Week 3— Lab Automation
Python Script for Opentrons Artwork
After making a copy of the collab doc into my drive, I searched for a reference image for my design. In the Designer Cells node there are two color options: red and green. So, I looked for a minimalistic design that could look good in green, my favorite color. I ended up choosing a hummingbird, as they come oftentimes to my garden to feed on the flowers’ nectar.
At the Automation Art Interface, there is the function of submitting an online image directly coping its direction, and it is suggested that its backdrop is white. The following is the selected reference image that meets the previous criteria:
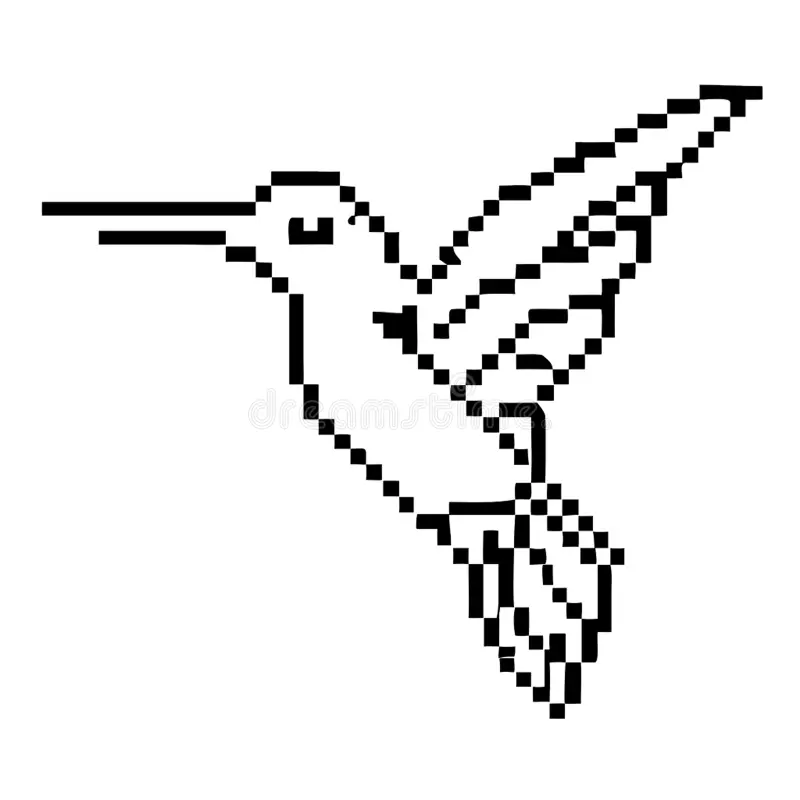
(This is the image’s web direction/link pasted in the Automation Art Interface)
With it i obtained this incredibly beautiful and complex design:
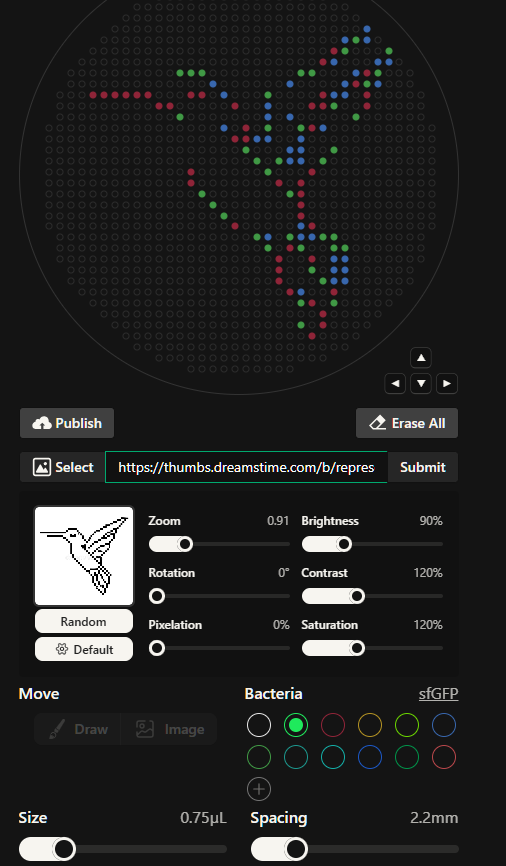
I tinkered manually with the dots and with the default settings (Brightness, Contrast and Saturation) to modify the original suggested design, plus I changed all the dots to the “mClover3” color as I initially intended. After that, the coordinates I obtained were:
