Week 3 HW: Lab Automation

I created two different agar art designs using two Arabic calligraphy styles. For the first design, I used a simple calligraphy style and created it directly using Python scripting in a Google Colab notebook. For the second design, I used the Opentrons Automation Art interface to design the calligraphy and obtain the coordinates.
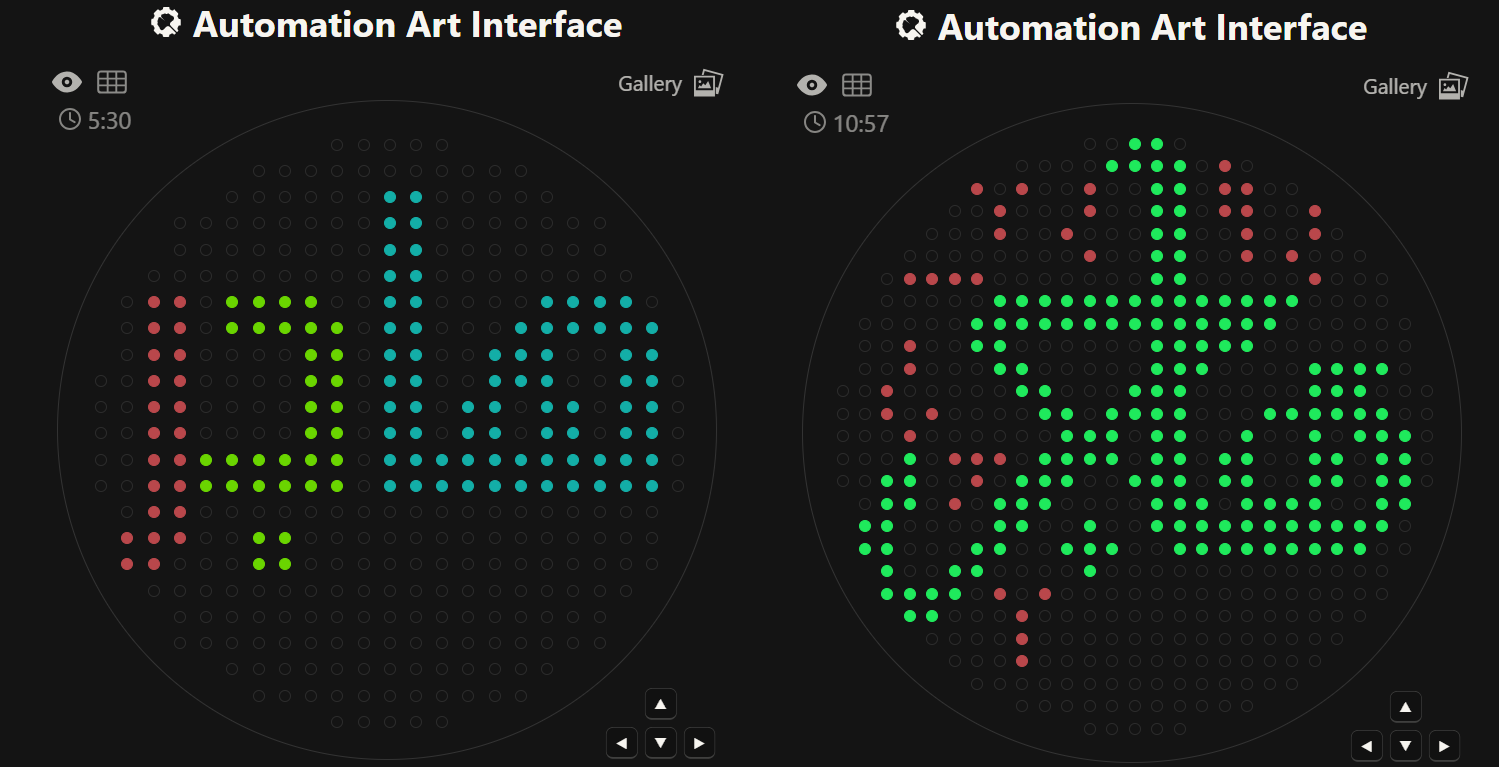

I used the Google Gemini AI tool in Colab to understand the logic of the example Opentrons scripts provided in the lab. It helped me understand how coordinates, loops, and pipetting commands work. I also used Gemini AI to help identify and correct mistakes in my Python script, such as indentation errors. I reviewed the suggestions and edited the final code myself.
