Week 2 HW: DNA Read, Write, and Edit
Part 1: Benchling and In-silico Gel Art
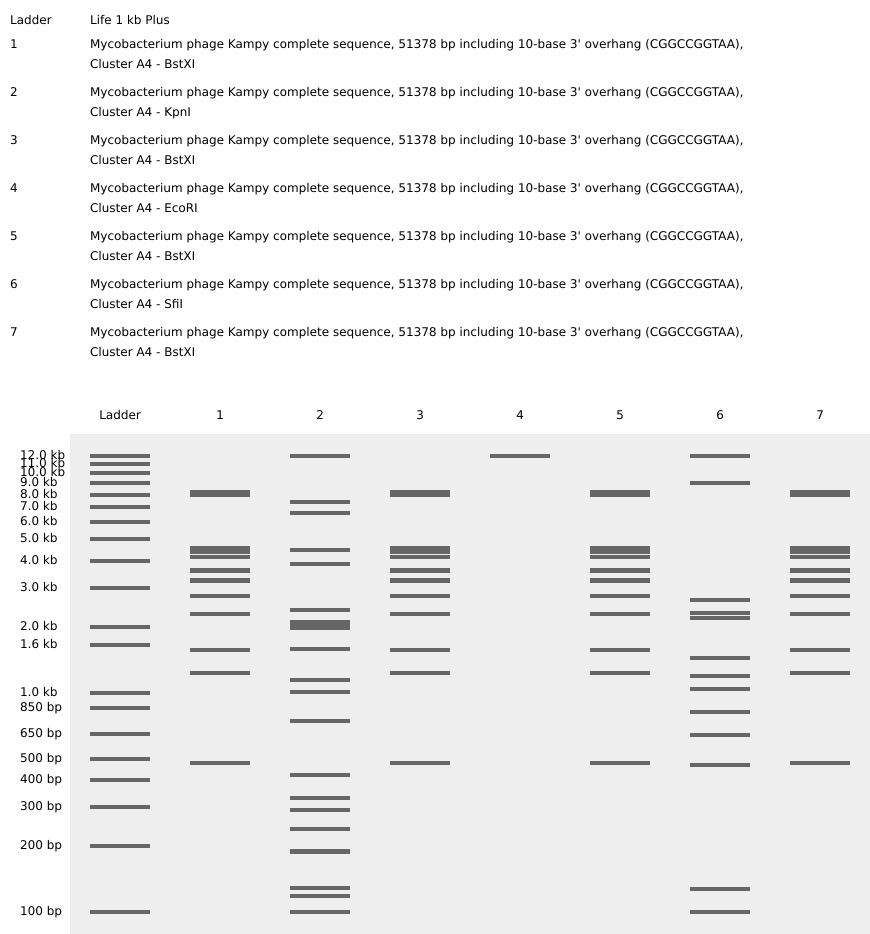


Virtual restriction enzyme digest designed with DNA from the bacteriophage Kampy (isolated at W&M!) and the restriction enzymes BstXI, KpnI, and SfiI to resemble two bacteriophages. The chosen restriction enzymes were selected because they were in stock at William & Mary, had multiple cut sites in the Kampy DNA, and could be combined to make a design resembling a bacteriophage.