mclover3_points = [(23.1,27.5), (18.7,25.3), (20.9,25.3), (29.7,25.3), (16.5,23.1), (27.5,23.1), (-12.1,20.9), (-5.5,20.9), (12.1,20.9), (23.1,20.9), (-34.1,18.7), (-31.9,18.7), (-29.7,18.7), (-27.5,18.7), (-25.3,18.7), (-23.1,18.7), (-20.9,18.7), (-18.7,18.7), (-16.5,18.7), (-14.3,18.7), (9.9,18.7), (18.7,18.7), (20.9,18.7), (-3.3,16.5), (7.7,16.5), (25.3,16.5), (-16.5,14.3), (-14.3,14.3), (-1.1,14.3), (5.5,14.3), (14.3,14.3), (16.5,14.3), (25.3,14.3), (-12.1,12.1), (1.1,12.1), (12.1,12.1), (23.1,12.1), (1.1,9.9), (14.3,9.9), (20.9,9.9), (-1.1,7.7), (1.1,7.7), (20.9,7.7), (1.1,5.5), (18.7,5.5), (-9.9,1.1), (14.3,1.1), (-9.9,-1.1), (7.7,-1.1), (12.1,-1.1), (-7.7,-3.3), (9.9,-3.3), (-3.3,-7.7), (-1.1,-9.9), (12.1,-9.9), (14.3,-9.9), (14.3,-12.1), (18.7,-12.1), (9.9,-14.3), (18.7,-14.3), (7.7,-16.5), (14.3,-16.5), (20.9,-16.5), (7.7,-18.7), (20.9,-18.7), (7.7,-20.9), (16.5,-20.9), (20.9,-20.9), (9.9,-23.1), (18.7,-23.1), (20.9,-23.1), (14.3,-25.3), (18.7,-25.3), (16.5,-27.5), (16.5,-29.7)]
Returning to the Google collab document, scrolling to the last celd, below the Your code comment, I copy pasted the tuple containing the coordinates. Below the pasted tuple, on the same level of indentatio, I