- Featured Article: Automated Assembly of Programmable RNA-Based Sensors


The research aimed to solve the challenge of rapidly designing and building large libraries of RNA sensors that can “sense” specific viral RNA signatures. These sensors are crucial for diagnostic applications and understanding RNA-protein interactions. The authors focused on the biological validation of these sensors in both in vivo (bacteria) and cell-free systems.


They used the following lab automation:
- Hardware: Hamilton Microlab STAR liquid-handling workstation.
- Software: Custom Python scripts integrated with the liquid handler’s control software to manage complex plate layouts and reaction conditions.
The researchers used the automated system as a tool to facilitate:
- High-Throughput Plasmid Assembly: The authors needed to construct 144 unique plasmids encoding different riboregulator designs. Doing this manually would be prone to pipetting errors and extremely time-consuming.
- Library Preparation: Automation was used to prepare DNA libraries and reaction mixes for cell-free protein synthesis assays, ensuring consistent reagent volumes across hundreds of samples.
- Normalization and Dilution: The Hamilton system handled the precise normalization of DNA concentrations across plates, which is critical for accurate comparative screening of sensor performance.
The study successfully identified several high-performing RNA sensors capable of detecting viral targets. The use of automation allowed the team to scale their construction phase by nearly 10-fold compared to manual workflows, enabling them to test a much wider range of biological designs than previously possible. For understanding the content of this artical and which type of Lab automation the authors used in their research , i used the AI tool “SCISPACE”.
- Final project Lab Automation:
My final project focuses on developing an in silico model of a lactose-responsive probiotic that produces lactase only when lactose is present. The physical implementation of this model would allow laboratory automation to verify its predicted results through experimental tests. A liquid-handling robot such as Opentrons could be used to prepare a multi-well plate containing a gradient of lactose concentrations. The robot would then inoculate each well with the engineered probiotic strain, and perform timed sampling to measure lactase activity or reporter output. The automated workflow enables scientists to perform systematic and repeatable tests on lactose responses of the genetic circuit. This helps them match their experimental results with their computer-based model. The project currently exists as a computational project which will use automation as a future extension of the project which does not require automation for its current research activities.
1st Idea: In-Silico Model of an Engineered Probiotic Producing Lactase in Response to Lactose
- Problematic:
