Week 6 HW: Genetic Circuits Part I
DNA Assembly
- What are some components in the Phusion High-Fidelity PCR Master Mix and what is their purpose?
- DNA polymerase: uses dNTP monomers to synthesize new DNA strands
- dNTPs: monomers of new DNA strand
- Buffer: stabilizes the pH of the reaction for optimal enzymatic function
- MgCl2: Cofactor for the polymerase; optimizes enzymatic function and primer annealing
- What are some factors that determine primer annealing temperature during PCR?
- GC content (higher GC content = more hydrogen bonds = stronger primer annealing = higher annealing temperature)
- Primer length (longer primers = more hydrogen bonds = stronger primer annealing = higher annealing temperature)
- There are two methods from this class that create linear fragments of DNA: PCR, and restriction enzyme digests. Compare and contrast these two methods, both in terms of protocol as well as when one may be preferable to use over the other.
- PCR:
- Enzyme: DNA polymerase
- DNA polymerases bind to primers and synthesize complementary DNA to the template strand
- Purpose: amplify!
- Polymerases synthesize DNA by recognizing primers (designed to flank the specific region of interest) and incorporating dNTPs into a novel DNA strand
- When to use:
- To detect a specific sequence within a mixed sample
- To create more of a specific DNA sequence (amplify)
- Enzyme: DNA polymerase
- Restriction enzyme:
- Enzyme: restriction endonuclease
- Restriction endonucleases recognize specific nucleotide sequences (4-8 bp) in double-stranded DNA and cut in a specific pattern (blunt or sticky ends, depending on the enzyme)
- Purpose: cut!
- Restriction enzymes cut existing template DNA, they do NOT amplify DNA fragments
- When to use:
- To linearize bacterial plasmids (ex. for in vitro transcription of capped mRNA for microinjection into X. laevis!)
- To cut DNA fragments to assemble/ligate together and transform into a plasmid
- Enzyme: restriction endonuclease
- How can you ensure that the DNA sequences that you have digested and PCR-ed will be appropriate for Gibson cloning?
- Primer design!