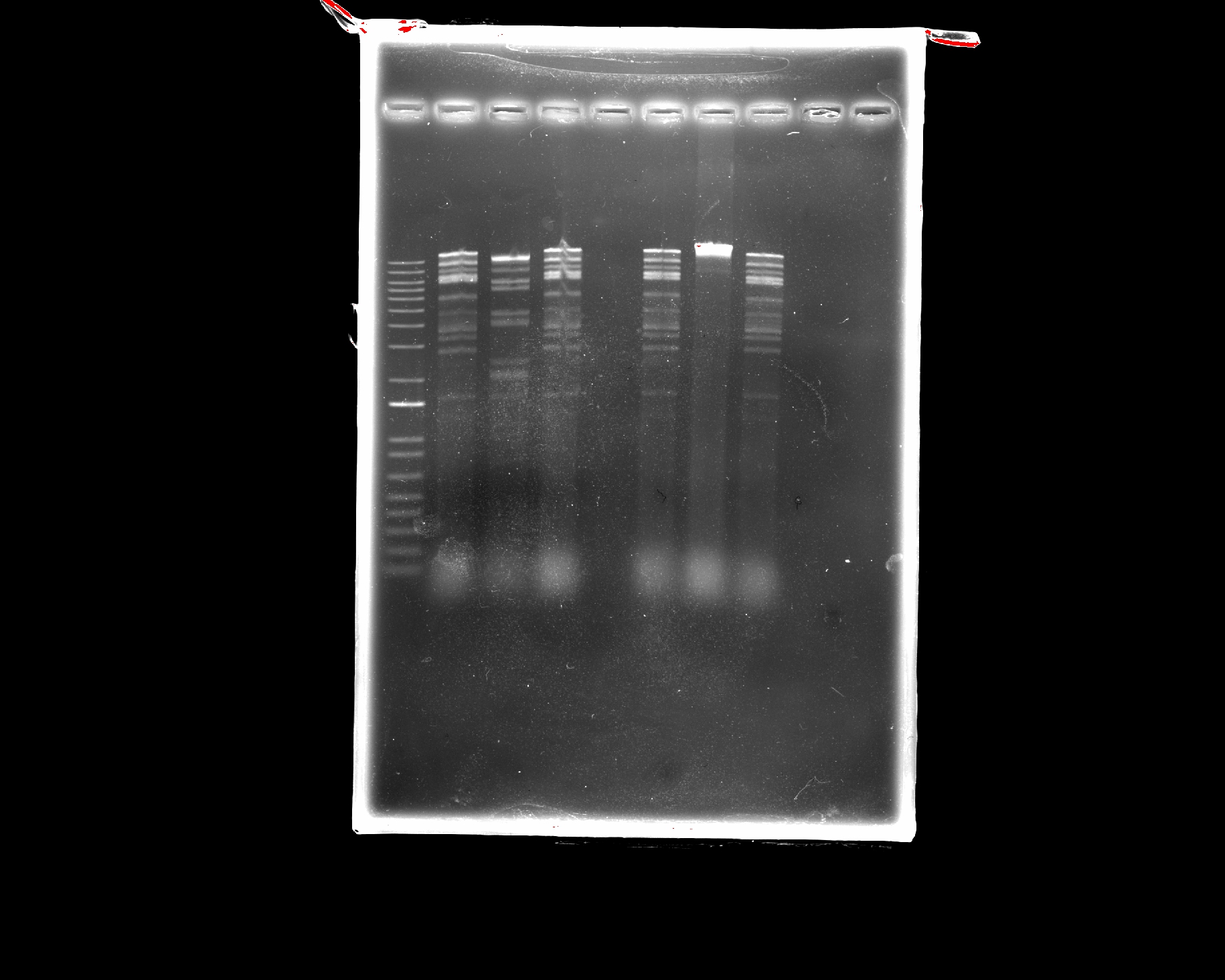
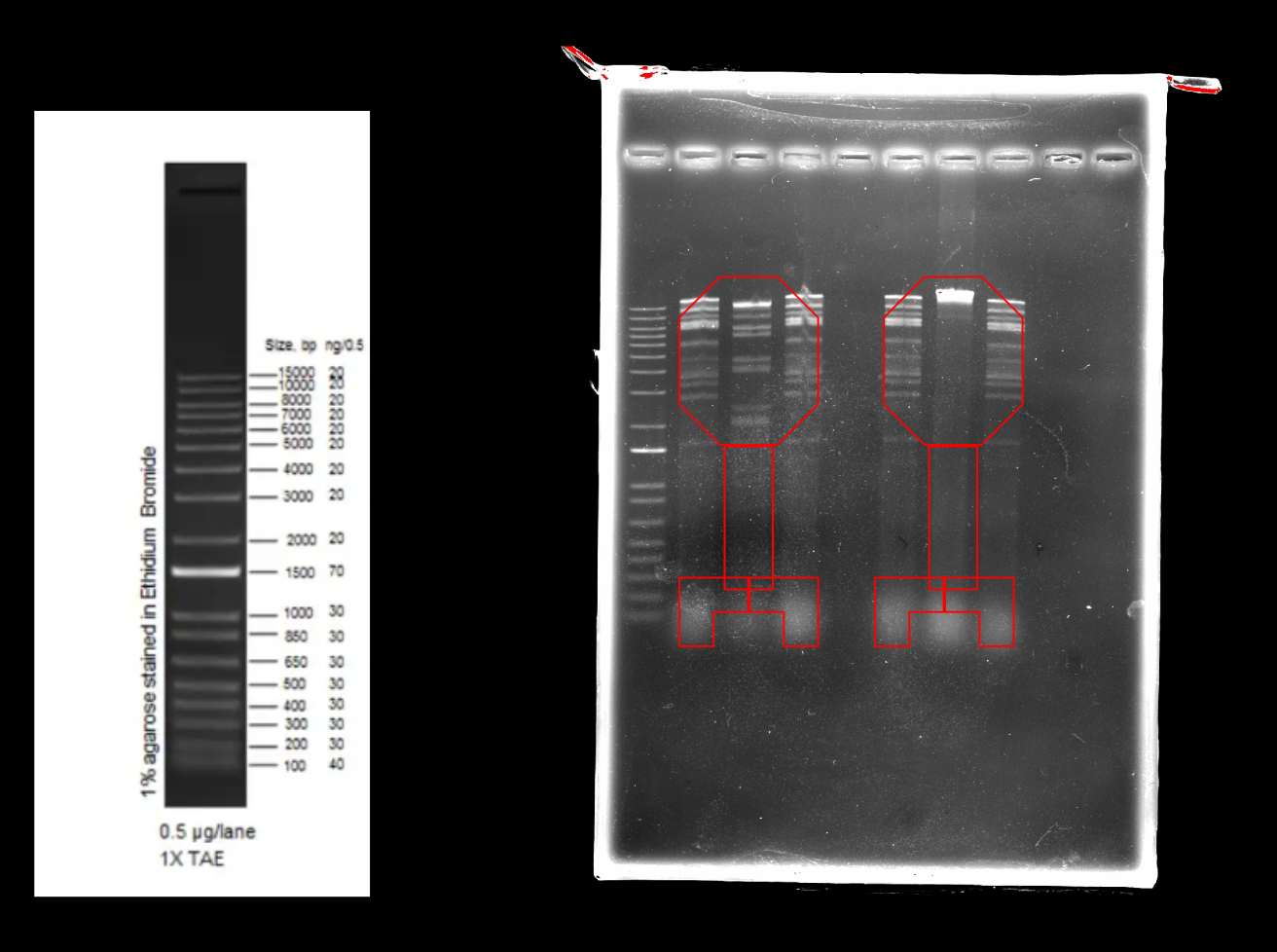
Part 2: Restriction Digests and Gel Electrophoresis
The imaged gel resembles the band lengths predicted by the virtual digest! While interpreting the gel image as two bacteriophages does require a bit of imagination, the digest was successful in that the true gel resembled the in silico prediction.
Part 3: DNA Design Challenge
3.1. Choose Your Protein
I chose the protein TTYH3 (tweety homolog 3) because it is the subject of my honors thesis research. ttyh3 encodes a calcium-dependent chloride channel and is a member of the tweety gene family- consisting of members ttyh1, ttyh2, and ttyh3- that is highly conserved across eukaryotes. ttyh3 is the least-characterized member of the tweety gene family, making it an intriguing subject of research. During neural development, the gene ttyh3 is primarily expressed in post-mitotic neurons. However, its precise function and role in neural development remain unknown. In my research, I aim to provide greater insight into the role of TTYH3 in neural development by overexpressing and knocking-out the ttyh3 gene in X. laevis and observing changes in expression of the downstream neural marker genes Sox2 and tubb2b.
