Week 4 HW: Protein Design Part I
Part A: Conceptual Questions
Part B: Protein Analysis and Visualization
Part C: Using ML-Based Protein Design Tools
C1: Protein Language Modeling
C2: Protein Folding
C3: Protein Generation
Part D: Group Brainstorm on Bacteriophage Engineering
I collaborated with Heather Qian on this assignment!
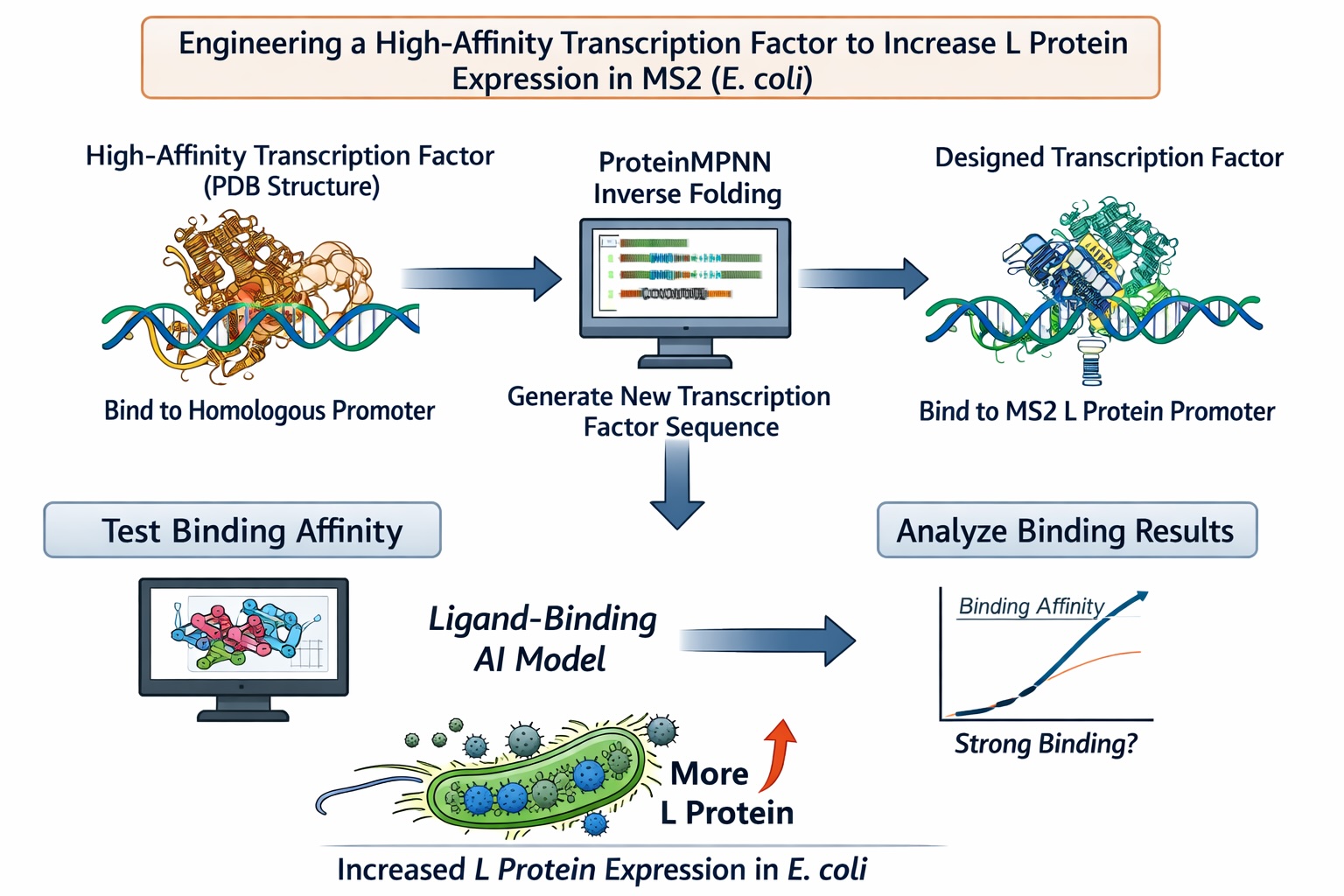
Computational Goal: We will attempt to increase the titer of the L protein expressed by MS2 in the E. coli host.
Overall solution: create a new transcription factor that binds very tightly to the promoter, increasing expression of the L protein
Inverse protein folding using ProteinMPNN
- Use structure of a transcription factor (a PDB file) that binds with very high affinity to a promoter that is highly homologous to the L protein promoter
- Input the structure of the aforementioned transcription factor into ProteinMPNN along with the amino acid sequence of a native transcription factor that binds to the L protein promoter
- This will theoretically generate the sequence of a protein that is structurally similar to a transcription factor with high DNA-binding affinity but is specific to the L protein promoter
Confirm the binding affinity between our designed transcription factor and the MS2 L protein promoter with a ligand-binding AI model
Why will these tools accomplish our computational goal?
- Protein MPNN is an inverse-folding algorithm. The sequence of the L protein promoter and other transcription factors that bind to this promoter are known. However, to generate a transcription factor with higher affinity for this promoter sequence than the native L protein transcription factors, we will model our inverse-folding after an existing transcription factor that behaves in the manner we envision (binds with high affinity) for our engineered transcription factor. As discussed in this week’s lecture, existing AI algorithms are good at designing structures similar to previously-characterized structures but less good at designing different (novel) structures. As such, our inverse-folding approach hinges on the existence of another high affinity transcription factor for a homologous promoter
- Using a ligand-binding AI model will provide a computational indication of the success of our engineering without the need to use an in vitro or in vivo model
Possible pitfalls:
- We are unable to find a transcription factor that binds to a highly homologous promoter sequence with high affinity :(
- Inaccuracies in AI predictions :(