- The 5’ tail overhang of each primer should be identical to the adjacent DNA fragment
- the 3’ end of the primer should be complementary to the DNA fragment to which it will anneal
- Restriction Digest:
- Restriction enzyme cut sites should be avoided
- PCR:
- Primers should be designed that flank the overhang regions
- How does the plasmid DNA enter the E. coli cells during transformation?
- Heat shock transformation - briefly raising temperature increases the permeability of the bacterial cell wall, allowing the plasmid DNA to enter into the E. coli cell
- E. coli cell sample placed on ice for ~15-30 minutes
- Sample heated to 42ºC for ~15-30 seconds
- Sample then returned to ice for ~5 minutes
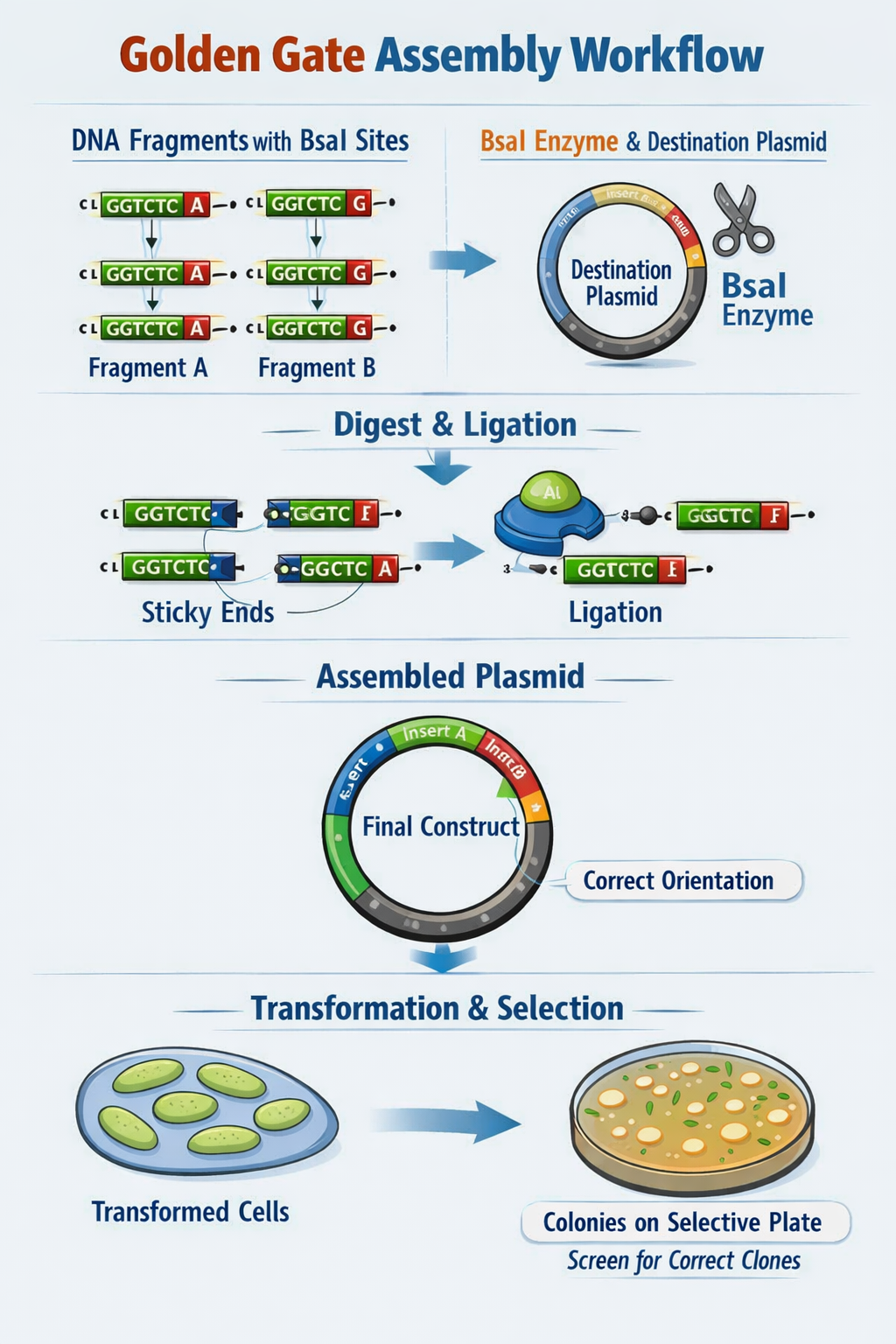
- Describe Golden Gate Assembly Golden Gate Assembly utilizes Type IIS restriction enzymes (such as BsaI) which recognize non-pallindromic sequences, cut outside the recognition site (to avoid damaging the DNA sequence of interest), and create sticky ends of variable lengths. First, template DNA is amplified using PCR with primers specified to include the TIIS recognition and cut sites. Next, a restriction digest with the TIIS enzyme is performed to create DNA fragments with sticky ends. The sticky ends of adjacent fragments should be complementary so the sequences can be ligated in the appropriate order. Following ligation of DNA fragments with a plasmid, the engineered plasmid can be transformed into a bacterial cell (ex. E. coli), and bacterial colonies containing the plasmid can be screened for furhter use.
Asimov Kernel
Repressilator
The protein concentrations generated from this recreation of the Repressilator meet my expectations in that they oscillate over time. However, unlike the in vivo system, the oscillations do not decay over time in this computational model.
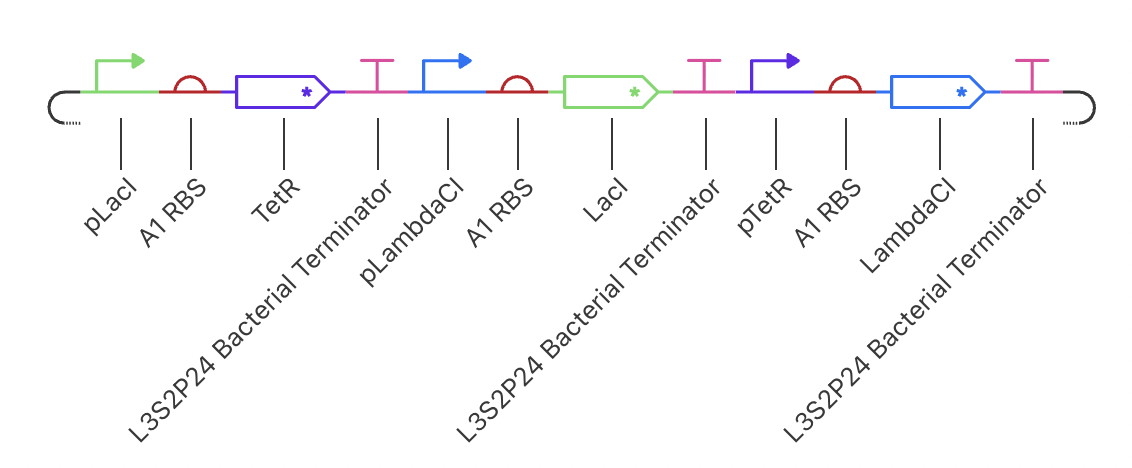


Construct 1: Negative Feedback Loop
I have designed a LacI genetic circuit in which the LacI gene should bind to its own promoter and inhibit its own expression. I would expect the RNA and protein concentration to fluctate slightly, then reach steady state.