Many people cannot digest lactose because they lack enough lactase in their intestine. A possible solution is to use engineered probiotics that produce lactase only when lactose is present.
Before building such probiotics in the laboratory, it is important to understand how the genetic system would behave. So, without computational modeling, designing these systems requires trial-and-error experiments that are slow and expensive.
There is a need for a simple computational model that can predict how a lactose-responsive genetic circuit would control lactase production over time. Image_ref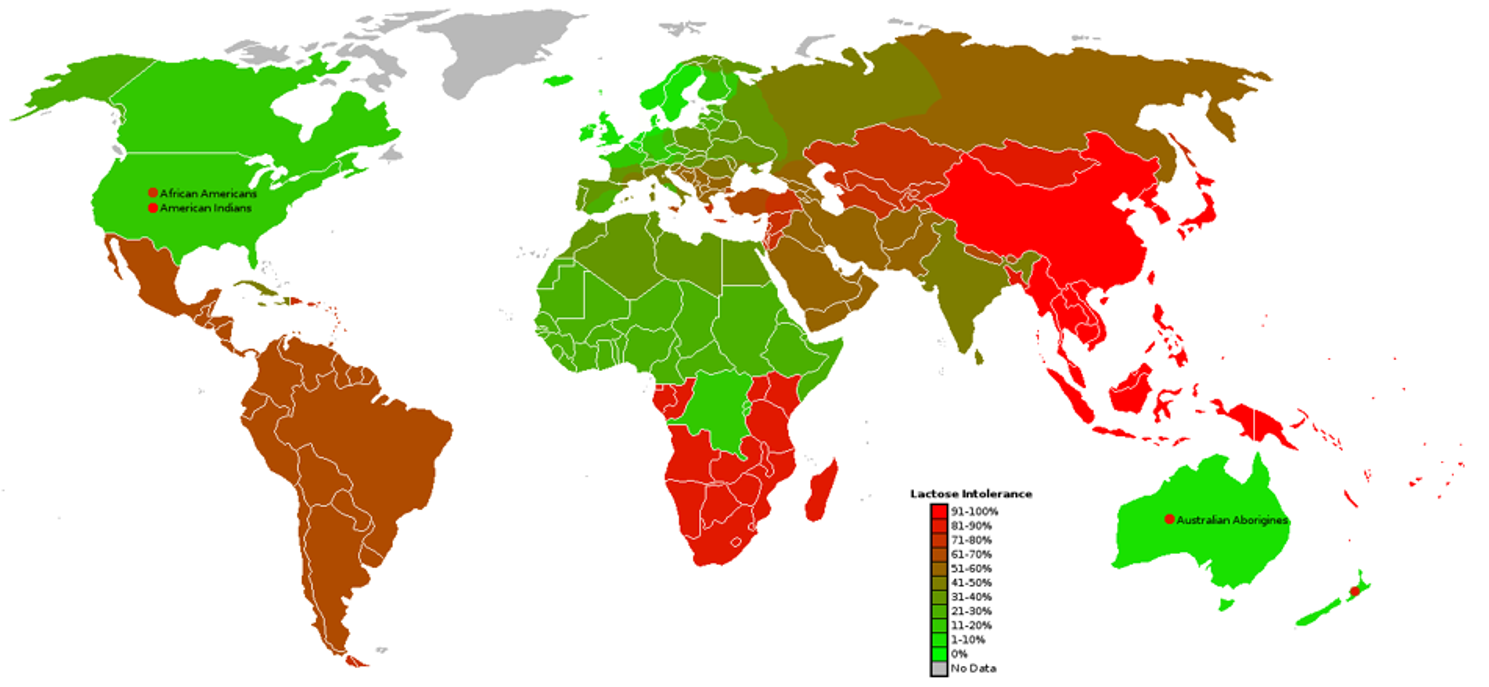

- Objectives:
The project is based on a lactose-responsive genetic cassette, who’s dynamic behavior is modeled as a genetic circuit in silico. The objectives of this project are:
- –> To build an in-silico model of a lactose-responsive genetic circuit.
- –> To simulate how lactose stimulate lactase production.
- –> To study how changing key parameters affects lactose degradation.
- –> To explore system behavior completely in silico.
- Project Description:
The project develops a purely computational model of an engineered probiotic strain. The model is based on a lactose-responsive genetic cassette, whose dynamic behavior is represented in silico as a genetic circuit: – a lacA promoter, operator, and native RBS from Lactococcus lactis for lactose sensing and regulation, – the lacZ gene from Escherichia coli K-12, encoding β-galactosidase (lactase).
I used an AI tool (ChatGPT) to guide me about the repression mechanism I should use, and its response was as follows: To ensure realistic behavior in the model, the lacA promoter includes a native operator, normally repressed by a LacR-like protein in Lactococcus lactis. In the simulation, a repression term is included to prevent unnecessary accumulation of lacZ (lactase) when lactose is absent.

 The model simulates how the presence of lactose activates the promoter, leading to lactase production, and how this enzyme then degrades lactose over time.
No DNA construction or wet-lab experiments are performed. All behavior is represented mathematically and simulated using a computer.
The model simulates how the presence of lactose activates the promoter, leading to lactase production, and how this enzyme then degrades lactose over time.
No DNA construction or wet-lab experiments are performed. All behavior is represented mathematically and simulated using a computer.
- Steps to Achieve the Project:
- Define simplified biological assumptions (single strain, constant environment).
- Represent lactose as the input signal.
- Model promoter activation based on lactose concentration.
- Model lactase production and degradation over time.
- Model lactose degradation by lactase.
- Run simulations to observe system behavior.
- Change parameters to study different scenarios.
- Limitations:
- The model does not include other gut microbes.
- The gut environment is assumed constant.
- Results are predictive, not experimentally validated.