Sequence from UniProt: MAGVSYAAPWWVSLLHRLPHFDLSWEATSSQFRPEDTDYQQALLLLGAAALACLALDLLFLLFYSFWLCCRRRKSEEHLDADCCCTAWCVIIATLVCSAGIAVGFYGNGETSDGIHRATYSLRHANRTVAGVQDRVWDTAVGLNHTAEPSLQTLERQLAGRPEPLRAVQRLQGLLETLLGYTAAIPFWRNTAVSLEVLAEQVDLYDWYRWLGYLGLLLLDVIICLLVLVGLIRSSKGILVGVCLLGVLALVISWGALGLELAVSVGSSDFCVDPDAYVTKMVEEYSVLSGDILQYYLACSPRAANPFQQKLSGSHKALVEMQDVVAELLRTVPWEQPATKDPLLRVQEVLNGTEVNLQHLTALVDCRSLHLDYVQALTGFCYDGVEGLIYLALFSFVTALMFSSIVCSVPHTWQQKRGPDEDGEEEAAPGPRQAHDSLYRVHMPSLYSCGSSYGSETSIPAAAHTVSNAPVTEYMSQNANFQNPRCENTPLIGRESPPPSYTSSMRAKYLATSQPRPDSSGSH
3.2. Reverse Translate: Protein Sequence to DNA Sequence
Using the “reverse translate” tool on bioinformatics.org, I generated the following nucleotide sequence from the TTYH3 amino acid sequence: atggcgggcgtgagctatgcggcgccgtggtgggtgagcctgctgcatcgcctgccgcattttgatctgagctgggaagcgaccagcagccagtttcgcccggaagataccgattatcagcaggcgctgctgctgctgggcgcggcggcgctggcgtgcctggcgctggatctgctgtttctgctgttttatagcttttggctgtgctgccgccgccgcaaaagcgaagaacatctggatgcggattgctgctgcaccgcgtggtgcgtgattattgcgaccctggtgtgcagcgcgggcattgcggtgggcttttatggcaacggcgaaaccagcgatggcattcatcgcgcgacctatagcctgcgccatgcgaaccgcaccgtggcgggcgtgcaggatcgcgtgtgggataccgcggtgggcctgaaccataccgcggaaccgagcctgcagaccctggaacgccagctggcgggccgcccggaaccgctgcgcgcggtgcagcgcctgcagggcctgctggaaaccctgctgggctataccgcggcgattccgttttggcgcaacaccgcggtgagcctggaagtgctggcggaacaggtggatctgtatgattggtatcgctggctgggctatctgggcctgctgctgctggatgtgattatttgcctgctggtgctggtgggcctgattcgcagcagcaaaggcattctggtgggcgtgtgcctgctgggcgtgctggcgctggtgattagctggggcgcgctgggcctggaactggcggtgagcgtgggcagcagcgatttttgcgtggatccggatgcgtatgtgaccaaaatggtggaagaatatagcgtgctgagcggcgatattctgcagtattatctggcgtgcagcccgcgcgcggcgaacccgtttcagcagaaactgagcggcagccataaagcgctggtggaaatgcaggatgtggtggcggaactgctgcgcaccgtgccgtgggaacagccggcgaccaaagatccgctgctgcgcgtgcaggaagtgctgaacggcaccgaagtgaacctgcagcatctgaccgcgctggtggattgccgcagcctgcatctggattatgtgcaggcgctgaccggcttttgctatgatggcgtggaaggcctgatttatctggcgctgtttagctttgtgaccgcgctgatgtttagcagcattgtgtgcagcgtgccgcatacctggcagcagaaacgcggcccggatgaagatggcgaagaagaagcggcgccgggcccgcgccaggcgcatgatagcctgtatcgcgtgcatatgccgagcctgtatagctgcggcagcagctatggcagcgaaaccagcattccggcggcggcgcataccgtgagcaacgcgccggtgaccgaatatatgagccagaacgcgaactttcagaacccgcgctgcgaaaacaccccgctgattggccgcgaaagcccgccgccgagctataccagcagcatgcgcgcgaaatatctggcgaccagccagccgcgcccggatagcagcggcagccat
3.3. Codon Optimization
Codon optimization is necessary because not all organisms use each codon with the same frequency due to differences in the abundance of various tRNAs, so optimizing the codons in the designed sequence will ideally increase the translational yield of my protein. I chose to optimize my sequence for X. laevis because that is the model organism in which I study the role of ttyh3 in neural development.