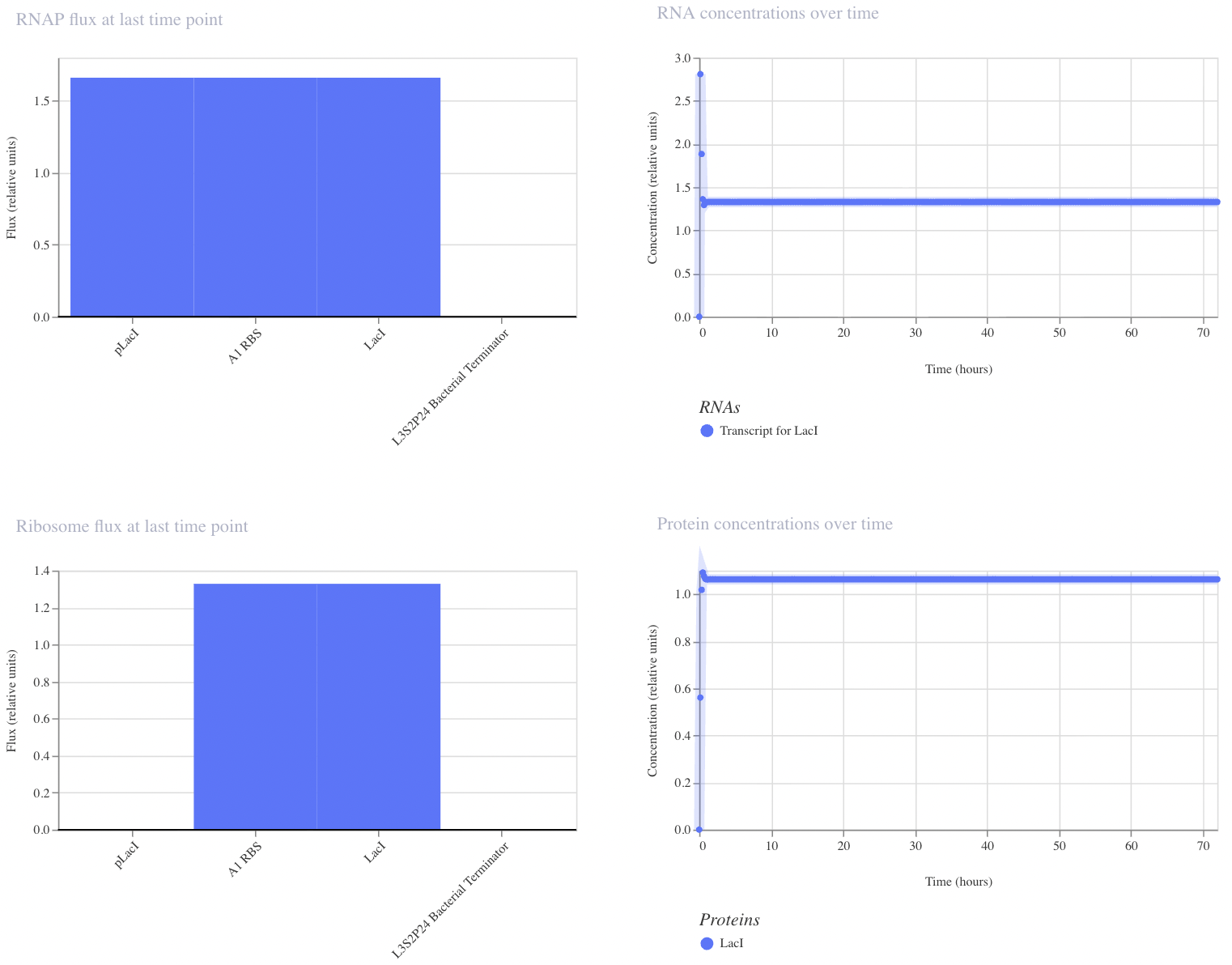

My system immediately reaches steady state, which is not what I predicted. However, I was unable to add an operator sequence in this circuit, so perhaps that is my issue.