defined the well for retrieving the green color based on the posted examples. Then, to dispense the “colorant” into the precise coordinate it was needed, I started with the following lines of code:
I selected Run all and after the run, an error message popped up: I was calling pipette_20ul.pick_up_tip(), which didn’t exist. The run stopped there, and I accepted the help of the Gemini assistant to understand where the mistake lay:
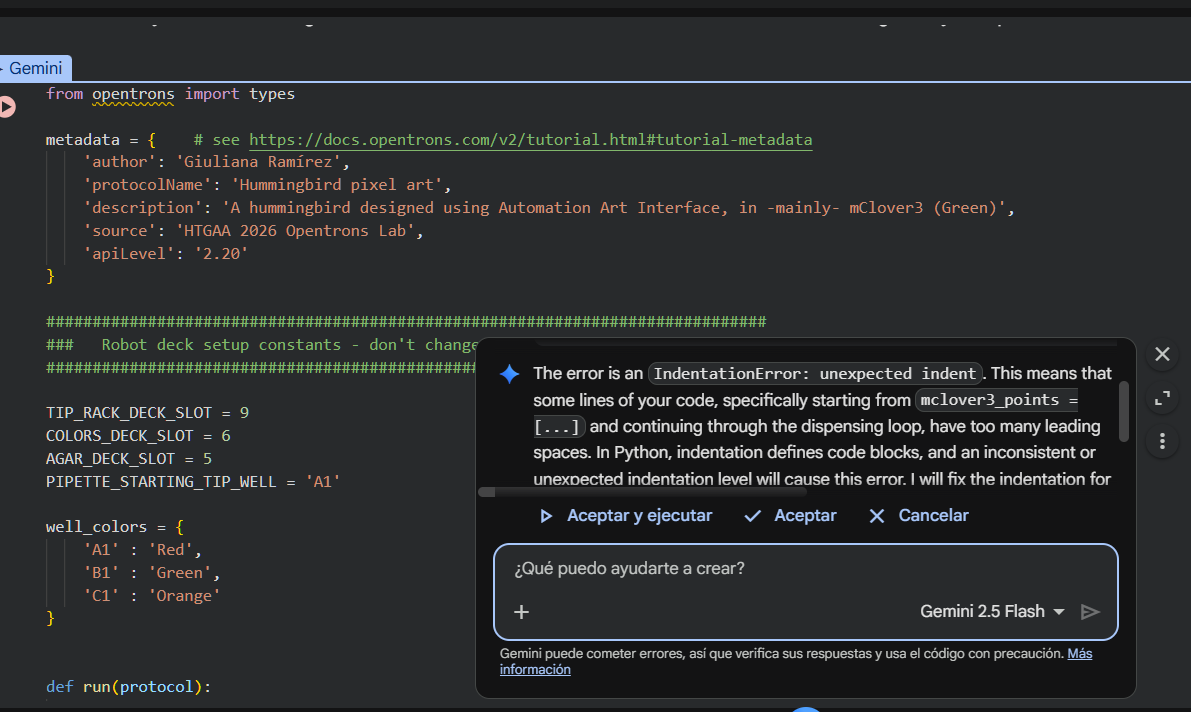

Gemini observed that I had an indentation error when pasting the tuple with my coordinates, due which I had called for pipette_20ul.pick_up_tip() outside of def run(protocol). From that point on, I got the indentation for my code wrong.
I corrected this by using tab at the start of every code line, and clicked on Run all again. However, for the second time an error message popped up:
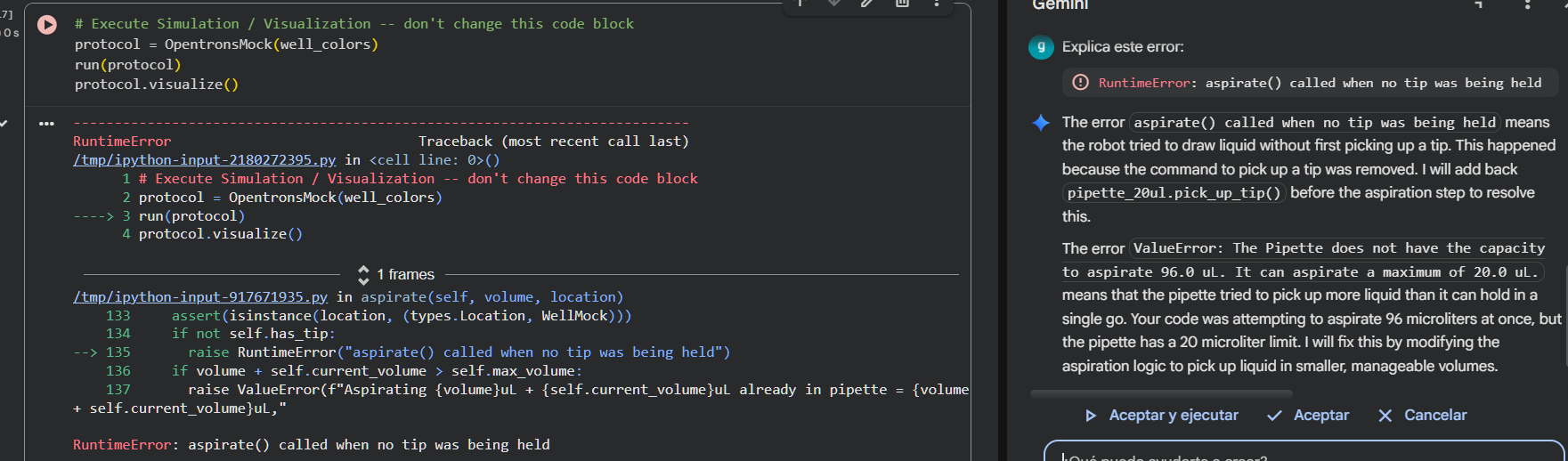

Gemini summed up the mistake in that I had coded pipette_20ul.aspirate(len(mclover3_points) * 1.2, location_of_color('Green')) forgetting that the robot was using 20 ul tips. With the current line of code, I was taking 90 ul at the start of the task, which was physically impossible.