Using the Twist Bioscience Codon Optimization tool, I generated the following codon-optimized sequence: ATGGCTGGTGTGTCTTATGCTGCTCCTTGGTGGGTCTCTTTATTACATCGGTTGCCACACTTCGACCTCTCCTGGGAAGCCACATCTAGTCAATTCCGACCAGAGGACACAGACTACCAACAAGCACTATTATTGCTAGGGGCTGCCGCTTTAGCTTGTTTGGCTCTTGACCTTCTCTTCCTTTTGTTCTACTCTTTCTGGTTATGTTGTAGAAGAAGGAAGTCAGAGGAGCACCTCGACGCAGACTGTTGTTGTACTGCTTGGTGTGTCATAATCGCTACTCTTGTATGTTCAGCAGGTATAGCAGTAGGATTCTACGGGAATGGTGAGACATCCGACGGAATCCACCGGGCAACTTACTCCCTCAGACACGCTAATAGAACTGTTGCTGGTGTACAAGACCGGGTATGGGACACTGCAGTAGGGTTGAATCACACAGCAGAGCCTTCATTACAAACTTTAGAGAGACAACTTGCTGGAAGACCTGAGCCACTTAGAGCTGTTCAAAGATTACAAGGATTGTTAGAGACGCTCCTAGGGTACACTGCAGCCATCCCATTCTGGCGAAATACTGCCGTATCCTTAGAGGTACTCGCAGAGCAAGTTGACCTCTACGACTGGTACCGATGGCTTGGATACCTTGGGTTGTTGTTGTTGGACGTTATCATATGTTTACTCGTATTAGTTGGACTCATCAGGTCATCTAAGGGAATACTTGTTGGGGTTTGTTTACTTGGGGTTCTTGCTCTCGTCATCTCTTGGGGAGCATTGGGTCTTGAGCTTGCTGTTTCAGTAGGGTCAAGTGACTTCTGTGTAGACCCCGACGCCTACGTCACAAAGATGGTCGAGGAGTACTCAGTTCTTAGTGGAGACATCTTACAATACTACCTCGCTTGTTCACCAAGGGCAGCTAATCCCTTCCAACAAAAGCTTTCAGGTTCTCACAAGGCACTCGTAGAGATGCAAGACGTTGTCGCAGAGTTGCTTAGAACAGTTCCTTGGGAGCAACCAGCAACAAAGGACCCATTGCTCAGAGTCCAAGAGGTCCTTAATGGAACTGAGGTTAATCTCCAACACCTAACAGCCCTTGTAGACTGTCGATCACTCCACTTGGACTACGTCCAAGCTTTGACAGGTTTCTGTTACGACGGAGTTGAGGGTCTAATATACCTCGCCCTTTTCTCCTTCGTTACAGCTCTAATGTTCTCCAGTATCGTTTGTTCTGTTCCCCACACTTGGCAACAAAAGAGAGGACCCGACGAGGACGGAGAGGAAGAGGCAGCACCCGGTCCCAGACAAGCACACGACTCTTTGTACCGGGTCCACATGCCAAGTTTGTACTCATGTGGGTCTTCTTACGGTAGTGAGACAAGTATACCAGCCGCTGCCCACACTGTTTCTAATGCCCCCGTTACAGAGTACATGTCCCAAAATGCAAATTTCCAAAATCCCCGATGTGAGAATACGCCTTTGATAGGACGGGAGAGTCCCCCACCTTCATACACATCATCAATGAGGGCAAAGTACCTTGCAACATCACAACCCCGACCCGACTCCAGTGGATCACAC