Construct 2: Toggle Switch
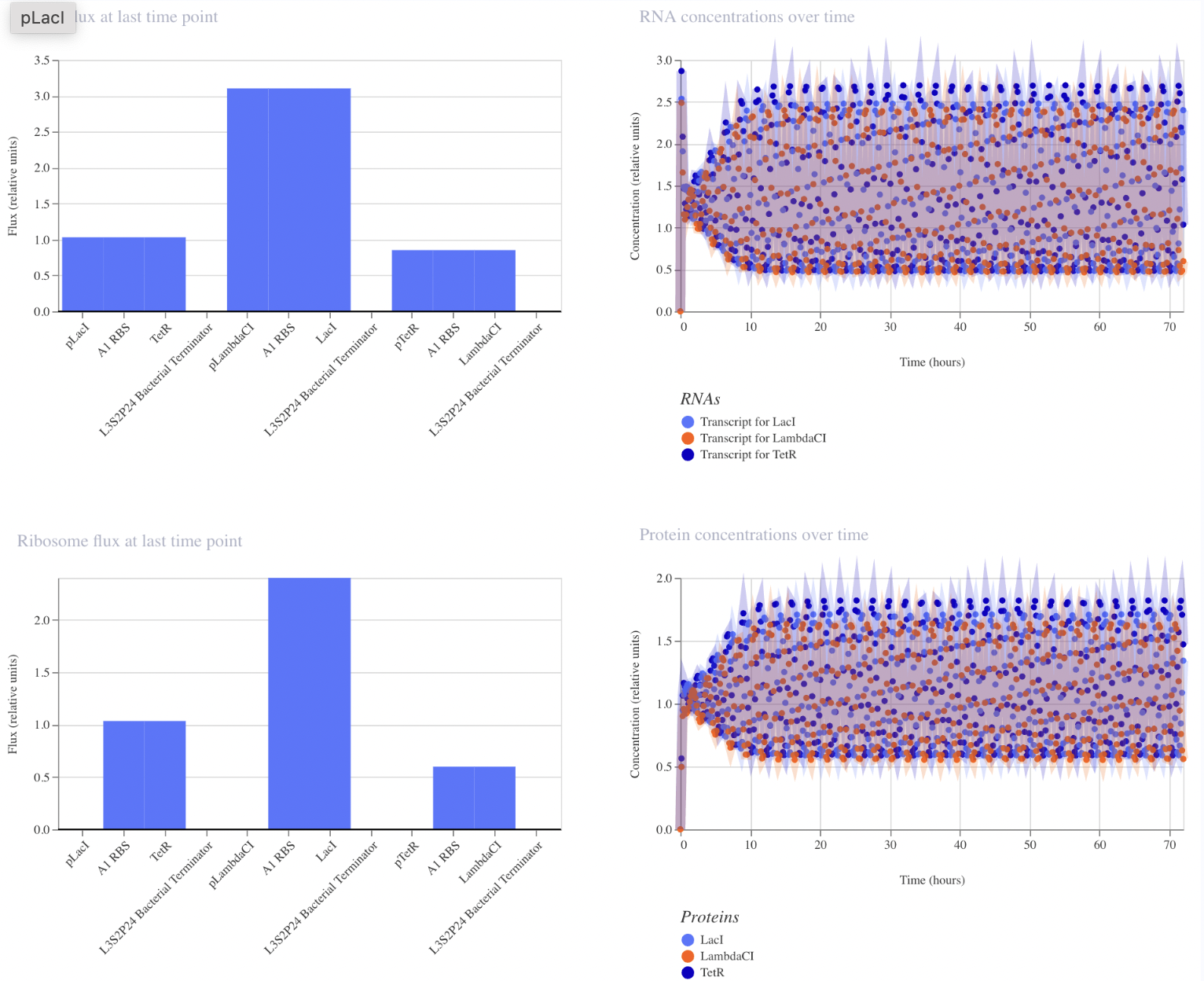
Modeled after the classical toggle switch genetic circuit of mutual repression, I expect this construct to reach a bistable state. If I were able to add an inducer to this system, I would expect to be able to switch which bistable state is selected by the system. However, in this instance, my system reached a single steady state.