2nd Idea:Engineering an E. coli Reporter Strain to Monitor Protein Aging During Heterologous Expression Using a Fluorescent Timer Protein
- Problematic:
Escherichia coli BL21(DE3) is one of the most widely used hosts for heterologous protein expression in research and biotechnology. Although protein expression levels can be easily measured, there are very limited tools to determine how long the expressed protein molecules have persisted inside the cell. During prolonged induction, proteins may accumulate, age, misfold, or lose functionality, even when expression appears successful. Most current methods detect protein quality only after purification, making optimization of expression conditions slow and inefficient. So, there is a need for a genetically encoded reporter system that can estimate protein aging in living cells during expression. Image_ref
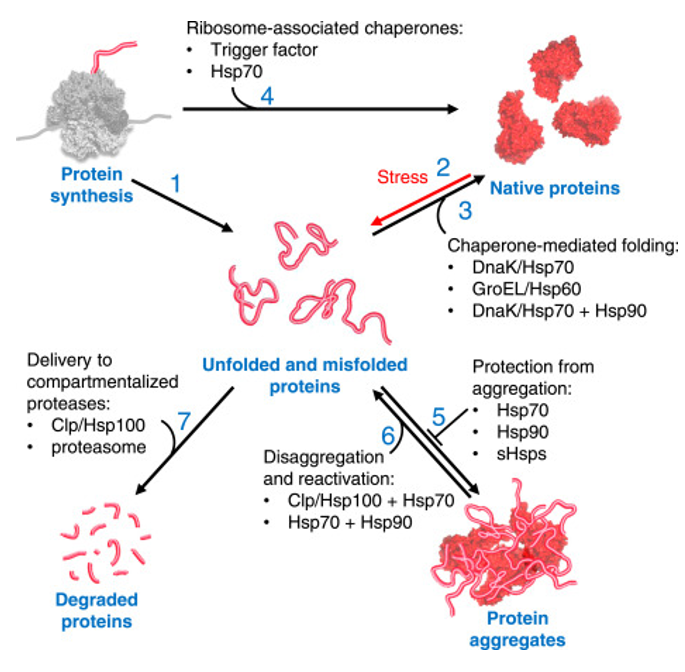

- Objectives:
This project is based on a fluorescent timer protein–based reporter system integrated into a heterologous protein expression strain. The objectives are:
- –> To engineer a reporter strain capable of estimating protein age in vivo.
- –> To use a fluorescent timer protein to distinguish newly synthesized and older proteins.
- –> To monitor protein aging during prolonged heterologous expression.
- –> To provide a practical tool for optimizing protein expression conditions.
- Project Description:
The project focuses on the genetic engineering of a protein expression strain of E. coli BL21(DE3). The reporter system is based on a genetic fusion between:
- a protein of interest (POI) expressed under the T7 promoter, and
- a fluorescent timer protein whose emission spectrum changes over time after synthesis. The genetic construct consists of:
- a T7 promoter and ribosome binding site,
- the gene encoding the protein of interest,
- a flexible linker sequence,
- the fluorescent timer protein gene,
- a transcriptional terminator. Image_ref
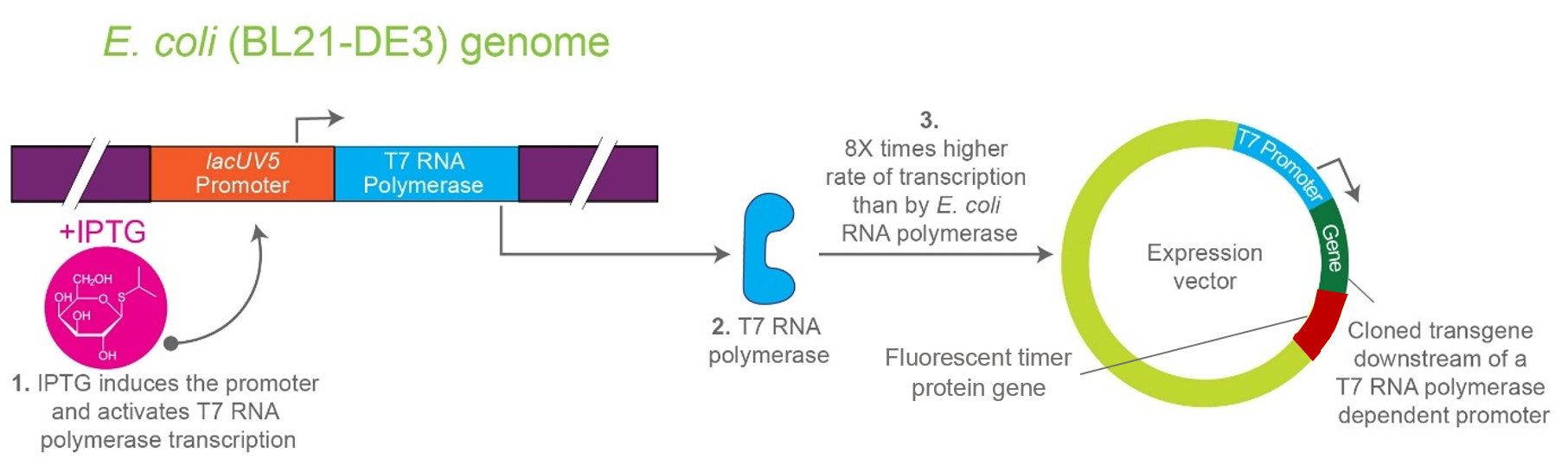

After induction, newly synthesized POI–timer fusion proteins initially emit one fluorescent signal. As time progresses, the timer protein matures and shifts to a second fluorescent signal. The ratio of the two fluorescence signals provides an estimate of the age distribution of the expressed protein population.