3.4. You have a sequence! Now what?
To synthesize the ttyh3 mRNA and protein, I would use an in vitro transcription and translation kit (such as the ones produced by ThermoFischer) to synthesize ttyh3 mRNA and TTYH3 protein, respectively.
3.5. How does it work in nature/biological systems?
Single genes can produce multiple proteins at the transcriptional level through the process of alternative splicing. In eukaryotic cells, transcription and mRNA processing occurs in the nucleus whereas translation occurs in the cytoplasm. After mRNA is synthesized via transcription, mRNA processing (5’ cap addition, poly(A) tail addition, and intron splicing) occurs and exons (which make the coding region of the protein) are annealed together after introns are removed. However, the combinations of different exons (alternative splicing) can produce different proteins from the same primary (immature) mRNA transcript.
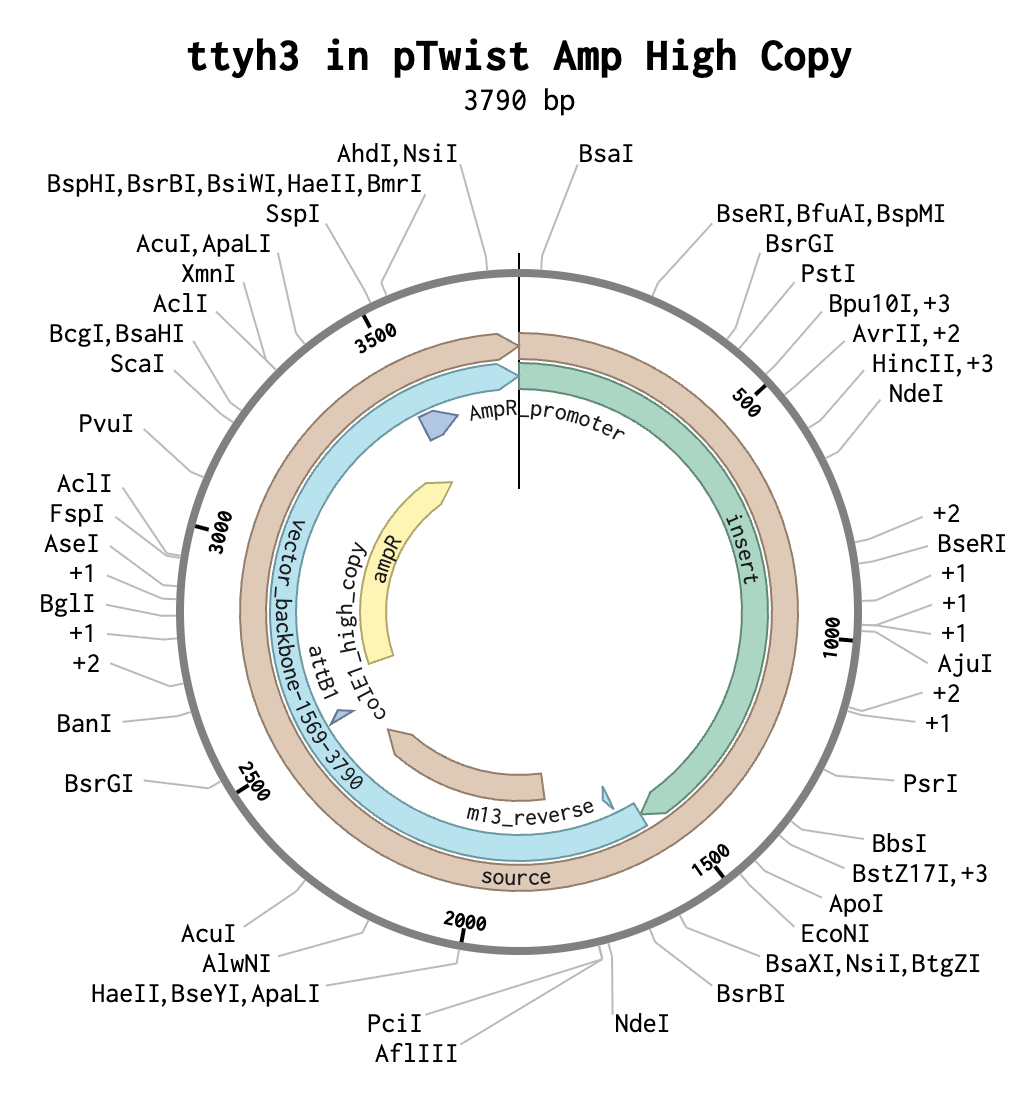
Part 4: Prepare a Twist DNA Synthesis Order

Part 5: DNA Read/Write/Edit
5.1. DNA Read
What DNA would you want to sequence (e.g., read) and why? I would like to sequence the DNA of a tumor sample from a cancer in which ttyh3 is observed to be misregulated (i.e. HCC, genitourinary cancers, colorectal cancer, etc.). I would be interested to observe which mutations in ttyh3 gene lead to functional changes that promote tumorigenesis, tumor invasiveness, etc. and which changes have little to no impact.
In lecture, a variety of sequencing technologies were mentioned. What technology or technologies would you use to perform sequencing on your DNA and why? I would use Illumina whole-genome sequencing (WGS), a second-generation sequencing method, because it is a relatively quick method of obtaining a base-by-base view of any single nucleotide variants (SNVs) in the genome which is essential information for understanding how mutations in the ttyh3 gene increase tumor proliferation, invaseiness, or aggression. The input for this sequencing is gDNA extracted from a tumor biopsy and prepared using the Illumina DNA Library Prep Kit. Following DNA extraction, the sample is sequenced using the “sequencing by synthesis” (SBS) technology which utilizes dNTPs bound to fluorescently-labeled reversible terminators to decode each base. The output of Illumina sequencing is a FASTQ file with quality scores.