Gemini suggested the next edit to correct the mistake while guaranteeing that the robot would not run out of “colorant”:
I hit the Run all option again and this time the code ran smoothly, and I could visualize my final design:
How AI aided me?: I am familiar with Python coding (but not with the Opentron API libraries). Gemini helped me to understand the logic/algorithm behind the coding. And this AI also became proofreader, noticing details that I missed but that directly affected my run.
Post-Lab
- Finding a published paper that utilizes Opentrons or an automation tool to achieve novel biological applications:
I found a published review paper: Automation and artificial intelligence in filamentous fungi-based bioprocesses: A review, that showcases how continuous data acquisition through intelligent sensors combined with Machine Learning (A.I) and automatization tools and protocols could boost fungi-based bioprocesses.
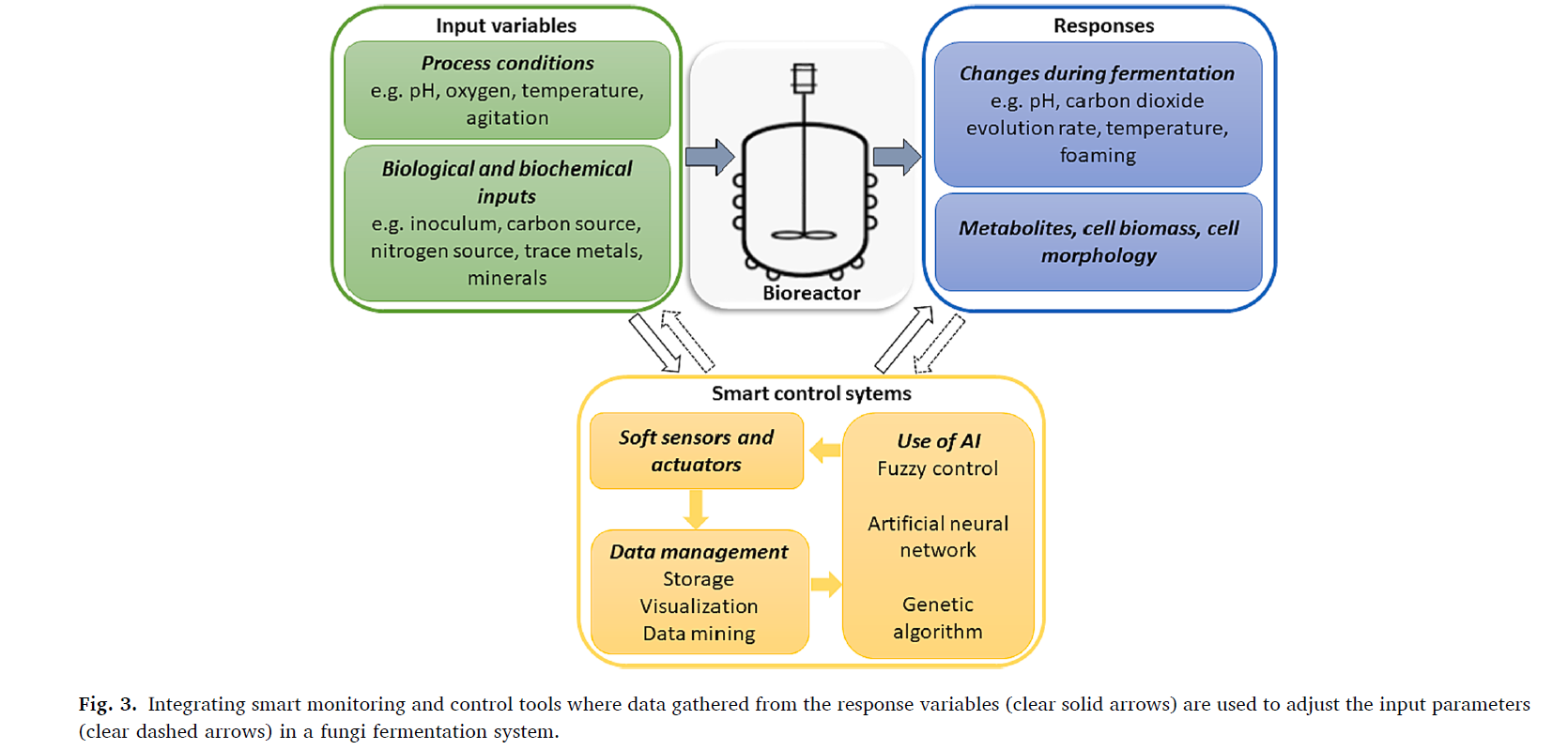

Taken from the paper by (Wainaina & Taherzadeh, 2023)
On their subchapter 3.4, they present the concept of the smart bioreactor in the context of filamentous fungi. The authors propose that such bioreactors should integrate the existing infrastructure of a bioreactor with the concept of a hierarchical structure control system (HSCS). HSCS is a “set of devices and software organized in a tiered manner” where “the layers on the higher levels are designed to perceive theoretical models and execute planning while the lower ones perform local tasks…”(Wainaina & Taherzadeh, 2023). The authors explain that automated sensors could help detect fermentation rate and related parameters, so researchers could monitor the bioreactor progress and make adjustments in real-time. This grabbed my attention, being the first time that I associated automation with bioreactors. HSCS prototyping could be a door to upscaling bioreactor production on low funded startups.