I used AI tool (ChatGPT) version to refine questions related to the necessary genetic elements required for T7-based heterologous expression in Escherichia coli BL21(DE3) and to determine the appropriate placement of a fluorescent timer gene for monitoring the age of the expressed protein.


- Steps to Achieve the Project:
- Select a heterologous protein suitable for expression in E. coli.
- Design a genetic fusion between the protein of interest and a fluorescent timer protein.
- Clone the fusion construct under a T7 promoter into an expression plasmid.
- Transform the plasmid into E. coli BL21(DE3).
- Induce protein expression using IPTG.
- Monitor fluorescence signals over time using appropriate excitation/emission settings.
- Calculate fluorescence signal ratios to estimate protein aging.
- Compare protein aging under different induction times and expression conditions.
- Limitations:
- Fusion of the timer protein may affect protein folding or function.
- Protein damage mechanisms are not directly measured.
3rd Idea:Engineering Houseplants for Atmospheric Carbon Monoxide (CO) Capture
- Problematic:
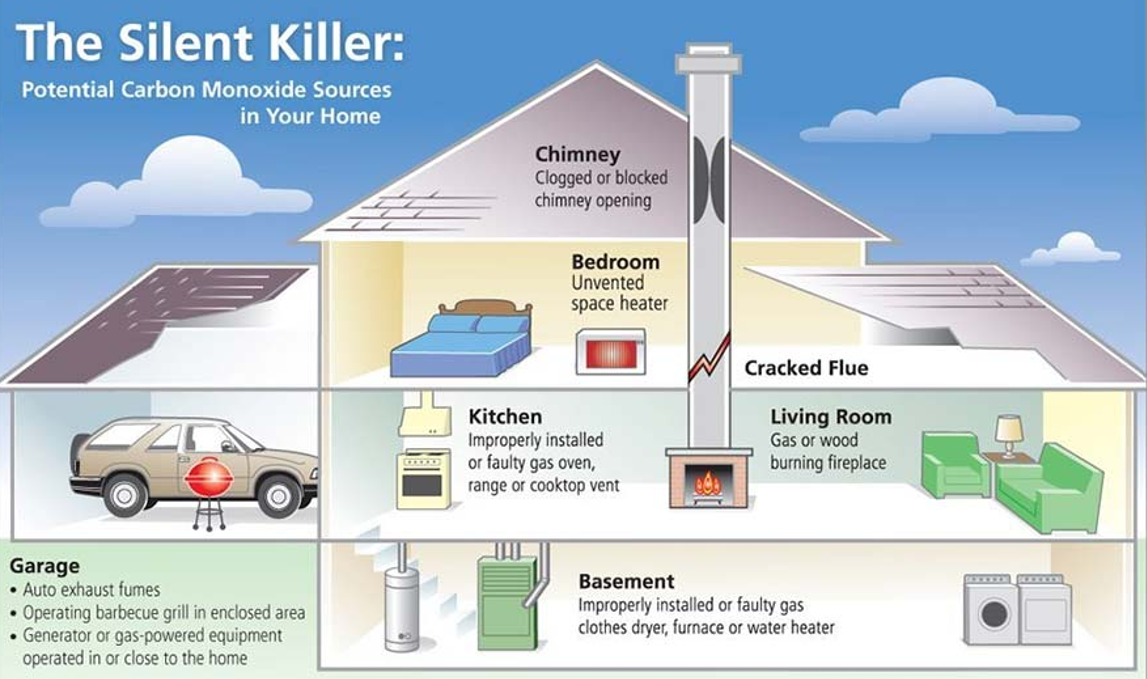
Carbon monoxide (CO) is a toxic gas produced by cars, heaters, and incomplete combustion. It is dangerous for humans, especially in indoor environments. Current solutions such as CO detectors can detect the gas but cannot remove it.