5.2. DNA Write
What DNA would you want to synthesize (e.g., write) and why? Completely unrelated to my work with X. laevis, I think it would be fascinating to design a phage with a novel gene (or perhaps an entirely synthetic genome!). The protein structure could be predicted with a tool like AlphaFold, then the amino acid sequence could be reverse transcribed into cDNA for the gene of interest and integrated into a plasmid. As advancements in phage therapy show promise in combating antibiotic resistance, it would be fascinating to design a protein that would increase phage DNA replication, phage entry, etc. to increase the efficacy of the treatment.
What technology or technologies would you use to perform this DNA synthesis and why? First, and I don’t know if this is possible, I would create the desired 3D protein structure with a tool like AlphaFold and obtain the amino acid sequence. Then, similarly to this homework assignment, I would generate cDNA by reverse transcribing the AA sequence and order a plasmid from Twist (or elsewhere) with the gene of interest.
5.3. DNA Edit
What DNA would you want to edit and why? I would like to develop a technology to perform somatic editing of the DNA of individuals with genetic diseases that result from mutations in a single gene (i.e. Huntington’s, cystic fibrosis, etc.). While many similar technologies already exist and some clinical trials have shown promise (i.e. sickle cell anemia!), I would like to expand these technologies to other genetic diseases.
What technology or technologies would you use to perform these DNA edits and why? I would use CRISPR/Cas9-mediated homlogy directed repair to excise the mutated gene and replace it with a nonmutated, functional gene. This would involve design of the nonmutated gene and the sgRNA to direct the Cas9 enzyme to the appropriate cut site. The limitations of CRISPR/Cas9 technology are possible off-target effects (low precision) or the construct not being effectively delivered to all cells (low efficiency).