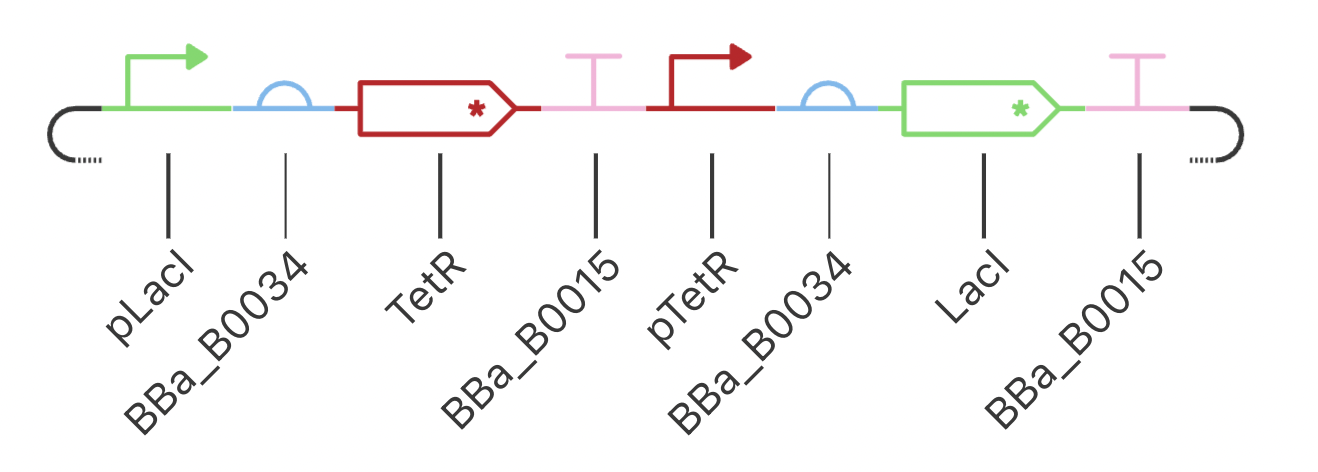

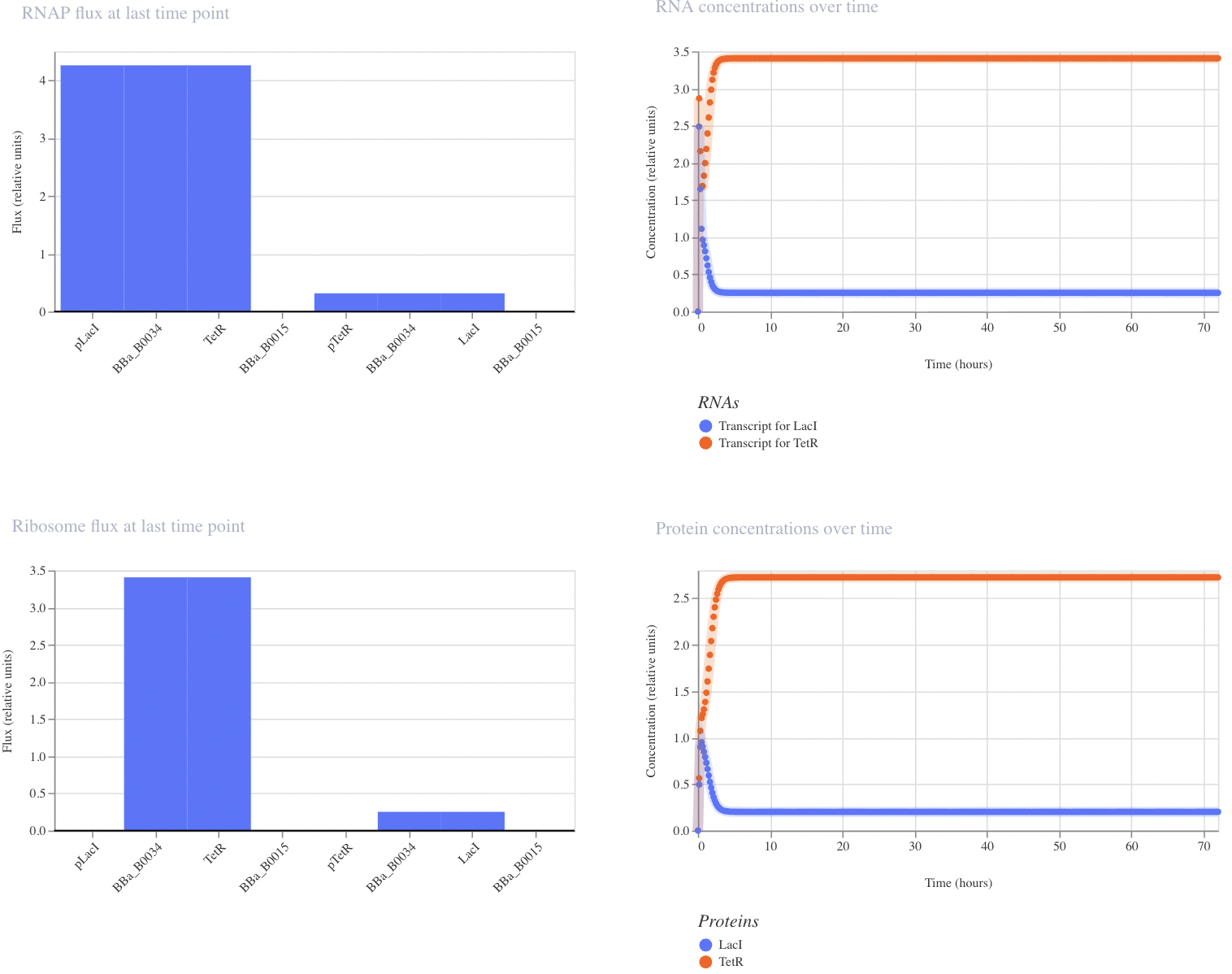

My system behaves the way I predicted it would!
Construct 3: Dual Repressors
My final circuit contains two repressors, LacI and