- Intended automation tools usage in my final project:
During Week 2, when I designed my TRIM28 expression cassette in Twist, the platform flagged my sequence as “high complexity” due to repetitive sequence parts (which I presume are part of the RBCC region’s domains). I have heard before that this is a common challenge when working with natural enzyme sequences that contain tandem repeats. This technical barrier directly impacts my goal of developing an autoregulatory circuit for lupus patients with a poor initial prognosis. I had to manually delete several high repetition regions for Twist being able to accept my order, and I wondered how this change could alter the original functionality of the enzyme. I also started to question the opposite: how much I could simplify the sequence and have the resulting protein performing its original function. A further simplified sequence would be cheaper and faster to sequence, optimizing at least one step of the process needed to reach my project´s goal.
So, I thought that I can use Opentron to make a trial in this regard:
I could order 3 to -5 variants of my TRIM28 construct, each with more high repetition sites manually deleted. However, to each construct I would add a ubiquitination sensor (Qin et al., 2022; Choi et al., 2019).
In the presence of TRIM28 and IRF7 ubiquitination should occur as part of IRF7 degradation process. I could gauge which construct is working by measuring the ubiquitination rate. Ubiquitination occurs naturally in the cells, so a control group of cells (without the construct expressing continuously TRIM28) should be added to the well, to have a contrast that can showcase the “extra” ubiquitination activity in the transfected cells. Literature suggests that after a cell culture is transfected, analysis should start after 24- 48 hours (only then cells could be harvested) (Qin et al., 2022).
I would use the Opentron robot to automate the process of adding each different construct to a well in a well plate containing mammalian cell lines. After robot preparation, cells would be manually transferred to a standard CO₂ incubator — a hybrid workflow that reduces human error in small-volume transfers. After the incubation period, the well plate would return to the Opentron robot to trypsinize cells for 2 purposes: 1) make a subculture; 2) Perform a fluorescence test.
- The cell subcultures of each cell group would be used to perform a western blot to determine the presence and relative abundance of TRIM28. The Opentron website mentions that the western blotting process could be automated , so this part could be left to the robot to increase throughput.
- There is also a link in the Opentron website about Fluorescence-Activated Cell Sorting (FACS) Sample Prep , however, in this protocol the robot adds staining fluorescent antibodies, which is not needed or wanted for my current design. If this step could be omitted, and some other steps edited to enhance coherence within steps, then the fluorescence assay could be automated too. If not, then after cell trypsinization the cell culture remaining in the well plate could be manually taken to the fluorescence microscopy equipment.
References:
- Choi, YS., Bollinger, S.A., Prada, L.F. et al. High-affinity free ubiquitin sensors for quantifying ubiquitin homeostasis and deubiquitination. Nat Methods 16, 771–777 (2019). https://doi.org/10.1038/s41592-019-0469-9
- Qin, W., Steinek, C., Kolobynina, K., Forné, I., Imhof, A., Cardoso, M. C., & Leonhardt, H. (2022). Probing protein ubiquitination in live cells. Nucleic acids research, 50(21), e125. https://doi.org/10.1093/nar/gkac805
- Wainaina, S., & Taherzadeh, M. J. (2023). Automation and artificial intelligence in filamentous fungi-based bioprocesses: A review. Bioresource Technology, 369(128421), 128421. https://doi.org/10.1016/j.biortech.2022.128421