α family actitvity would trigger preemptive TRIM28 circuit activation before clinical symptoms manifest—transforming lupus management from reactive to predictive. (ii) Sequencing technology choice In my expereience Ecuador, Illumina NovaSeq is the to go technology for biomedical research (second-generation sequencing). It is second-generation sequencing (sequencing-by-synthesis with reversible terminators), and works fine when needing to balances accuracy (Q30+ scores) with scalability—critical (e.g. when screening hundreds of rural patients). For the input (taken from the “QIAseq cfDNA All-in-One Kit Handbook”): Extract 10 ng cfDNA from 1 mL plasma (using silica-column kits compatible with low-input Twist workflows) Fragment to 300 bp via enzymatic shearing (avoiding sonication that degrades already-short cfDNA) Ligate Twist Universal Adapters with unique dual indexes (UDIs) to enable multiplexing of 96 samples per lane PCR amplify with 8 cycles (minimizing chimera formation noted in LeProust’s slides, p. 38) Base calling mechanism: Fluorescently labeled nucleotides (A/C/G/T with distinct dyes) are incorporated sequentially. After each cycle, lasers excite the dyes and cameras capture emission wavelengths. Software converts fluorescence patterns into base calls (A=green, C=blue, G=yellow, T=red), with quality scores reflecting signal-to-noise ratios. Output: 150 bp paired-end reads (300 bp total coverage per fragment) in FASTQ format. 5.2 DNA Write (i) What DNA would you want to synthesize and why? I would synthesize the TRIM28 expression cassette I designed in Part 3—a 1.2 kb sequence containing CMV promoter, Kozak sequence, human-optimized TRIM28 coding sequence, His tag, and SV40 poly-A signal. This cassette represents the effector module of my lupus circuit: when IFN-α levels rise during pre-flare states, TRIM28 would be expressed to SUMOylate and deactivate IRF-7 (Liang et al., 2011), breaking the pathogenic IFN-α amplification loop without broad immunosuppression. Unlike current therapies (e.g., anifrolumab antibody injections requiring monthly clinic visits), this circuit could provide continuous, autonomous regulation—critical for patients in Chimborazo province who travel 6+ hours for care. (ii) Synthesis technology choice I would use Twist Bioscience’s silicon-based phosphoramidite synthesis.
- In silico design: Finalize cassette in Benchling with codon optimization for human cells (avoiding rare codons like AGG that stall ribosomes)
- Chip synthesis: 150-mer oligos synthesized in parallel on silicon wafer (1 million oligos/chip)
- Enzymatic assembly: i would opt to join independtly synthesized oligos into a full-length cassette.
- Error correction: there must be a MutS protein to bind mismatched bases for removal
- As stablished, I choosed cloning as to make it easier for the vector (pTwist CMV OriP) to reach mammalian (host cells) expression. A limitation encountered was that during my simulation (Part 4), Twist flagged my TRIM28 sequence as “high complexity” due to repetitive domains. This reflects a fundamental constraint: phosphoramidite chemistry has ~99% coupling efficiency per cycle (LeProust slides), making synthesis of long repeats error-prone. For my final project, I’d need to: -Break repeats with synonymous codon substitutions (preserving amino acid sequence) -Use Twist’s Ultra-complex Genes service (which is costly!!) for sequences with homopolymers >20 bp Accept that some functional domains may require post-synthesis assembly via Golden Gate cloning 5.3 DNA Edit (i) What DNA would you want to edit and why? My project is not about directly editing the patients DNA, as I am very concerned about incidentally damaging the patient. What I aim to do is to provide an external aid that is not as invasive as the current pharmacological guidelines. If I would given to choose what DNA to directly edit, I would be interested in modifying plants to become natural biorreactors: instead of producing their wildtype fruits (e.g. compounds required for the development of the seed), they could be edited to produce an specific compound (such as insulin for example).
References Baker T, Sharifian H, Newcombe PJ, et al. Type I interferon blockade with anifrolumab in patients with systemic lupus erythematosus modulates key immunopathological pathways in a gene expression and proteomic analysis of two phase 3 trials. Annals of the Rheumatic Diseases. 2024;83:1018-1027. Liang Q, Qi H, Hou F, et al. TRIM28 is a SUMO E3 ligase for IRF7. Journal of Biological Chemistry. 2011;286(37):32448-32456. McCarty NS, Graham AE, Studena L, et al. Rapid and scalable characterization of CRISPR technologies using an E. coli cell-free transcription-translation system. Molecular Cell. 2023;83(1):1-15.