Some bacteria naturally use CO as an energy source and convert it into carbon dioxide (CO₂). However, common houseplants cannot metabolize CO. If plants could be engineered to convert CO into CO₂, they could act as natural biological air filters. Image_ref

- Objectives:
The objectives of this project are:
- –> To engineer a houseplant capable of converting carbon monoxide into carbon dioxide.
- –> To use microbial genes that naturally perform CO oxidation.
- –> To ensure the system works safely in oxygen-rich (indoor) environments.
- –> To allow the produced CO₂ to be reused by the plant’s normal photosynthesis.
- –> To design a genetically stable and safe indoor plant system.
- Project Description:
This project engineers a plant to express a bacterial enzyme called carbon monoxide dehydrogenase (CODH). This enzyme converts carbon monoxide (CO) into carbon dioxide (CO₂). The CO₂ produced by this reaction is not wasted. Instead, it enters the plant’s natural photosynthetic pathway (Calvin cycle), where it can be fixed into sugars. The plant therefore detoxifies CO while continuing its normal metabolism. The system is designed to work only when CO is present, to avoid unnecessary energy use.
- Genetic Elements for construct design:
- CO Oxidation Enzymes
The core of the system is the carbon monoxide dehydrogenase (CODH) enzyme, which is responsible for converting carbon monoxide (CO) into carbon dioxide (CO₂). This enzyme is composed of three subunits encoded by the genes coxL, coxM, and coxS. The coxL gene encodes the large catalytic subunit, coxM encodes a subunit involved in electron transfer, and coxS encodes a structural subunit that stabilizes the enzyme complex. These genes originate from Oligotropha carboxidovorans, a bacterium that can oxidize CO in the presence of oxygen, making it suitable for expression in plant cells.
- Promoter (Gene Expression Control)
To drive the expression of the CODH genes in plant cells, the CaMV 35S promoter is used. This promoter originates from the Cauliflower mosaic virus and is one of the most widely used promoters in plant biotechnology. It enables strong and constitutive gene expression across many plant tissues and is well characterized, making it a reliable choice for this project.
- Subcellular Targeting Signal
A chloroplast transit peptide is included to ensure that the CODH proteins are transported into the chloroplast after synthesis. This targeting signal is derived from the small subunit of the plant enzyme Rubisco, which naturally localizes to the chloroplast. By directing the CODH enzymes to the chloroplast, the CO₂ produced from CO oxidation is generated close to the photosynthetic machinery, allowing it to be efficiently reused by the plant during photosynthesis.
- Transcription Terminator
The NOS terminator is used to ensure proper termination of transcription and stable gene expression. This terminator originates from Agrobacterium tumefaciens and is commonly used in plant genetic constructs. Its function is to signal the end of transcription, improving mRNA stability and ensuring reliable expression of the introduced genes.