TetR, controlled by the QacR promoter. Additionally, the expression of QacR is controlled by the expression of either LacI or tetR (two separate QacR genes). I expect the system to reach a single steady state. If I was able to design an inducible promoter, I could be able to switch between stable states.
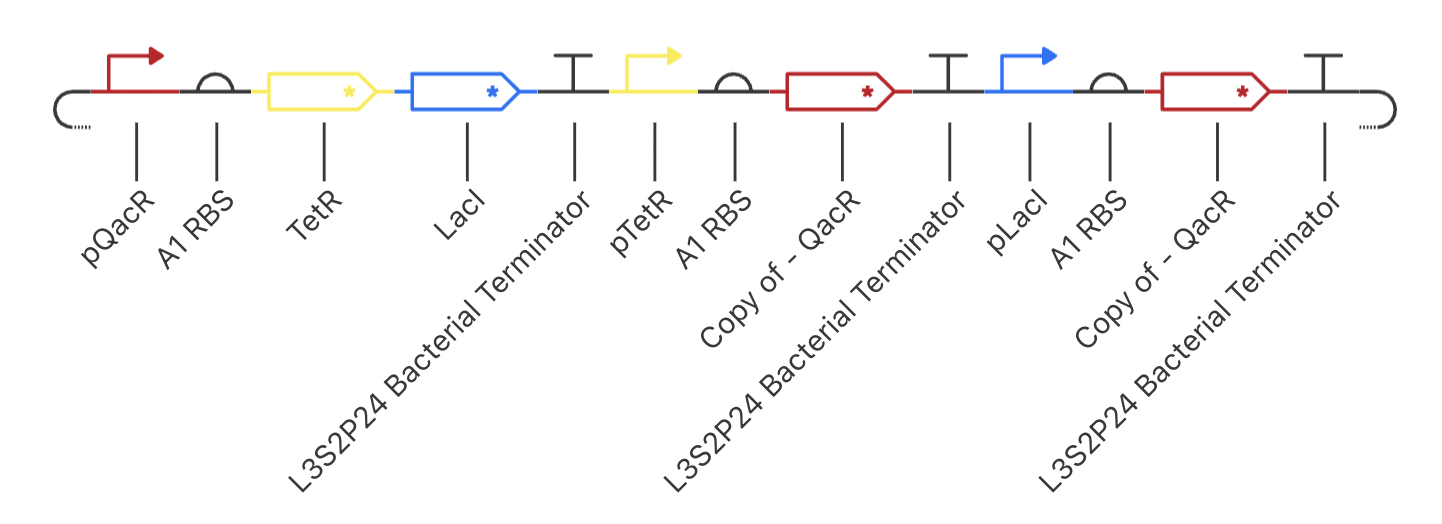

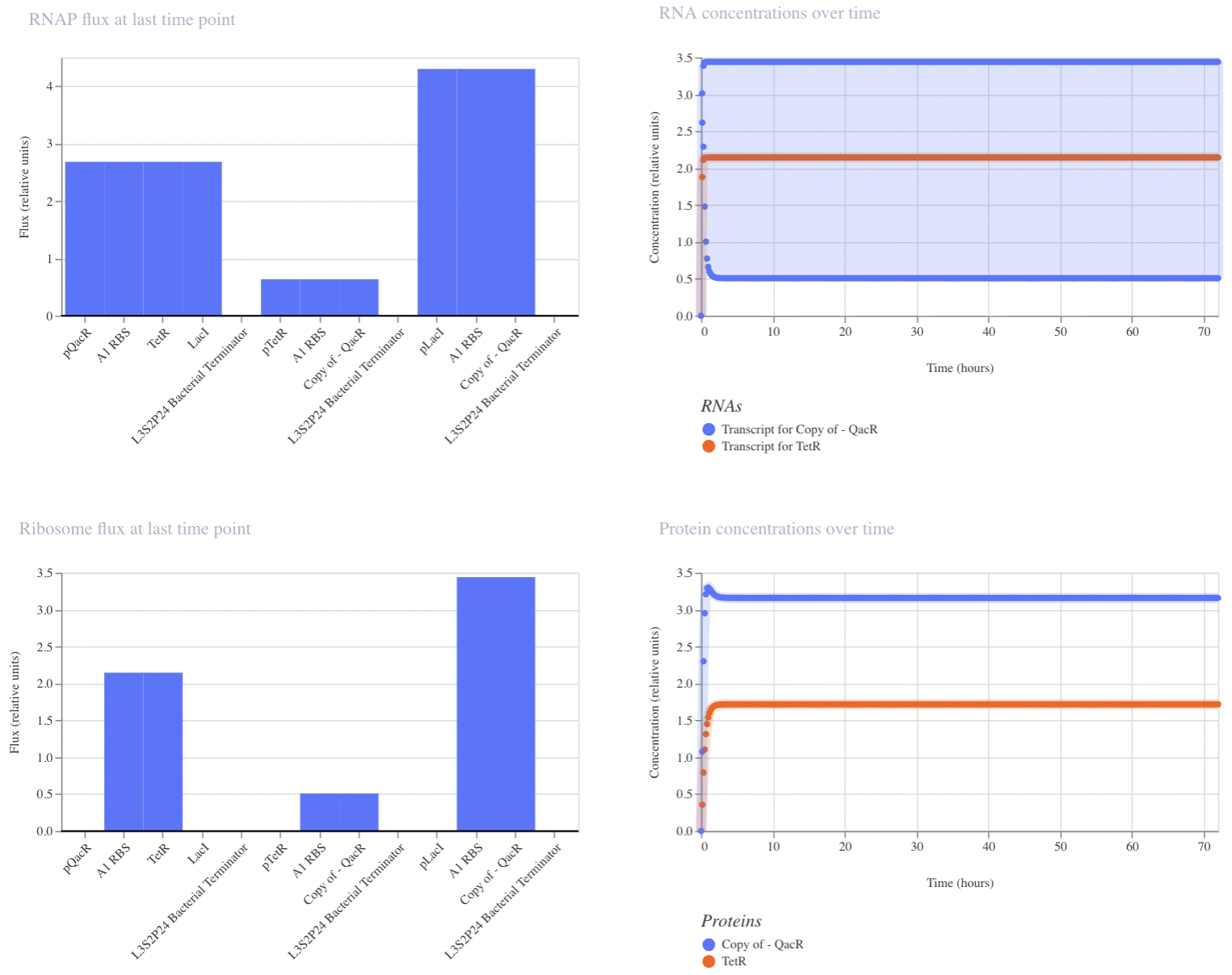

My system behaved as predicted :)