- Steps to Achieve the Project:
- Select CO-oxidation genes from aerobic bacteria.
- Adapt bacterial gene sequences for plant expression (codon optimization).
- Design a plant expression construct containing:
- Plant promoter
- CODH genes
- Chloroplast targeting signal
- Transcription terminator Image_ref
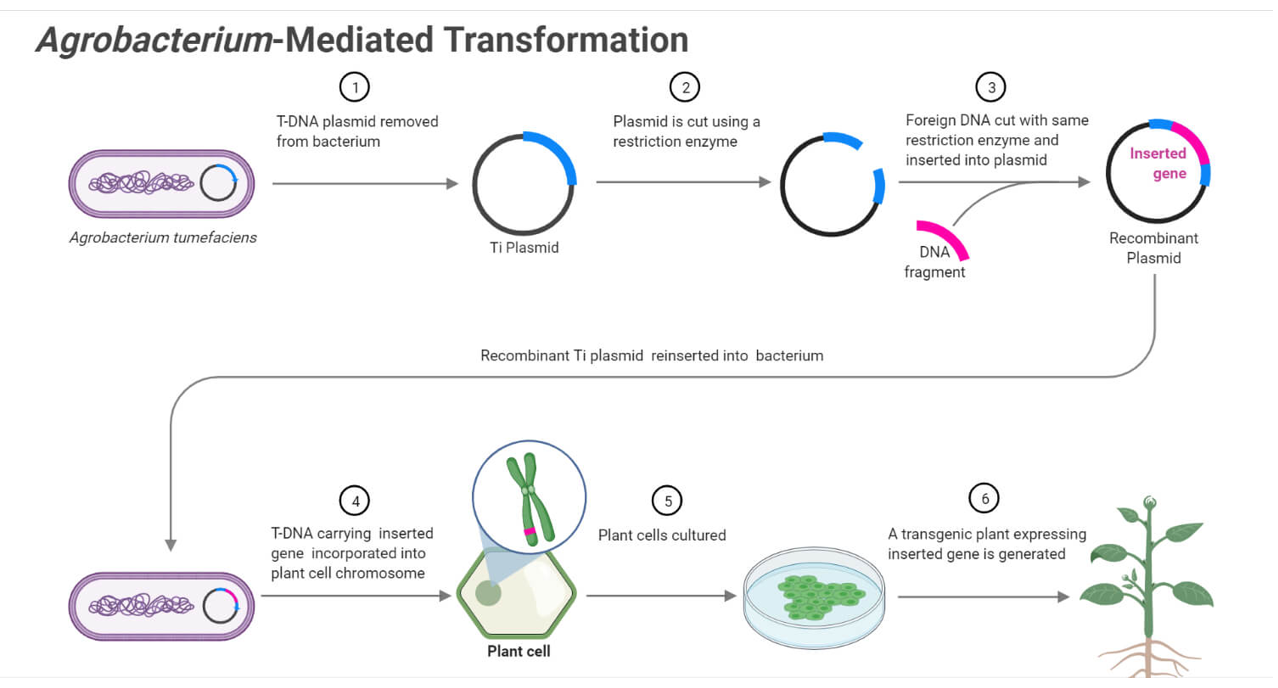

- Introduce the construct into the plant genome.
- Confirm expression of CODH proteins in plant cells.
- Evaluate CO removal and plant health in controlled conditions.
- Assess whether produced CO₂ supports normal photosynthesis.
- Limitations:
- Plant genetic engineering is slow and complex.
- CO uptake by plants may be limited.
- CO metabolism efficiency may be low.
in this homework, AI ChatGPT also assisted me in organizing and clearly articulating my answers and descriptions, ensuring that the content is well-structured and easy to understand.
Sources:
- Bidwell, R., & Bebee, G. (2011). Carbon monoxide fixation by plants. Canadian Journal of Botany, 52, 1841–1847. https://doi.org/10.1139/b74-236 Buckley, S. (2024, September 12). Bioengineered Plants Offer Superior Indoor Air Purification. Sustainable Brands. https://sustainablebrands.com/read/bioengineered-plants-indoor-air-purification
- DesignWorksGarage. (n.d.). NCHH. NCHH. Retrieved February 24, 2026, from https://nchh.org/information-and-evidence/learn-about-healthy-housing/health-hazards-prevention-and-solutions/carbon-monoxide/
- Hartl, F. U., Bracher, A., & Hayer-Hartl, M. (2011). Molecular chaperones in protein folding and proteostasis. Nature, 475(7356), 324–332. https://doi.org/10.1038/nature10317
- Heiss, S., Hörmann, A., Tauer, C., Sonnleitner, M., Egger, E., Grabherr, R., & Heinl, S. (2016). Evaluation of novel inducible promoter/repressor systems for recombinant protein expression in Lactobacillus plantarum. Microbial Cell Factories, 15(1), 50. https://doi.org/10.1186/s12934-016-0448-0
- Orina, F., Amukoye, E., Bowyer, C., Chakaya, J., Das, D., Devereux, G., Dobson, R., Dragosits, U., Gray, C., Kiplimo, R., Lesosky, M., Loh, M., Meme, H., Mortimer, K., Ndombi, A., Pearson, C., Price, H., Twigg, M., West, S., & Semple, S. (2024). Household carbon monoxide (CO) concentrations in a large African city: An unquantified public health burden? Environmental Pollution, 351, 124054. https://doi.org/10.1016/j.envpol.2024.124054
- Promoter and terminator considerations for gene integration into the genome of E.coli? (n.d.).ResearchGate. Retrieved February 24, 2026, from https://www.researchgate.net/post/Promoter_and_terminator_considerations_for_gene_integration_into_the_genome_of_Ecoli
- Robson, J. M., Arevalos, N. R., & Green, A. A. (2025). Automated Assembly of Programmable RNA-Based Sensors. bioRxiv, 2025.08.12.669972. https://doi.org/10.1101/2025.08.12.669972